Clear Sky Science · en
Prognostic impact of spatial niches in prostate cancer
Why the tumor’s “neighborhoods” matter
Prostate cancer is common, but not all tumors behave the same way. Some grow slowly, while others progress quickly and threaten patients’ lives. Doctors currently rely on markers like PSA levels and microscope-based grading to guess which tumors are dangerous. This study asks a new question: can the exact place where cancer cells live inside a tumor—the busy outer rim versus the deeper core—help predict how a patient will do?
Seeing the tumor as a small ecosystem
Modern research shows that a tumor is not a uniform ball of identical cells. Instead, it is more like a small ecosystem with distinct “neighborhoods,” or spatial niches. In prostate cancer, the inner core of the tumor and the outer edge where cancer meets normal tissue can host different mixes of cancer cells, immune cells, and supporting structures. The authors used a technology called digital spatial profiling, which lets them measure dozens of proteins directly in thin slices of stored tumor tissue, while keeping track of exactly where in the tumor each measurement came from.
Measuring proteins in tumor center and edge
The team analyzed tumor samples from 49 men with high-risk prostate cancer who had undergone surgery. For each tumor, they selected regions in the center and at the periphery and measured the levels of 46 different proteins related to immune cells, cell death, and growth signals. This produced tens of thousands of data points. When they looked at the tumor center and periphery separately, they confirmed that these areas behave differently: many proteins, especially those linked to immune activity, were more abundant at the tumor’s outer edge, while certain cell-death regulators were higher in the center.
When separate views fall short
The researchers then asked whether protein patterns in just the center or just the periphery could predict how long patients would remain free from disease progression after surgery. Using statistical methods to group patients based on protein patterns in each compartment alone, they found two main patient clusters for both the center and the periphery. However, neither set of clusters lined up with how patients actually fared over time, nor did they correlate clearly with classic risk factors such as Gleason score or known gene mutations like BRCA1/2 and TP53. In other words, looking at either neighborhood in isolation did not yield a useful new prognostic marker.
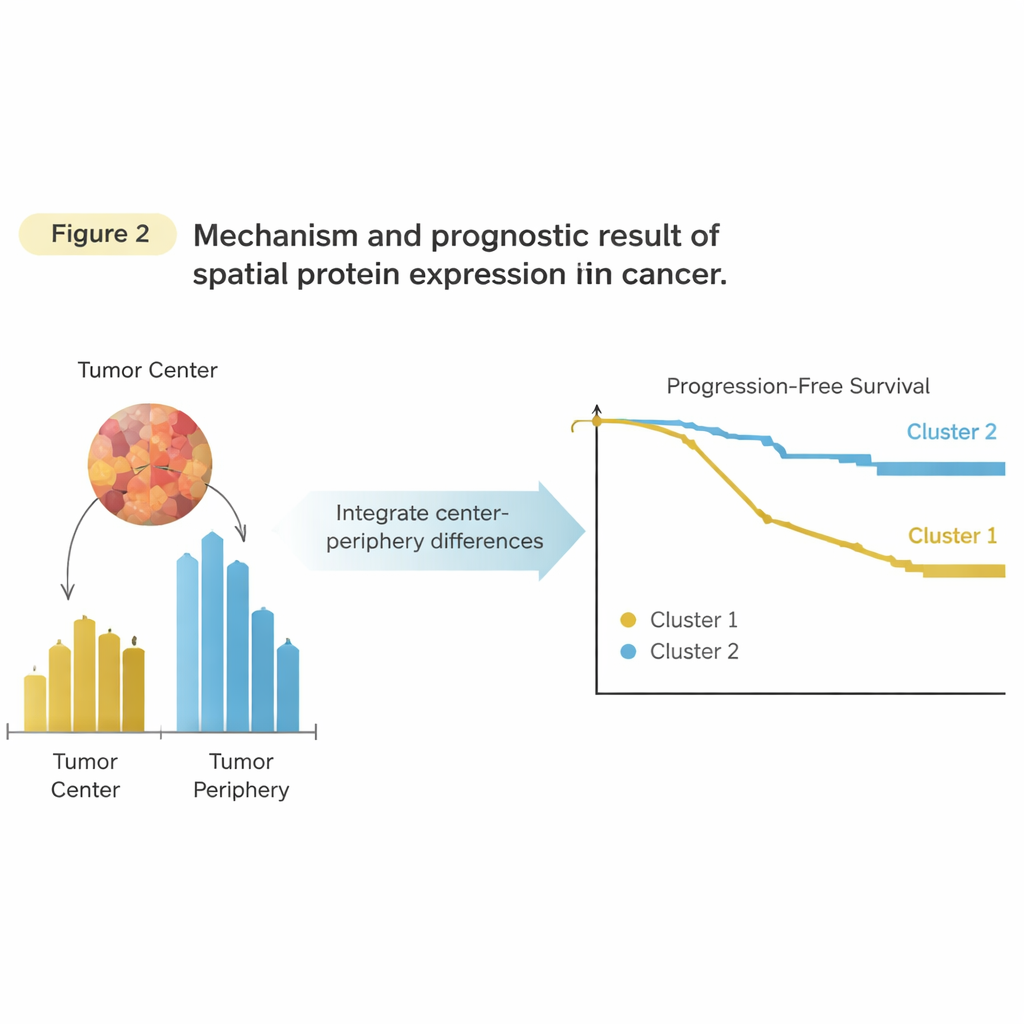
Combining both neighborhoods reveals a signal
The key step was to combine information from the center and the periphery rather than treating them separately. For each protein and each patient, the researchers calculated how much more (or less) that protein was expressed in the periphery compared with the center. This produced a single “balance value” per protein that still captured where the protein was higher inside the tumor. When they grouped patients based on these spatial balances, two new clusters emerged. This time, the clusters did matter: one group had a much shorter time before the cancer progressed than the other. Importantly, the power of this approach did not come from any single standout protein but from the overall spatial pattern across many proteins, especially those linked to immune cells and tissue remodeling at the tumor edge.
What this could mean for patients
For people facing prostate cancer, these findings suggest that where proteins are expressed inside the tumor—not just how much in total—may carry valuable clues about future disease behavior. While the study is small and uses a specialized, costly technology, it provides proof of concept that spatially aware protein measurements can uncover prognostic information that standard tests miss. With further validation and simpler laboratory methods, this kind of spatial biomarker could one day help doctors better distinguish which high-risk prostate cancers are likely to progress quickly and which may follow a more manageable course, leading to more tailored treatment decisions.
Citation: Schneider, F., Böning, S.H., Antunes, B.C. et al. Prognostic impact of spatial niches in prostate cancer. Sci Rep 16, 2598 (2026). https://doi.org/10.1038/s41598-026-35720-1
Keywords: prostate cancer, spatial profiling, tumor microenvironment, prognostic biomarker, intratumoral heterogeneity