Clear Sky Science · en
Structured dissociative PCA methods for high dimensional neuroimaging signal decomposition
Untangling the Brain’s Hidden Signals
Modern brain scans can record activity from hundreds of thousands of locations every second, but turning this torrent of numbers into clear, meaningful networks is extremely hard. Different brain processes often overlap in space and time, like multiple radio stations broadcasting on nearby frequencies. This paper presents new mathematical tools that help separate these tangled signals more cleanly, promising sharper maps of brain function for both basic research and clinical studies.
Why Usual Methods Fall Short
For decades, researchers have relied on techniques called principal component analysis (PCA), sparse PCA, and independent component analysis (ICA) to compress and separate functional MRI (fMRI) data. PCA finds patterns that explain most of the variation in the data, but each pattern mixes information from almost every brain location, making interpretation difficult. Sparse PCA tries to fix this by forcing each pattern to use only a subset of locations, and ICA goes further by assuming the underlying brain signals are statistically independent. In practice, however, real brain networks overlap and influence one another. When that happens, the independence and sparsity assumptions break down. The result can be fragmented maps and noisy time courses that no longer match how brain networks actually behave.
A New Way to Pull Signals Apart
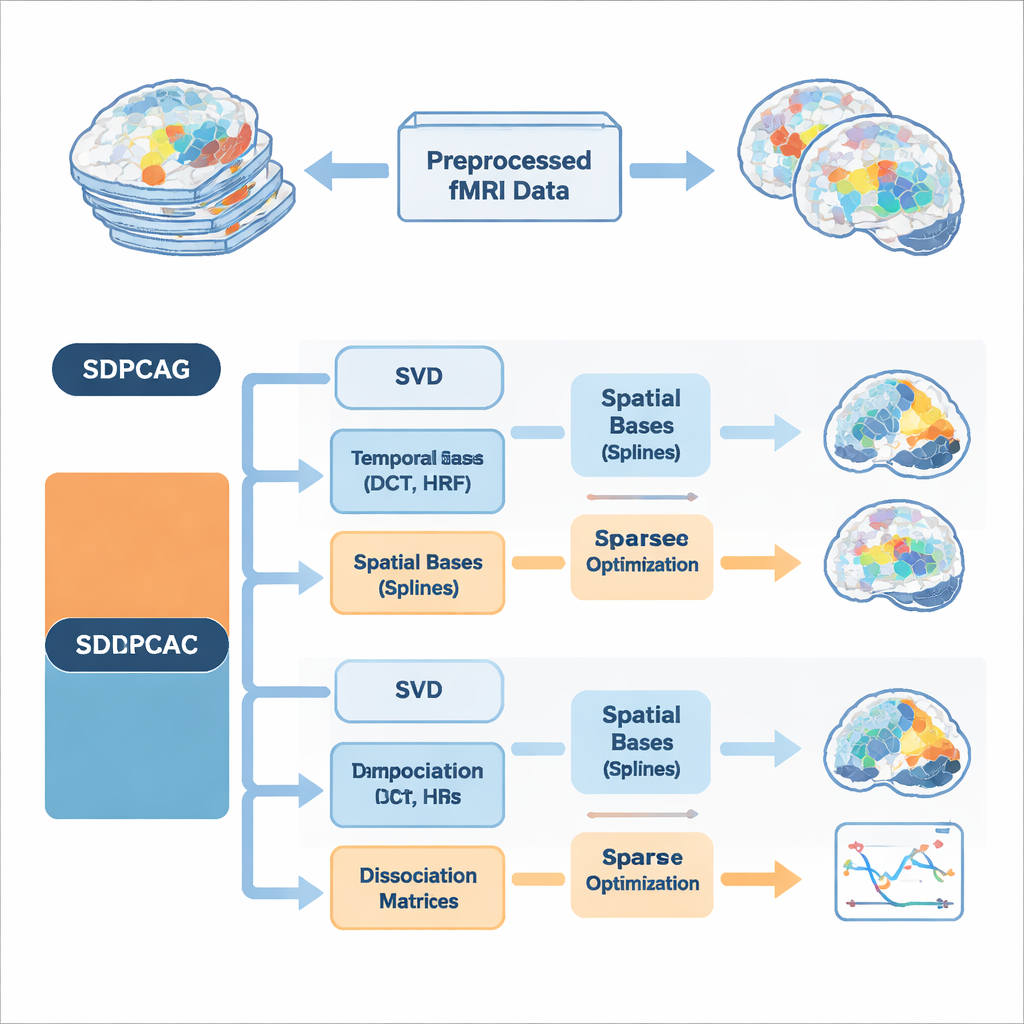
The authors propose a unified framework called structured dissociative PCA, implemented in two algorithms named SDPCAG and SDPCAC. Instead of treating space and time separately, the method reshapes the entire fMRI dataset through a single, carefully designed decomposition. It starts from a standard low-rank breakdown of the data, then introduces special "dissociation" matrices that reweight and rotate both spatial maps and time courses together. At the same time, it represents these components using sets of smooth building blocks: cosine-like temporal waves, models of the brain’s blood-flow response to neural activity, and smooth spatial curves called splines. By learning how to combine these building blocks while enforcing sparsity—only keeping the most important pieces—the method can tease apart overlapping networks without imposing unrealistic independence.
Building in Brain Knowledge from the Start
A key innovation is that the algorithms bake prior knowledge about brain signals directly into the decomposition, rather than cleaning things up afterward. On the temporal side, they use discrete cosine functions to encourage smooth time courses and include standard models of the hemodynamic response, the delayed blood-oxygen signal measured by fMRI. On the spatial side, spline bases help favor contiguous, coherent activation patterns instead of scattered dots. Extra constraints limit how many temporal and spatial basis functions each component may use, which reduces overfitting to noise and mirrors the idea that real brain networks are relatively compact in both space and time. Two complementary optimization strategies are presented: SDPCAG updates whole matrices in blocks, while SDPCAC refines one component at a time using residual errors, trading slightly higher cost for more flexible adjustments.
