Clear Sky Science · en
IL2Pepscan: A machine learning framework for predicting IL-2 inducing peptides and their identification across global viral proteomes
Teaching the Immune System with Tiny Protein Fragments
Modern vaccines and cancer therapies increasingly rely on precisely nudging the immune system rather than carpet-bombing disease with drugs. This study explores how tiny fragments of proteins, called peptides, can be chosen to switch on a powerful immune messenger, interleukin‑2 (IL‑2). By using advanced computer models, the authors search through both known immune data and the protein catalogs of thousands of viruses to find peptide “needles” in a molecular haystack that may help design better vaccines and immunotherapies.
Why IL-2 Matters for Health and Disease
IL‑2 is a small signaling molecule that acts like a growth factor for key immune cells known as T cells. When these cells first encounter a threat—such as a virus or cancer cell—they can release IL‑2, which then encourages T cells to multiply, specialize, and remember the invader. IL‑2 also helps maintain regulatory T cells that keep the immune system from turning against the body’s own tissues. Because of this dual role, IL‑2 has been used as a drug to treat cancers like melanoma, and is being explored for autoimmune diseases. But giving IL‑2 directly can be harsh on patients, so there is growing interest in designing safe peptides that make the body produce IL‑2 in a more controlled, targeted way.
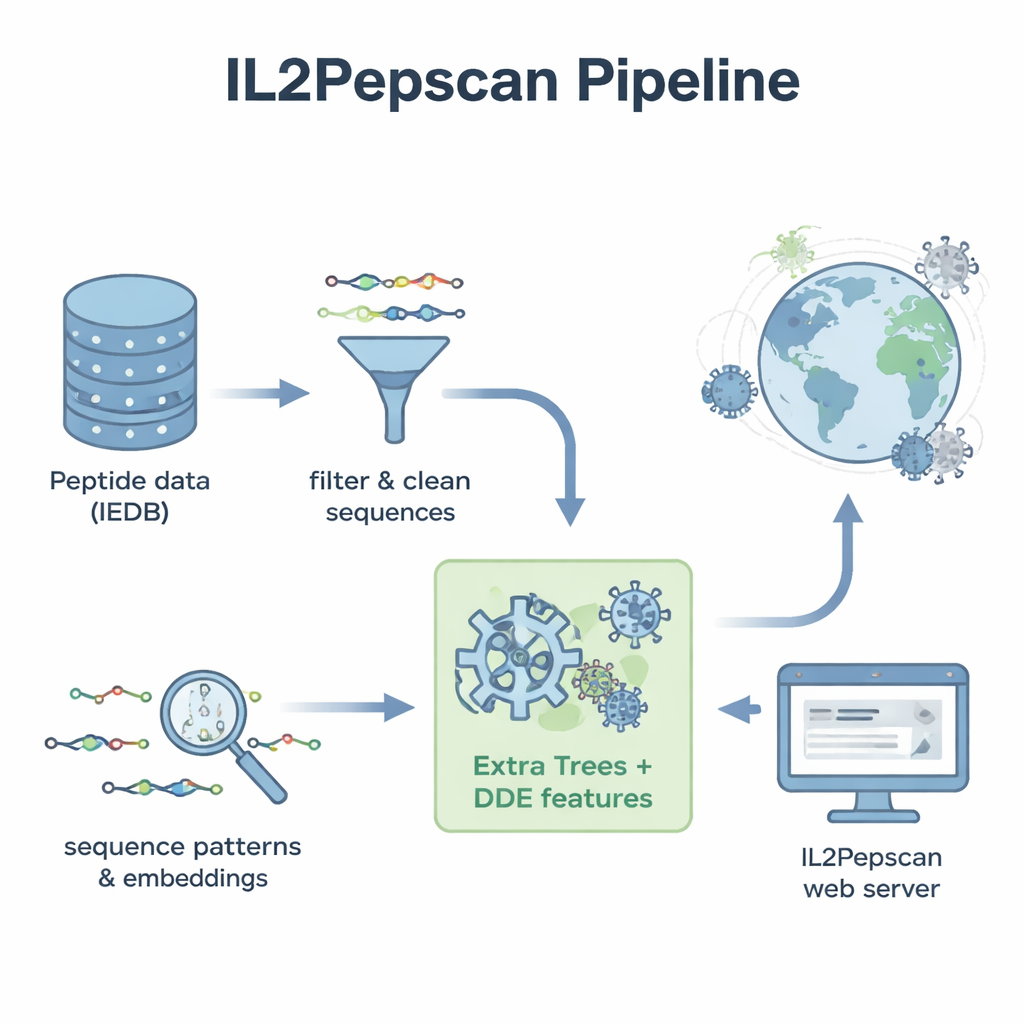
Learning the “Flavor” of IL-2–Inducing Peptides
The researchers began with thousands of peptide sequences that had already been tested in laboratory experiments and labeled as either IL‑2–inducing or not. They cleaned this dataset to remove duplicates, unusual building blocks, and peptides that were too short or too long, ending with over 6,000 well-characterized examples. By examining the building blocks (amino acids) that make up these peptides, they discovered clear differences between the two groups. IL‑2–inducing peptides tended to be richer in hydrophobic, or water‑repelling, amino acids like leucine and alanine, while non‑inducing peptides leaned toward more polar and charged residues. Certain short patterns, or motifs, such as “LEGS” and “ALEG,” appeared only in IL‑2–inducing peptides, hinting at structural signatures that may help trigger immune activation.
Training Machines to Spot Immune-Boosting Patterns
