Clear Sky Science · en
Prevalence of antibiotic resistance gene in different wastewater treatment systems and effluent-irrigated soils through metagenomic analysis
Why the Water Coming Out of Treatment Plants Still Matters
Antibiotics have saved countless lives, but the microscopic genes that make bacteria resistant to these drugs do not disappear when we flush them away. This study looks at what happens to antibiotic resistance genes as wastewater passes through two modern treatment systems, and what occurs when the cleaned water is then used to irrigate soil in a dry region of China. The findings show that even well‑run plants can become hubs where resistance genes change, spread, and enter the wider environment.
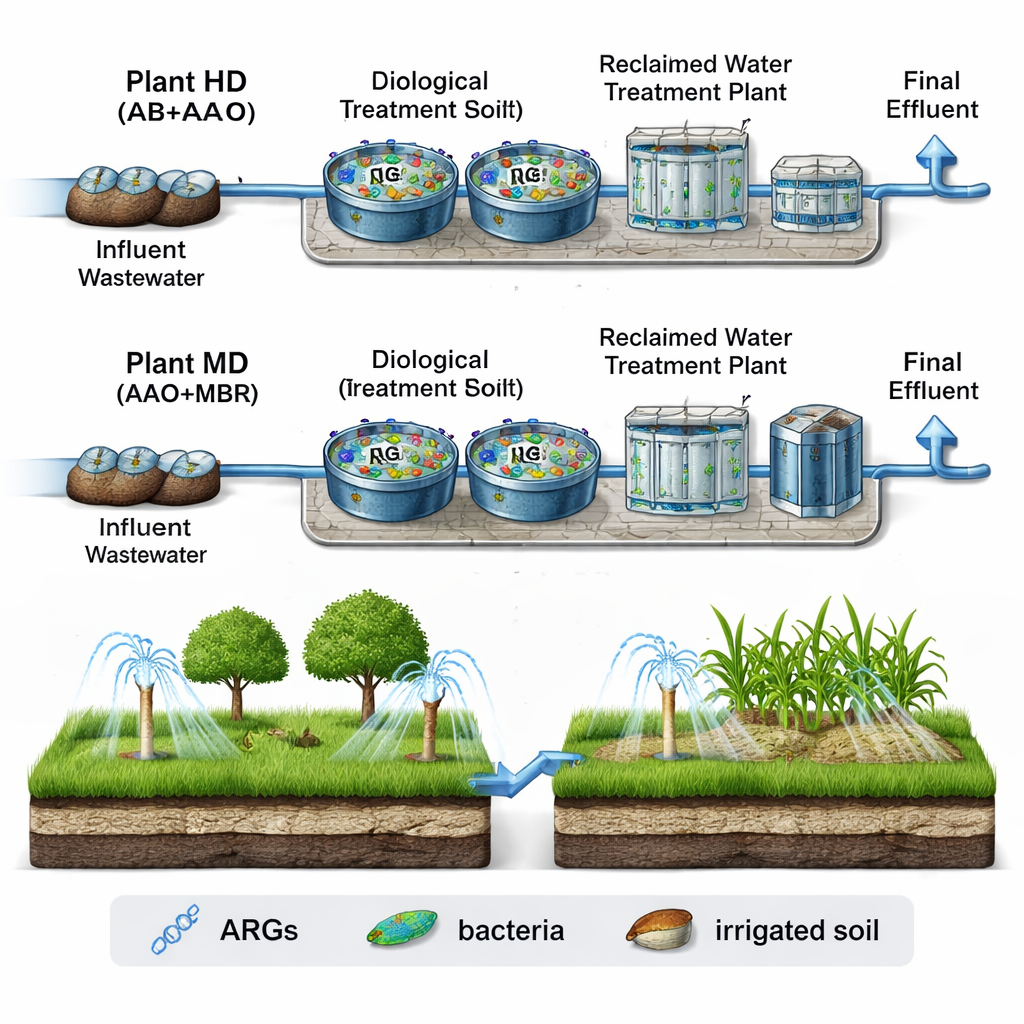
From Sewers to Sprinklers
The researchers examined two full‑scale wastewater treatment systems, called HD and MD, in the city of Urumqi. Both receive a mix of household, hospital, and industrial wastewater, but they use different process chains. One combines an adsorption–bio‑oxidation step with an anaerobic–anoxic–oxic tank system; the other uses the same three‑zone biology paired with a membrane bioreactor. In both cases, a separate reclaimed water plant polishes the partially treated water before it is used to irrigate parks, lawns, and roadside greenery. By sampling the incoming sewage, the biologically treated water, the final effluent, and nearby soils in both winter and summer, the team could track how resistance genes move along this entire path.
