Clear Sky Science · en
Lightweight SwiM-UNet with multi-dimensional adaptor for efficient on-device medical image segmentation
Smarter scans at the bedside
Brain scans can reveal life‑threatening tumors, but turning raw images into clear outlines that doctors can act on is still slow and computer‑hungry. This study introduces SwiM‑UNet, a new algorithm that can segment brain tumors from 3D MRI scans with state‑of‑the‑art accuracy while running efficiently on local devices, bringing precise image analysis closer to the patient’s bedside or even into mobile clinics. 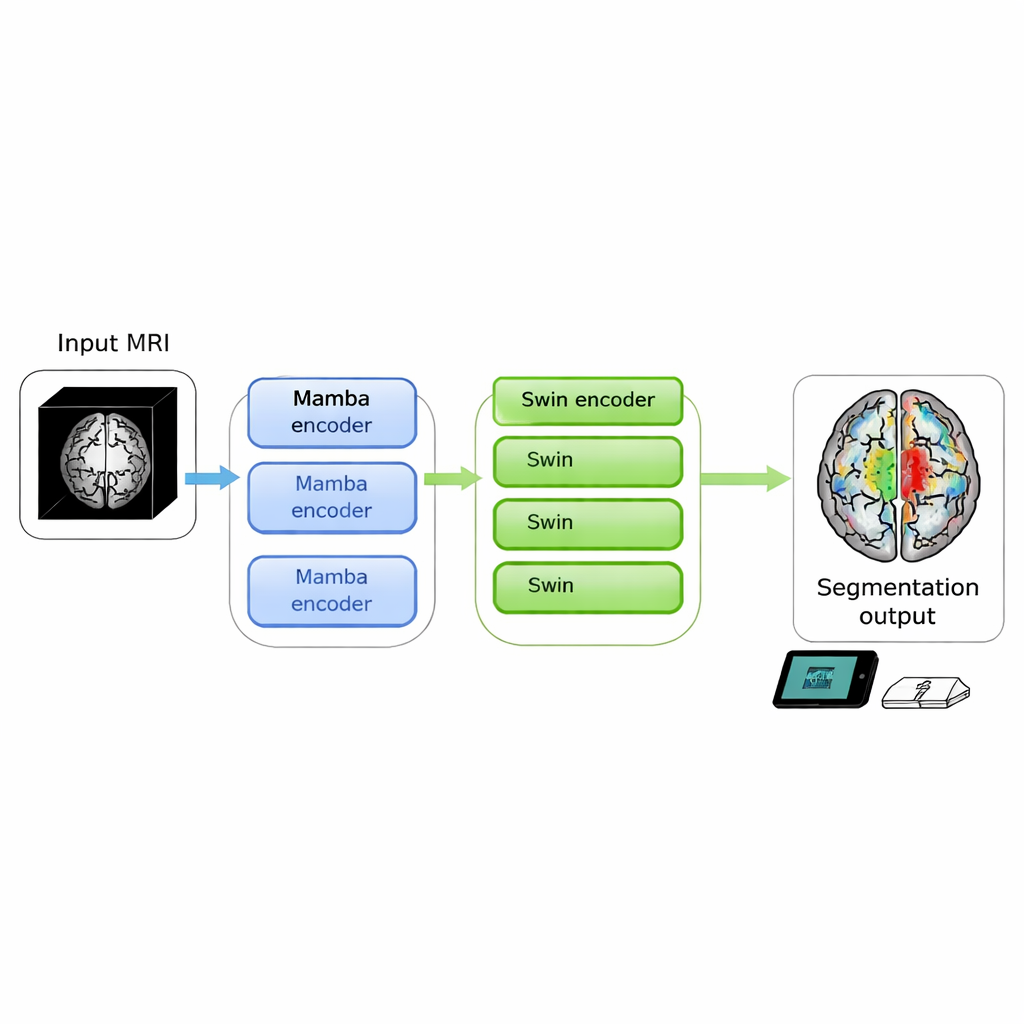
Why precise outlines of tumors matter
Modern medicine relies heavily on imaging to plan surgery, radiation therapy, and drug treatments. For brain tumors, doctors need not only to see that a tumor is present, but to know exactly where different parts of it begin and end. This task, called segmentation, is usually done or refined by specialists slice by slice—a time‑consuming process that can delay decisions and varies from one expert to another. Automated segmentation tools based on deep learning have improved this situation, but the most accurate ones often demand large graphics processors, making them hard to use directly in the clinic, especially on smaller or portable machines.
Two powerful ideas that don’t easily fit on small machines
Recent breakthroughs in computer vision come mainly from two families of models. Transformer‑based systems, such as the Swin Transformer, are excellent at seeing global patterns across an entire 3D scan and have powered top‑performing brain tumor segmenters. However, their core operation, called self‑attention, becomes extremely costly as image size grows, limiting their use in real‑time or on compact hardware. A newer family, known as Mamba and based on state‑space models, offers a clever workaround: it processes sequences in effectively linear time, reducing the number of computations needed. Early experiments in medical imaging showed that Mamba‑style models are fast and efficient but usually lag behind transformers in segmentation quality, especially when it comes to complex tumor shapes.
