Clear Sky Science · en
Machine learning framework for mRNA alternative splicing analysis identifies a signature of progression in colorectal adenocarcinoma
Why this research matters for patients
Colorectal cancer is one of the most common and deadly cancers, yet doctors still struggle to predict which patients’ tumors will quietly stay in check and which will come roaring back after treatment. This study introduces a new way to read hidden signals in tumor RNA – the messages cells use to make proteins – and uses machine learning to turn those signals into a simple risk score that may help tailor how aggressively each patient is treated.
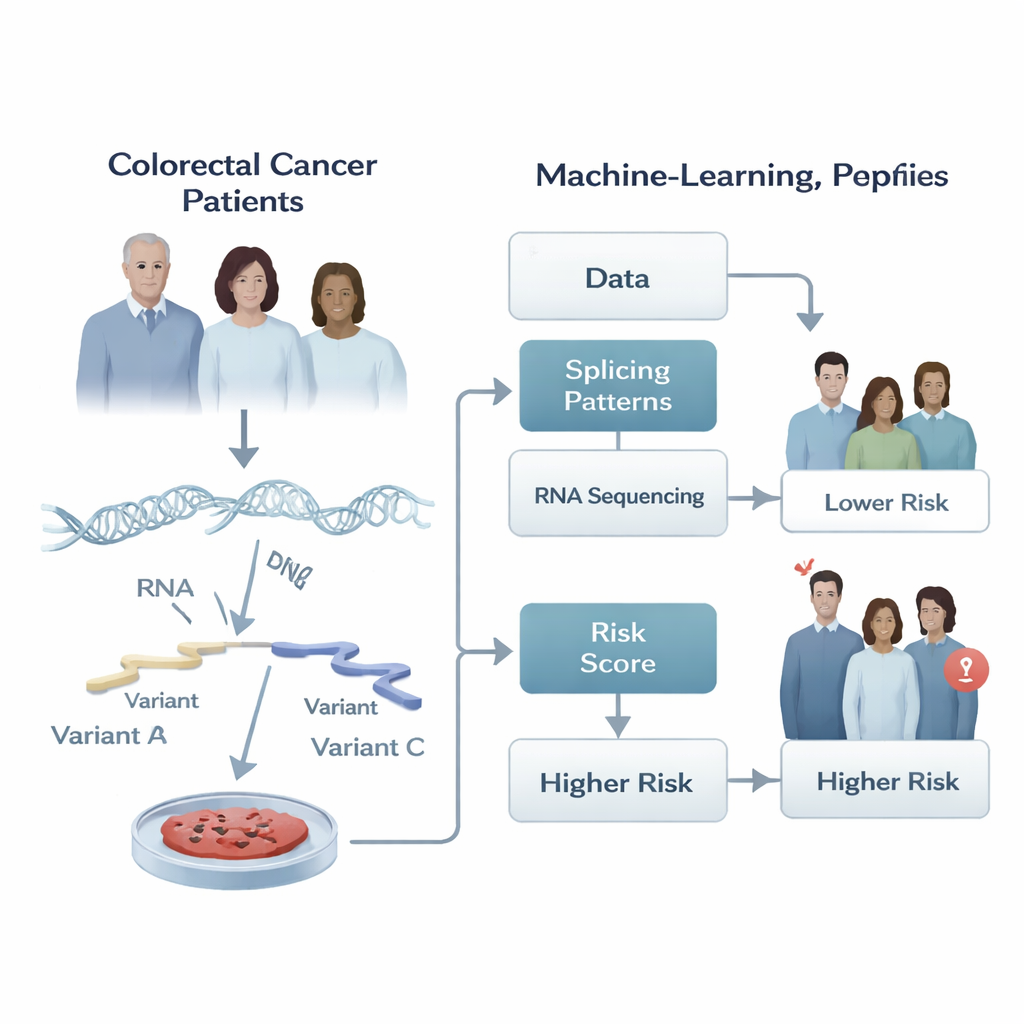
Hidden cuts and edits in cancer genes
Our genes are not read out in a fixed way. When a cell copies DNA into RNA, it can cut and paste pieces of the RNA message in different combinations, a process called alternative splicing. This editing lets a single gene produce several versions of a protein, like different tools from the same toolkit. In healthy cells, this flexibility is tightly controlled. In cancer, however, the cutting and pasting can go awry, creating versions of proteins that help tumors grow, spread, and resist treatment. The authors reasoned that the pattern of these RNA edits across a tumor might carry powerful clues about how that cancer is likely to behave over time.
Turning RNA patterns into a risk score
The researchers analyzed RNA sequencing data from tumors of 266 patients with colorectal adenocarcinoma from The Cancer Genome Atlas and another 348 patients from an independent study. For each tumor, they quantified how often particular splicing choices were used, summarizing them with a number between zero and one. They then built a stepwise machine learning pipeline that first screened thousands of splicing events for any link to how long patients stayed free of tumor progression, and then carefully narrowed this list down while avoiding redundant, overlapping signals. The end result was a compact “signature” of just five specific splicing events whose combined behavior best tracked whether a patient’s cancer progressed sooner or later.
