Clear Sky Science · en
The functional organization of chromosome territories in single nuclei during zygotic genome activation
A Busy Neighborhood Inside Baby Cells
When a fertilized egg begins to develop, its DNA must spring to life in a matter of hours. During this awakening, called zygotic genome activation, each chromosome claims its own “territory” inside the cell nucleus. This study peeks into that crowded space, one nucleus at a time, to see how parental chromosomes find each other, how tightly they are packed, and how these physical changes relate to turning genes on and off—insights that may ultimately help us understand certain cancers and developmental disorders.
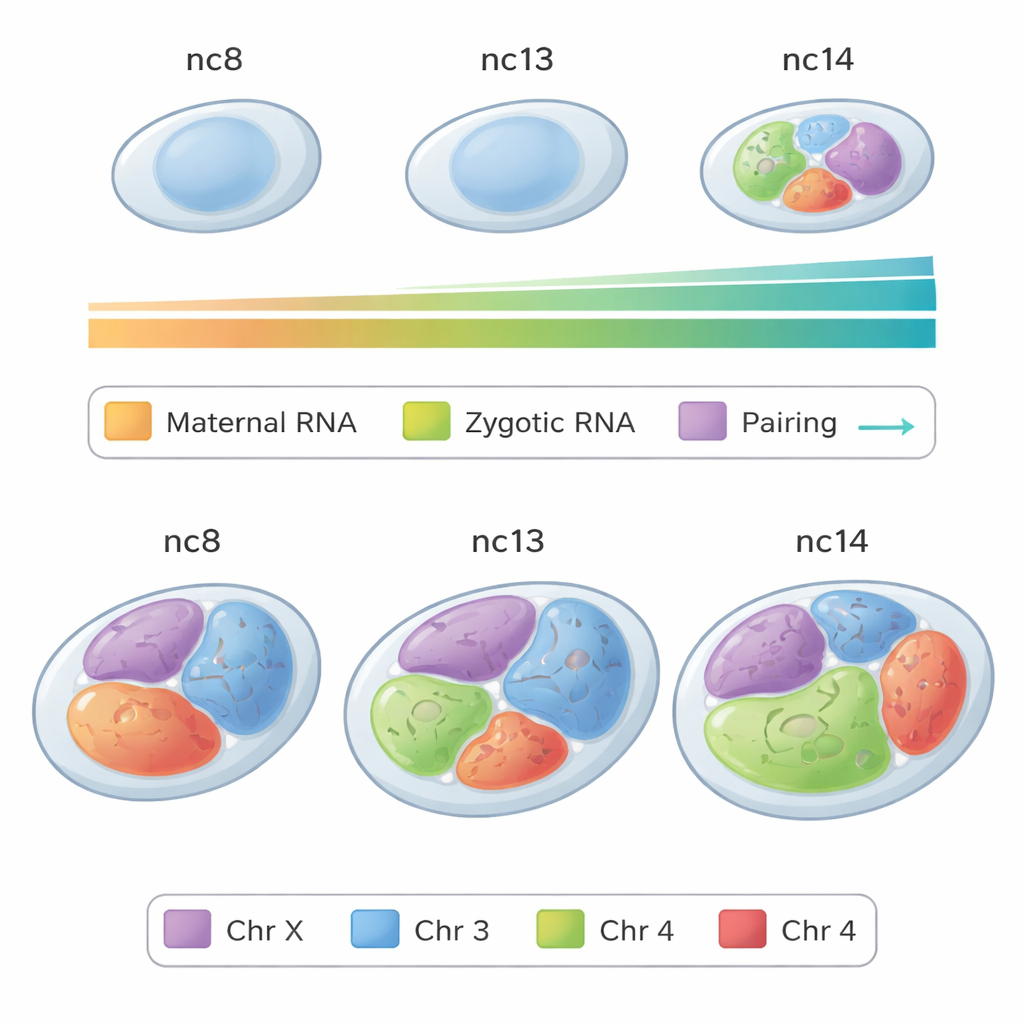
Chromosomes as Rooms in a Nuclear House
Rather than floating randomly, each chromosome tends to occupy its own zone, or chromosome territory, inside the nucleus. Using fruit fly embryos as a model, the researchers applied a high-resolution imaging method called Oligopaints to paint entire chromosomes and their arms in different colors. They focused on a critical window when the embryo’s own genes begin to take over from maternal instructions, moving from a minor early wave of gene activity to a major wave later on. By measuring the three-dimensional shape and size of these painted territories in hundreds of individual nuclei, they could watch the genome’s large-scale architecture change in real time as development progressed.
From Tightly Packed to More Open DNA
As embryos shifted from the minor to the major wave of genome activation, all major chromosomes became noticeably larger relative to the nucleus, and their shapes grew less perfectly round. At the same time, the territories for different chromosomes overlapped more with one another. These trends held both for entire chromosomes and for their individual arms. Such growth in volume, loss of compact spherical shape, and increased intermingling are hallmarks of more open, active chromatin—DNA that is easier for the cell’s machinery to read. In short, as the embryo turns on many more genes, its chromosomes loosen and spread out within the nuclear space.
