Clear Sky Science · en
Structural analysis of OCT4 binding to human LIN28B nucleosomes
How Cells Access Hidden Genetic Instructions
Every cell in your body carries the same DNA, yet only a fraction of those genetic instructions are used in any given cell type. Much of this control comes from how DNA is tightly packed around proteins called histones, forming structures known as nucleosomes. This tight packing can hide key stretches of DNA from the cellular machinery that reads genes. The study described here looks at how a special protein, OCT4, manages to find and bind its target DNA even when that DNA is wrapped and partly hidden inside nucleosomes, a process central to stem cell identity and cellular reprogramming.
Why Pioneer Factors Matter for Stem Cells
OCT4 belongs to a small but powerful group of proteins called pioneer transcription factors. Unlike most DNA-binding proteins, pioneers can slip into compacted, “off” regions of the genome and help turn genes on, playing a crucial role in shaping cell identity and in reprogramming adult cells back into stem-cell-like states. OCT4 works together with partners such as SOX2, KLF4, and c-MYC to induce pluripotency, the property that allows stem cells to become almost any cell type. Understanding exactly how OCT4 latches onto DNA wrapped in nucleosomes is essential for decoding how cells switch fates, and for designing more precise ways to manipulate cell identity in research and medicine.
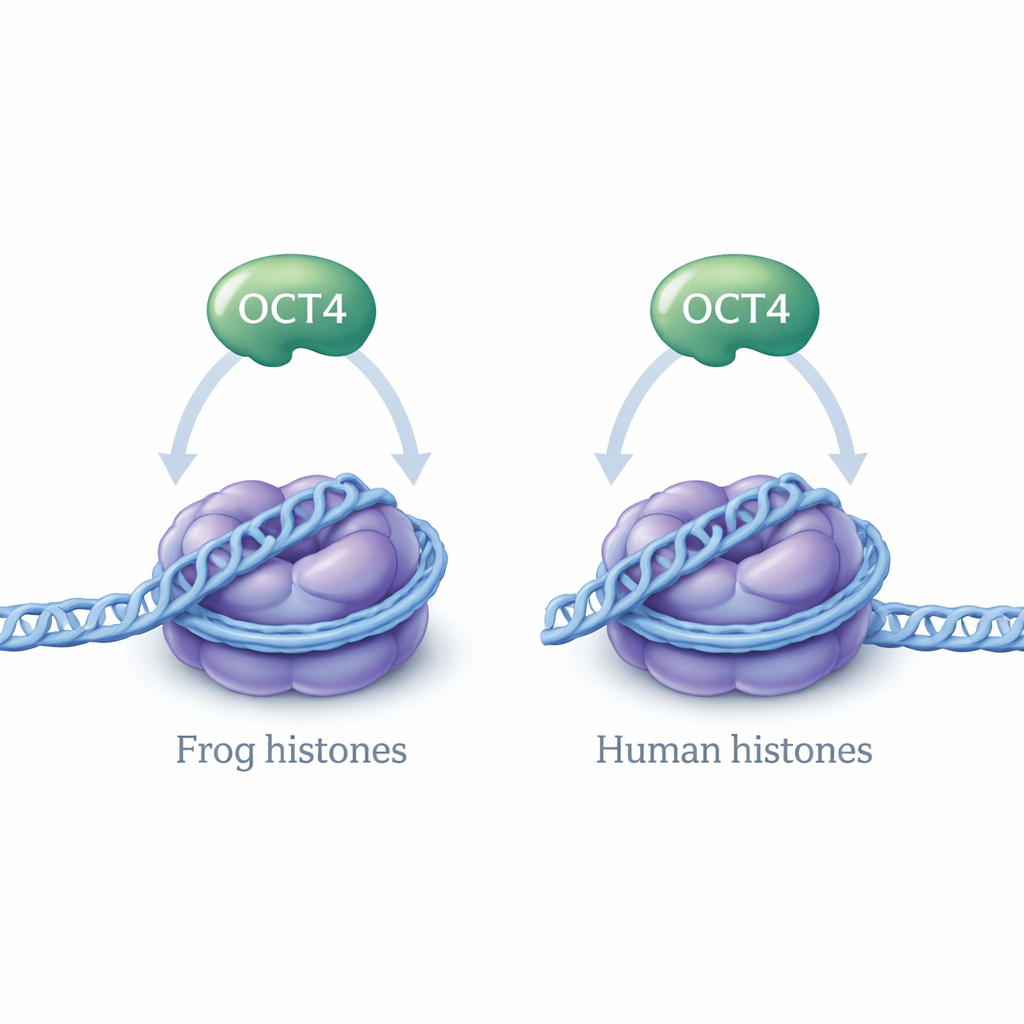
Testing Whether Species Differences Matter
Most structural studies of nucleosomes use histone proteins from either frogs (Xenopus laevis) or humans, because these proteins are very similar but not identical. Small differences in their amino acid sequences might, in principle, alter how DNA is wrapped or how regulatory proteins like OCT4 bind. In earlier work, the authors showed that OCT4 binds to a specific regulatory DNA sequence from the human LIN28B gene when that DNA is wrapped in nucleosomes made with frog histones. In the new study, they asked a simple but important question: does OCT4 behave the same way when the nucleosomes are built from human histones instead of frog histones?