Blending speed and accuracy in one design
The authors set out to merge the strengths of both worlds in a single 3D model. Their SwiM‑UNet keeps the familiar U‑shaped structure widely used in medical imaging, with an encoder that compresses information and a decoder that rebuilds detailed segmentations. In the early stages, where the scan is still large and high‑resolution, they use efficient Mamba blocks to keep computations manageable. Deeper in the network, after the data have been downsampled, they switch to streamlined Swin Transformer blocks that can now afford to model long‑range relationships without overwhelming the hardware. A custom bridge called the MS‑adapter links these two regimes. It looks at features along the width, height, and depth of the volume separately, and also across channels, then learns through small gating units how much each view should influence the final representation. 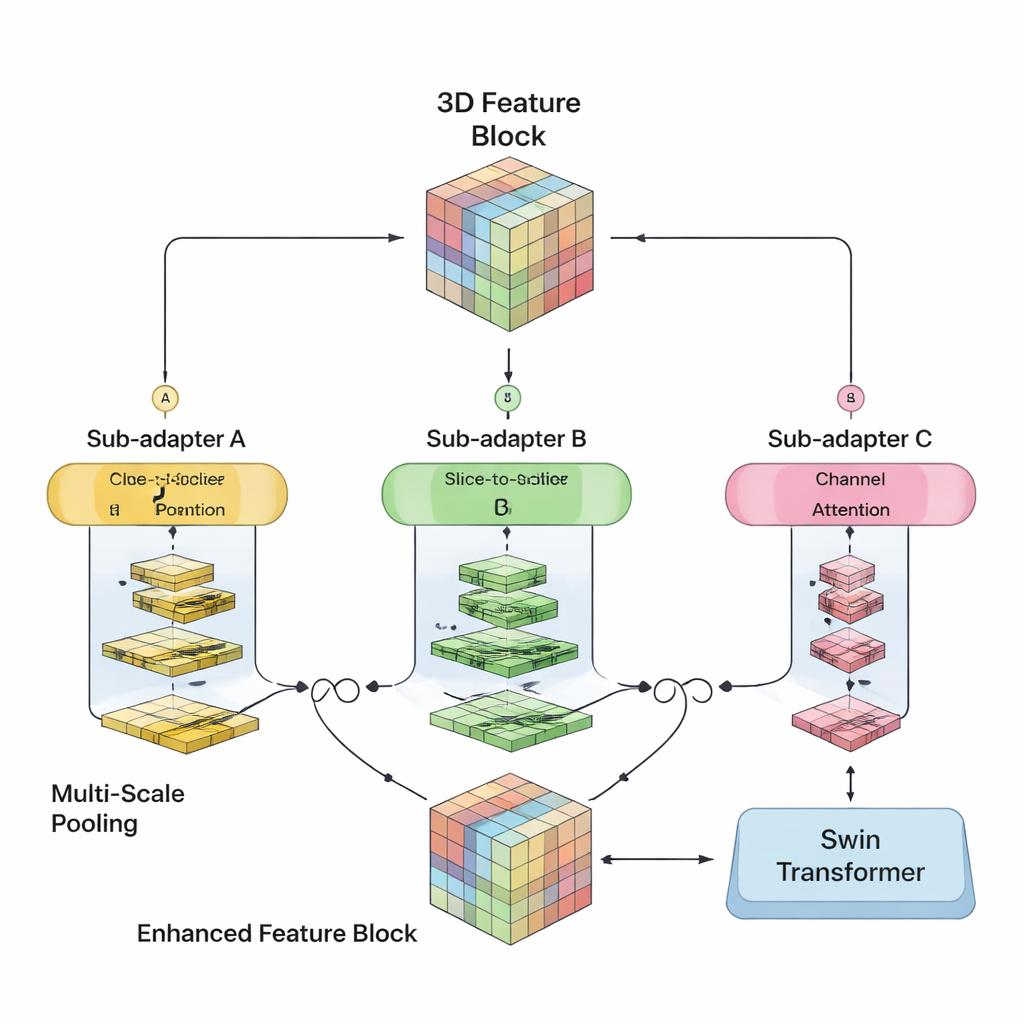
Doing more with less computation
Beyond this hybrid layout, the team trims excess computation by using low‑rank fully connected layers and a decoder that deliberately reduces the number of channels it needs to handle. They tested several variants with different mixes of Mamba and Swin layers and found that using Mamba in the first three encoder stages and Swin only at the deepest stage gave the best balance of speed and accuracy. On two large public brain tumor datasets from the BraTS 2023 and 2024 challenges, SwiM‑UNet achieved higher accuracy and more precise boundaries than leading transformer‑only, Mamba‑only, and previous hybrid models, while using far fewer floating‑point operations and cutting inference time to around 45 milliseconds per scan patch on a modern graphics card.
Ready for real‑world devices
To check whether these gains matter outside the lab, the authors compared the model’s computational demands with the capabilities of typical clinical edge systems—portable MRI consoles, point‑of‑care computers, and operating‑room workstations. Their analysis suggests that, unlike heavier transformer models, SwiM‑UNet fits comfortably within the power, memory, and speed limits of such devices, often meeting real‑time requirements. It also performed strongly on a separate abdominal CT dataset, indicating that the approach can generalize beyond brain tumors and even beyond MRI.
What this means for patients and clinicians
In practical terms, SwiM‑UNet shows that it is possible to approach the accuracy of the most sophisticated segmentation models while keeping computation light enough for on‑device use. That could enable faster, more consistent tumor outlines in emergency rooms, rural hospitals, or mobile imaging units without sending sensitive scans to distant servers. While further work is needed to adapt the method to different scanners and conditions, this hybrid design points toward a future in which high‑quality image analysis runs wherever the patient is, not just in data centers.
Citation: Noh, Y., Lee, S., Jin, S. et al. Lightweight SwiM-UNet with multi-dimensional adaptor for efficient on-device medical image segmentation. Sci Rep 16, 5807 (2026). https://doi.org/10.1038/s41598-026-35771-4
Keywords: brain tumor segmentation, medical imaging AI, hybrid neural networks, on-device inference, 3D MRI analysis