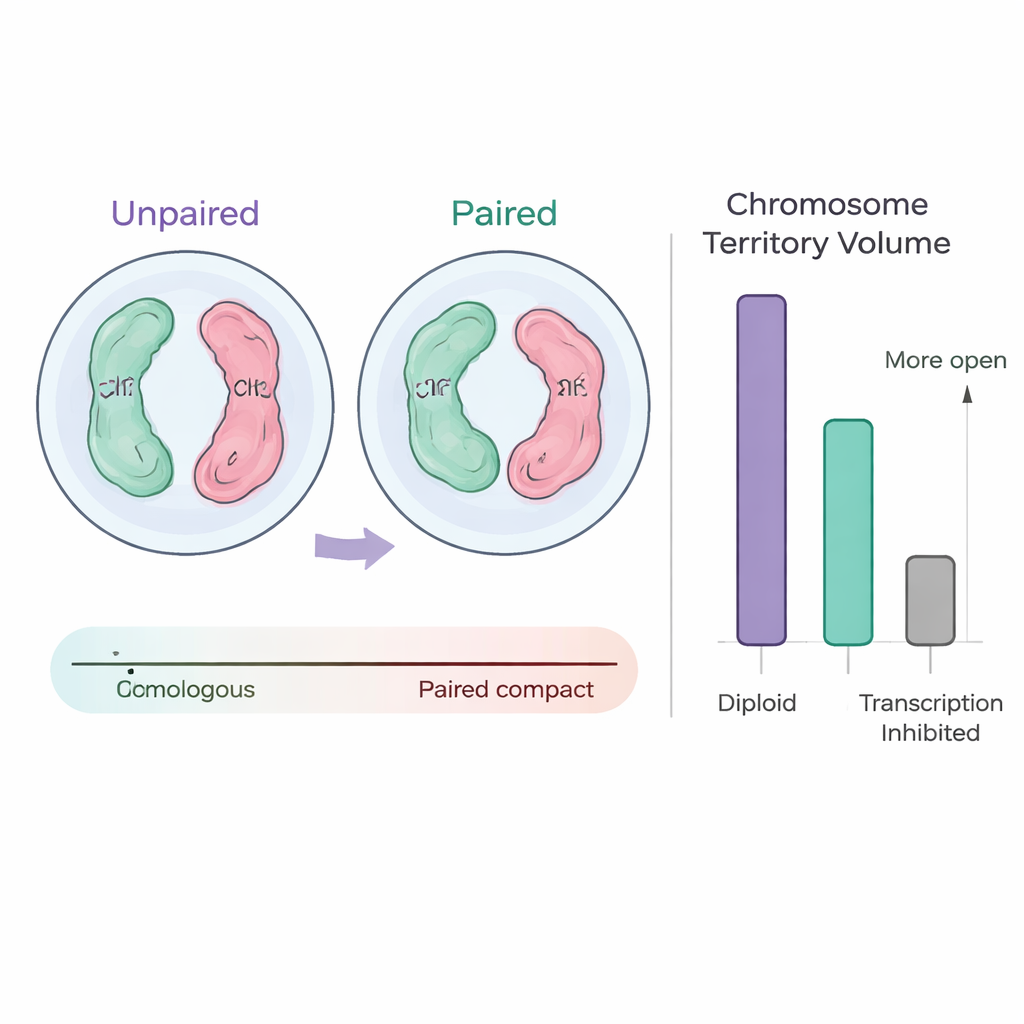
Parental Chromosomes Meet, But Not Perfectly
An important twist in this work is the focus on pairing between the two parental copies of each chromosome. At the scale of whole chromosomes, the maternal and paternal copies often appear as a single blended signal, meaning they are highly paired in the same region of the nucleus. Yet, when the team looked more closely at chromosome arms and centromeric regions, the pairing became less precise. Arms might be partially paired while the central region remained separate, or vice versa, and some arms showed signs of both tight and looser association. This suggests that although parental chromosomes come together globally, their fine-scale alignment is flexible and can adopt multiple configurations in different nuclei.
What Happens When Copies Are Missing or Genes Are Silenced
To test how these physical arrangements relate to gene activity, the researchers manipulated the system in two ways. In haploid embryos, which carry only one copy of each chromosome instead of two, all pairing between homologs is eliminated. These embryos have smaller nuclei, but in early stages their single chromosome territories occupy a relatively large fraction of the space and intermingle more, matching a period of unusually high RNA production. Later, as certain genes quiet down, both chromosome territories and specialized RNA polymerase II “hubs” shrink. In a complementary experiment, the team chemically blocked transcription in normal diploid embryos. Nuclear size stayed the same, but chromosome territories became smaller and more compact—consistent with reduced chromatin opening—while the overall level of pairing between homologs barely changed.
Why This Nuclear Choreography Matters
Taken together, the findings paint a picture of a highly dynamic nuclear interior during early development. Chromosome territories swell and intermingle as gene activity rises, and shrink again when transcription is dampened, but the tendency of parental chromosomes to pair at the whole-chromosome level remains surprisingly robust. This means that large-scale pairing is not simply a byproduct of active gene expression, yet changes in chromosome shape and packing do track with how much RNA the embryo is making. Understanding this choreography—how chromosomes fold, pair, and compact—may help explain why errors in chromosome number or structure can disrupt gene regulation and contribute to cancers and developmental diseases, and it offers a framework for thinking about therapies that target chromosome organization itself.
Citation: Shankar Ganesh, A., Orban, T.M., Raj, R. et al. The functional organization of chromosome territories in single nuclei during zygotic genome activation. Sci Rep 16, 5668 (2026). https://doi.org/10.1038/s41598-026-35953-0
Keywords: chromosome territories, zygotic genome activation, homolog pairing, 3D genome organization, Drosophila embryogenesis