Building and Probing LIN28B Nucleosomes
To answer this, the researchers reconstituted nucleosomes in the lab using a 182-base-pair segment of LIN28B DNA and either human or frog histone proteins, assembled by a careful “slow-dialysis” method that mimics conditions in cells. Gel-based assays confirmed that both types of histone octamers formed nucleosomes efficiently. They then examined how these nucleosomes sit on the LIN28B DNA. Both gel experiments and DNA-cutting followed by sequencing (MNase-seq) showed that the LIN28B DNA can adopt multiple positions on the histone core, and this behavior did not change if the nucleosomes were warmed to body temperature. Crucially, whether the histones came from frogs or humans made no noticeable difference to how the DNA was positioned or how tightly OCT4 bound.
Visualizing OCT4 on the Nucleosome
The team next used cryogenic electron microscopy (cryo-EM) to visualize the structure of OCT4 bound to LIN28B nucleosomes containing human histones. From about 15,000 particles, they reconstructed a three-dimensional map at roughly 6-angstrom resolution. The images revealed that both DNA-binding parts of OCT4 contact the same exposed stretch of DNA near the entry and exit point of the nucleosome, the so-called linker region. This is exactly the site and orientation previously seen when OCT4 was bound to nucleosomes made with frog histones. When the older model was fitted into the new cryo-EM map, the match was excellent, indicating that the overall architecture of the OCT4–LIN28B nucleosome complex is essentially identical for both species.
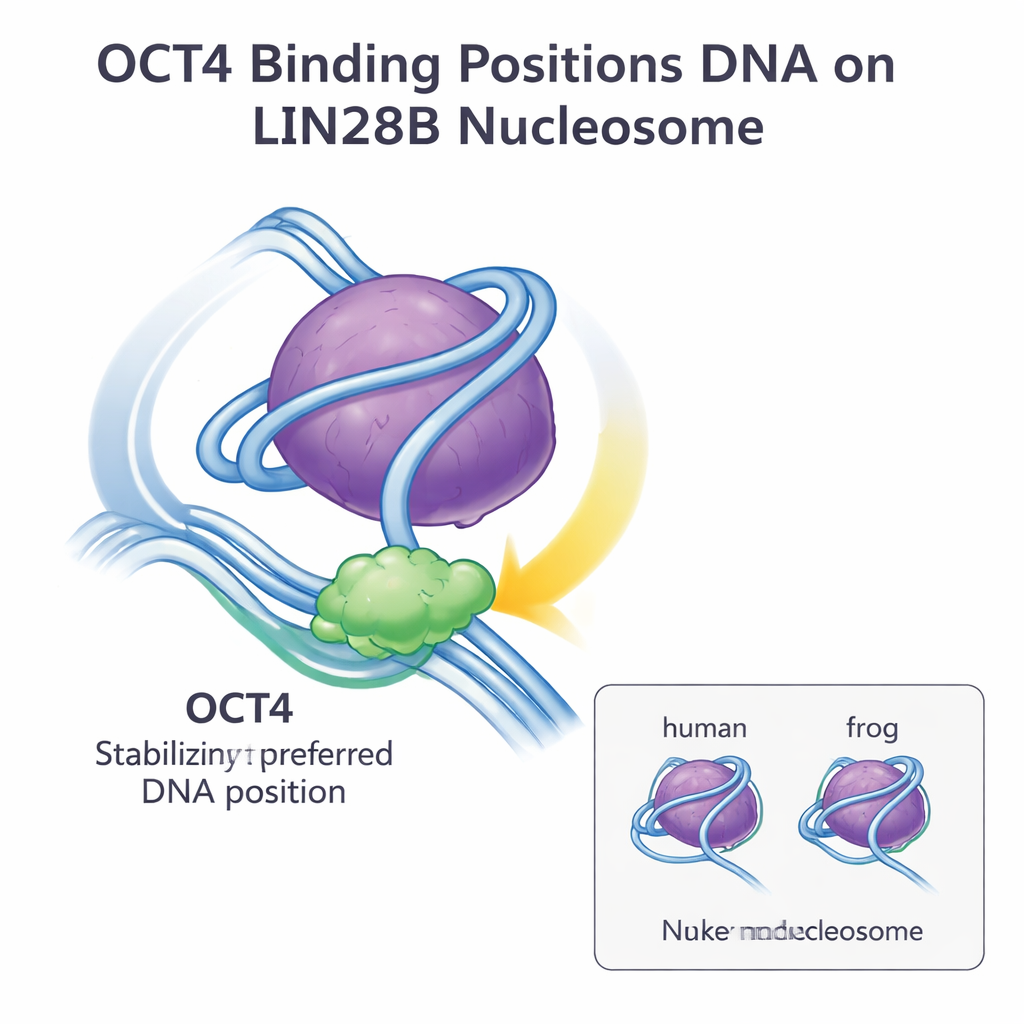
A General Strategy for Opening Chromatin
Together, these findings show that small sequence differences between frog and human histones do not alter how OCT4 recognizes and binds the LIN28B nucleosome. LIN28B DNA naturally sits in several possible positions on the histone core, but OCT4 binding selects and stabilizes one preferred position, making additional binding sites more accessible to OCT4 and its partners. This “DNA positioning and stabilization” strategy appears to be a general way that pioneer factors gain access to closed chromatin and promote cooperative binding of other regulators. For a lay audience, the bottom line is that the basic design of the nucleosome and the way key regulatory proteins like OCT4 work with it are highly conserved across species, reinforcing the idea that insights from model organisms can reliably inform our understanding of human gene control and stem cell biology.
Citation: Sinha, K.K., Halic, M. Structural analysis of OCT4 binding to human LIN28B nucleosomes. Sci Rep 16, 5704 (2026). https://doi.org/10.1038/s41598-026-35959-8
Keywords: pioneer transcription factors, OCT4, nucleosome structure, chromatin accessibility, stem cell gene regulation