Putting the Method to the Test
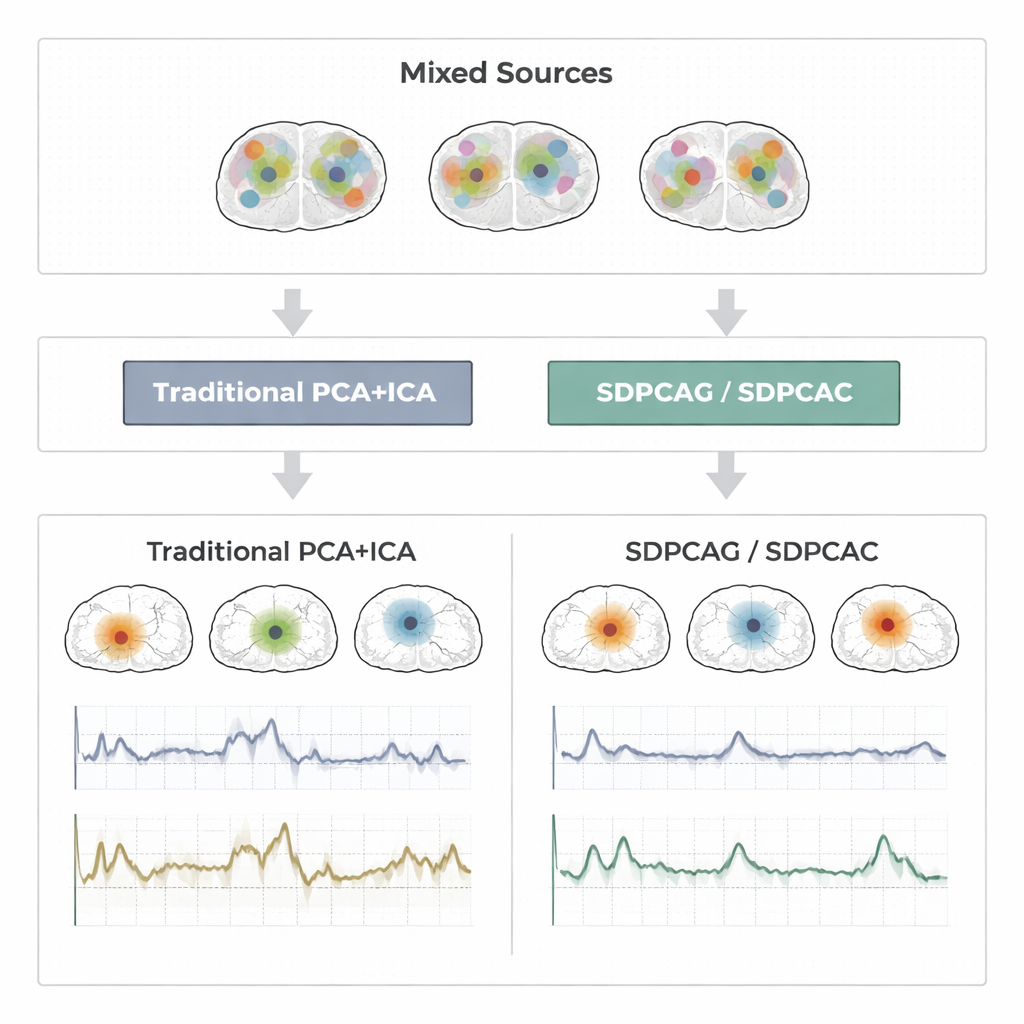
To judge how well these ideas work, the authors run extensive tests on three types of data: carefully designed synthetic fMRI signals with known ground truth; a block-design motor experiment from the Human Connectome Project; and an event-related finger-tapping study from an independent lab. Across these settings, they compare SDPCAG and SDPCAC against leading alternatives that combine penalized matrix decomposition, sparse PCA, ICA, and dictionary learning. They measure how closely recovered time courses match known task patterns, how well spatial maps line up with established resting-state networks, and how accurately sources are recovered under varying levels of noise. The new methods consistently produce cleaner, more localized brain maps and less noisy time series, maintaining strong performance even when the data are heavily corrupted. One of the algorithms, SDPCAG, improves source recovery accuracy by about 22% over a strong competing method, while also running faster than its more detailed sibling SDPCAC.
What This Means for Brain Research
In plain terms, this work offers a better way to "unmix" the signals inside fMRI data. By jointly modeling space and time, using realistic priors about how brain activity and blood flow behave, and enforcing smart sparsity, SDPCAG and SDPCAC produce brain networks that are both sharper on the images and truer to their underlying time courses. This can lead to more reliable detection of task-related activations and more precise mapping of resting-state networks, which in turn supports studies of conditions such as Alzheimer’s disease, psychiatric disorders, and other brain illnesses. While there is still room to speed up and extend the approach—for example to multi-subject studies or multimodal imaging—it represents a significant step toward turning high-dimensional scan data into trustworthy, interpretable pictures of the working human brain.
Citation: Khalid, M.U., Nauman, M.M., Rehman, S.U. et al. Structured dissociative PCA methods for high dimensional neuroimaging signal decomposition. Sci Rep 16, 6911 (2026). https://doi.org/10.1038/s41598-026-35764-3
Keywords: fMRI signal decomposition, sparse PCA, brain network mapping, blind source separation, resting-state connectivity