Genes that Defy Many Medicines
Using metagenomic sequencing—a method that reads the combined DNA of all microbes in a sample—the team cataloged 31 types of antibiotic resistance genes. The clear leaders were genes that protect bacteria from multiple drug classes at once, followed by genes that resist tetracyclines, macrolides, and aminoglycosides, which are widely used in human and veterinary medicine. Certain specific gene subtypes, such as msrE, mphE, and ANT(6)-Ia, were especially common in raw sewage and remained detectable even after treatment. Interestingly, the overall mix of resistance types changed little between winter and summer, although winter samples tended to carry slightly more resistance genes and a wider variety of them.
Treatment Helps—but Also Reshapes the Problem
The biological tanks in these systems are designed to break down organic pollution, but they also create dense, nutrient‑rich communities of bacteria under constant pressure from leftover antibiotics and even heavy metals. In the HD system, this stage actually increased the total amount and variety of resistance genes, turning the tanks into hotspots of genetic exchange. In the MD system, biological treatment reduced total gene abundance somewhat, but in both plants some key genes survived. The final polishing step—using processes such as additional biological treatment, membranes, and disinfection with ultraviolet light or ozone—did lower the overall number of resistance genes in the effluent. Yet this step could also favor new gene subtypes that cope well with these conditions, meaning total gene counts fell while diversity sometimes rose.
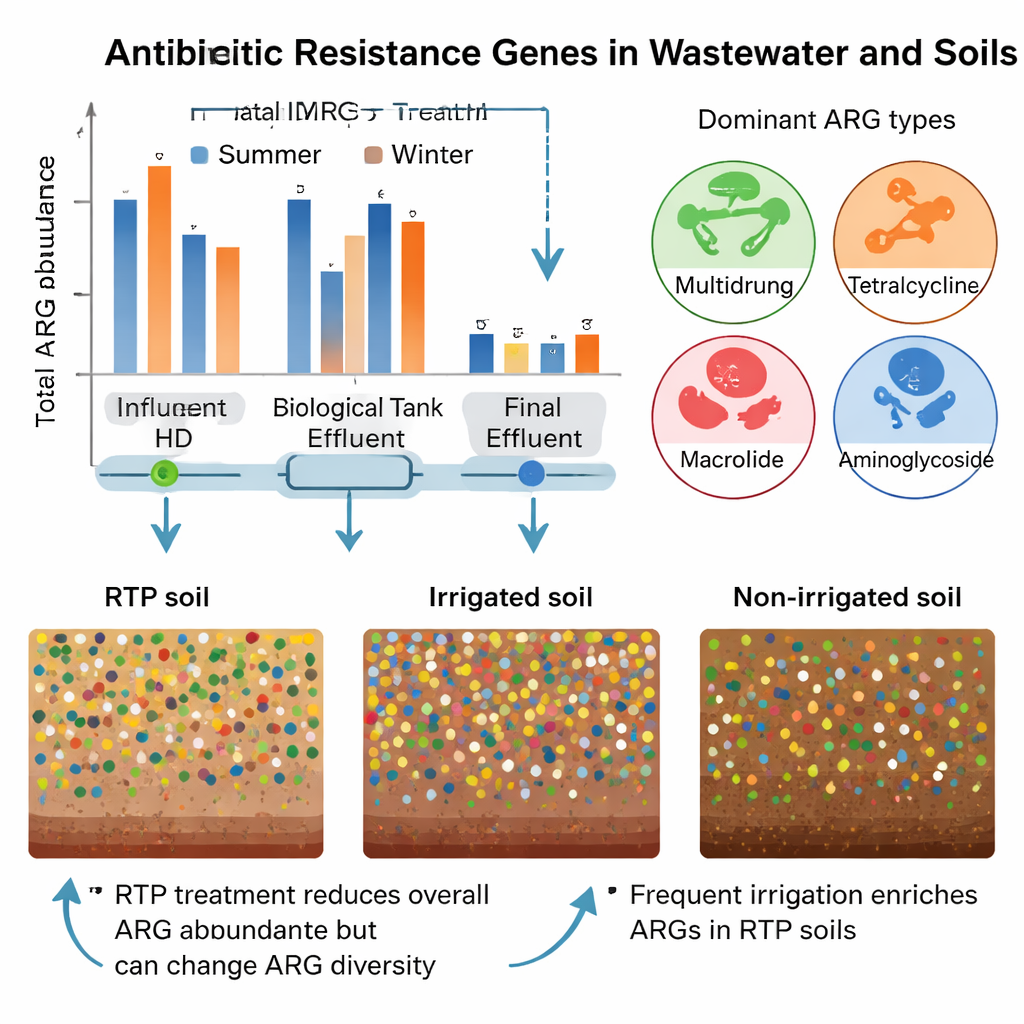
What Happens When Effluent Reaches the Soil
To see how reuse of treated water affects land, the researchers compared soils with different irrigation histories: within the reclaimed water plant itself, in nearby green spaces regularly irrigated with effluent, and in a reference area that had never received this water. All soils contained resistance genes, reflecting both natural background and past human activities. Soils within the plant, which were irrigated most often, had the highest gene abundance and diversity. Surprisingly, total resistance in irrigated park soils was not dramatically higher than in non‑irrigated soils over the short study period. However, the makeup of the resistance “community” shifted: some specific genes, such as certain tetracycline resistance subtypes, were more common in irrigated plots, hinting that reclaimed water nudges the soil toward particular resistance profiles rather than simply increasing all genes at once.