Sorting patients into lower- and higher-risk groups
Using this five‑event signature, the team defined a numerical risk score for each patient by adding up the splicing measurements, weighted by how strongly each one related to progression. Patients whose tumors favored three of the splicing patterns tended to do worse, while two patterns were linked to better outcomes. The score neatly split patients into low- and high‑risk groups: in both the original cohort and the independent validation group, those with high scores experienced cancer progression significantly earlier. When the researchers plotted time-to-progression curves, the two lines separated clearly, indicating that this small set of RNA edits captured meaningful differences in tumor behavior across hundreds of individuals.
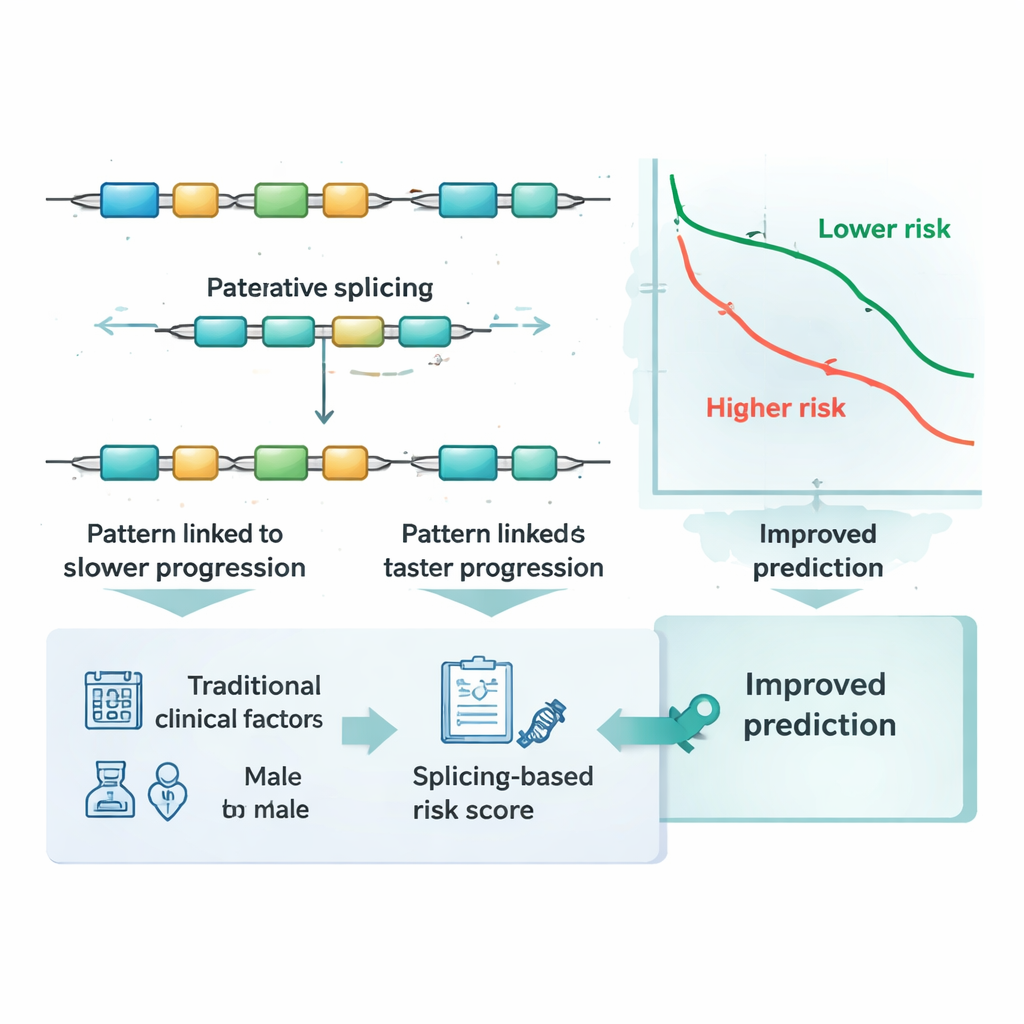
Beyond standard staging and known markers
Doctors currently rely on tumor stage, age, and other clinical features to estimate risk, and sometimes on specific DNA changes or gene activity levels. The researchers asked whether their splicing-based score added anything on top of these established measures. Using time‑dependent accuracy tests, they showed that predictions based only on stage, age, and gender were noticeably improved when the splicing risk score was included. They also compared the score to dozens of well-known molecular markers in colorectal cancer and to several common statistical modeling approaches. In both main patient groups, the five‑event splicing signature either matched or outperformed these alternatives, and improved prediction when used alongside them, suggesting it captures information that other markers miss.
What this could mean for future care
For a layperson, the key message is that the way a tumor “edits” its RNA can reveal how dangerous it is likely to be. This study shows that tracking just five specific RNA edits in colorectal tumors can sort patients into groups that differ meaningfully in their chances of remaining free from progression. While this work still needs to be translated into practical lab tests and evaluated in prospective clinical trials, it points toward a future in which doctors could use such a score at diagnosis to decide who needs more aggressive treatment and closer follow-up, and who might safely avoid overtreatment. More broadly, it offers a reusable framework for mining RNA splicing patterns in other cancers to refine prognosis and guide truly personalized therapy.
Citation: Maimekov, U., Nosrati, M., Mahmoud, A. et al. Machine learning framework for mRNA alternative splicing analysis identifies a signature of progression in colorectal adenocarcinoma. Sci Rep 16, 7106 (2026). https://doi.org/10.1038/s41598-026-35903-w
Keywords: colorectal cancer, alternative splicing, RNA sequencing, machine learning, cancer prognosis