Clear Sky Science · en
Disentangling direct and pleiotropic SNP effects in alfalfa (Medicago sativa L.) using causal graph learning
Why this matters for farms and food
Alfalfa is a workhorse of modern agriculture, feeding dairy cows and helping build healthy soils. Yet breeding better alfalfa—plants that stand strong through winter, resist damage, and provide high-quality feed—has been slowed by the sheer complexity of its genetics. This study introduces a new way to move from long, confusing lists of DNA markers to clear, cause-and-effect maps that show which pieces of the genome truly drive important stem traits, and which simply tag along for the ride.
From loose links to cause-and-effect
Traditional genome-wide association studies scan the genome for DNA variations, called SNPs, that tend to appear together with a trait, such as stem color or winter survival. In alfalfa, however, the situation is especially tangled: it has four copies of each chromosome, large stretches of DNA move together, and plants are highly mixed genetically. This creates a "fog of correlation" where many markers look important but only a few truly influence the trait. The authors argue that breeders need more than simple statistical links; they need to know which markers lie on the actual causal paths from genotype to visible plant traits.
How the new framework works
The researchers built a two-stage framework that combines modern machine learning with ideas from causal graph theory. First, they used a technique called Double Machine Learning to screen about 2,400 SNPs in 500 alfalfa genotypes. This step removes the influence of hidden factors such as family background and geography, using principal components of the genome as proxies. The result is a cleaner view of which markers still show a direct effect on traits like stem color after accounting for these confounding influences. In this filtered view, strong, stable peaks of signal appeared mainly on chromosomes 2 and 4, and key markers showed effect sizes whose confidence ranges clearly excluded zero, suggesting real causal influence.
Turning markers into genetic road maps
In the second stage, the team used a causal graph learning algorithm, known as the PC algorithm, to connect the most promising markers into a directional network. In these diagrams, nodes represent SNPs and the trait, and arrows show the most likely direction of influence. By trimming away edges that conflict with basic biology (for example, traits cannot change the underlying DNA) and keeping only SNPs that feed into the trait, the authors obtained compact, biologically sensible maps. These "sunflower" networks reveal a layered structure: an inner ring of Direct Parent SNPs that connect straight to the trait, and an outer ring of Upstream Hub SNPs that influence multiple parents but do not touch the trait directly.
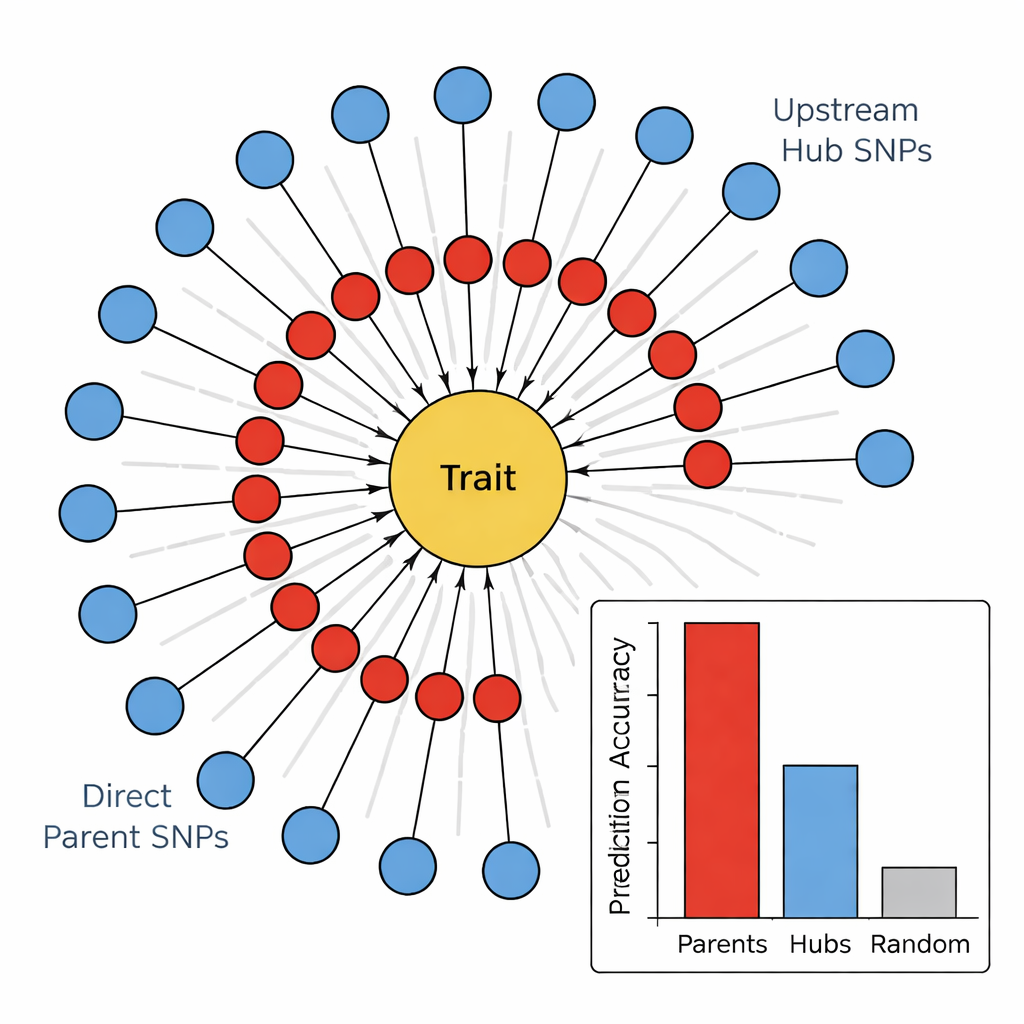
Executors versus directors in the genome
To test whether this hierarchy was meaningful, the authors compared how well different groups of markers could predict four stem-related traits: stem color, stem fill, stem strength, and winter injury. Across all traits, the Direct Parent SNPs were consistently the best predictors, often explaining several times more variation than either random markers or the Upstream Hubs. In contrast, the hubs showed weak or even negative predictive power, despite being highly connected in the network. When the team linked these SNPs to known genes, a pattern emerged: Direct Parents often matched enzymes or structural proteins that act directly on cell walls, pigments, or stress damage, while Hubs tended to correspond to transcription factors and regulatory proteins that broadly adjust many pathways at once.
What this means for future alfalfa breeding
For breeders and geneticists, the study offers a way to cut through noisy association results and focus on DNA changes that truly move the needle for specific traits. The authors show that combining de-confounded screening with causal graphs can serve as a built-in safeguard against overfitting, turning long candidate lists into small, interpretable networks aligned with known biology. In practical terms, Direct Parent SNPs become high-precision markers for selecting plants with better stems or winter survival, while Upstream Hubs point to master switches that might reshape broader stress responses, but with possible trade-offs. This structural view of the genome lays a foundation for more reliable genomic selection in complex crops and for integrating future layers of data, such as gene expression and metabolism, into coherent, cause-and-effect models of plant performance.
Citation: Lee, Y., Medina, C.A. & Xu, Z. Disentangling direct and pleiotropic SNP effects in alfalfa (Medicago sativa L.) using causal graph learning. Sci Rep 16, 5216 (2026). https://doi.org/10.1038/s41598-026-35876-w
Keywords: alfalfa genetics, causal graph learning, genomic selection, plant breeding, polyploid crops