Who Carries These Genes—and Why That Matters
The team also mapped which bacterial groups tended to carry which genes. Common wastewater genera such as Arcobacter, Acinetobacter, Pseudomonas, and Sphingomonas showed strong links to multiple resistance genes, including those that fend off many drugs at once. In both water and soil, some genes were tied to several different bacterial hosts, suggesting that horizontal gene transfer—the swapping of DNA between microbes—is an important driver of spread. When the scientists compared resistance genes with measured antibiotic levels, they found both positive and negative correlations. This indicates that while antibiotics can clearly favor resistant microbes, other environmental pressures, like metals or general pollution, also help shape where and how these genes persist.
What This Means for Everyday Life
For a non‑specialist, the main message is that wastewater treatment plants do reduce antibiotic resistance genes, but they do not erase them. Some treatment steps can even enrich certain genes before later stages bring totals back down. When the resulting effluent is used to irrigate parks and other green spaces—as is increasingly common in water‑scarce regions—resistance genes can accumulate in nearby soils and may slowly change the invisible microbial community underfoot. The study suggests that improving the final polishing and disinfection steps, closely monitoring key resistance genes, and paying special attention to frequently irrigated sites could all help curb the environmental spread of antibiotic resistance without abandoning the valuable practice of water reuse.
Citation: Fang, H., Pu, M., jiang, A. et al. Prevalence of antibiotic resistance gene in different wastewater treatment systems and effluent-irrigated soils through metagenomic analysis. Sci Rep 16, 5167 (2026). https://doi.org/10.1038/s41598-026-35758-1
Keywords: antibiotic resistance genes, wastewater treatment, reclaimed water irrigation, soil microbiome, water reuse