To turn these patterns into a practical prediction tool, the team converted each peptide into numerical descriptions that capture its composition and the order of its amino acids. They tested a range of machine‑learning methods—including popular algorithms like random forests, support vector machines, and boosted trees—along with deep‑learning architectures that are often used for language and image tasks. They also tapped into a large protein “language model” called ProtBERT, originally trained on hundreds of millions of protein sequences, and fine‑tuned it to better recognize IL‑2–related signals. After extensive testing with cross‑validation and an independent test set, the standout performer was a model called Extra Trees combined with a feature set known as dipeptide deviation from expected mean (DDE). This model achieved close to 80% accuracy and a strong correlation score, outperforming multiple deep‑learning approaches.
Scanning the Viral World for Hidden Immune Triggers
Armed with their best model, the authors cast a much wider net. They gathered reference protein sequences from more than 14,000 viruses, sliced these proteins into about 156 million overlapping peptides, and asked the model to predict which ones might induce IL‑2. Among the highest‑scoring candidates were peptides from well‑known viral families, including flaviviruses such as West Nile, Zika, Yellow Fever, and Hepatitis C viruses, as well as from Influenza and SARS‑CoV‑2. Many promising peptides came from viral envelope or nucleocapsid proteins—the same types of proteins that other studies have shown can provoke IL‑2 responses in animals. The model also flagged potential IL‑2‑inducing peptides encoded by bacteriophages, viruses that infect bacteria, hinting at an even broader landscape of immune‑relevant sequences.
From Algorithm to Accessible Tool
To make their work usable beyond the computing lab, the authors built a public web server called IL2Pepscan. Researchers can paste peptide or protein sequences into the site to estimate their IL‑2‑inducing potential, design new variants by mutating positions, scan entire proteins for hotspots, or search for known IL‑2–linked motifs. While the study does not yet experimentally confirm each predicted peptide, the agreement with existing laboratory findings suggests that IL2Pepscan can reliably narrow down candidates for further testing. For non‑specialists, the takeaway is that carefully trained algorithms can sift through huge biological datasets to pinpoint small protein fragments that may someday help vaccines and immunotherapies coax the immune system into a more powerful—and more precise—response.
Citation: Arora, P., Abhigyan, R., Periwal, N. et al. IL2Pepscan: A machine learning framework for predicting IL-2 inducing peptides and their identification across global viral proteomes. Sci Rep 16, 6701 (2026). https://doi.org/10.1038/s41598-026-35977-6
Keywords: interleukin-2, peptide vaccines, machine learning, viral proteome, immunotherapy