Clear Sky Science · en
Correlations of m6A methylation-related mRNAs with thyroid cancer
Why tiny changes in RNA matter for thyroid cancer
Thyroid cancer is often curable, yet some patients still face recurring or spreading disease. This study looks at an emerging layer of biology called RNA “marking,” where small chemical tags are added to RNA molecules inside our cells. The researchers asked a focused question: in people with papillary thyroid cancer who do not have autoimmune thyroid disease, do these RNA tags change in a way that might help drive the cancer, and could those changes point to new ways to diagnose or treat it?
Zooming in on thyroid tumors without immune disease
Many people with thyroid cancer also have autoimmune thyroid disease, an immune attack on the thyroid that can muddy the biological picture. To avoid this, the team studied 26 patients whose papillary thyroid cancers arose without signs of autoimmunity. From each patient, they collected a piece of the tumor and a nearby piece of noncancerous thyroid tissue. In a subset of three patients, they performed broad surveys of which genes were switched on or off, and how heavily those RNAs carried a specific chemical tag called m6A, the most common internal modification on RNA in human cells.
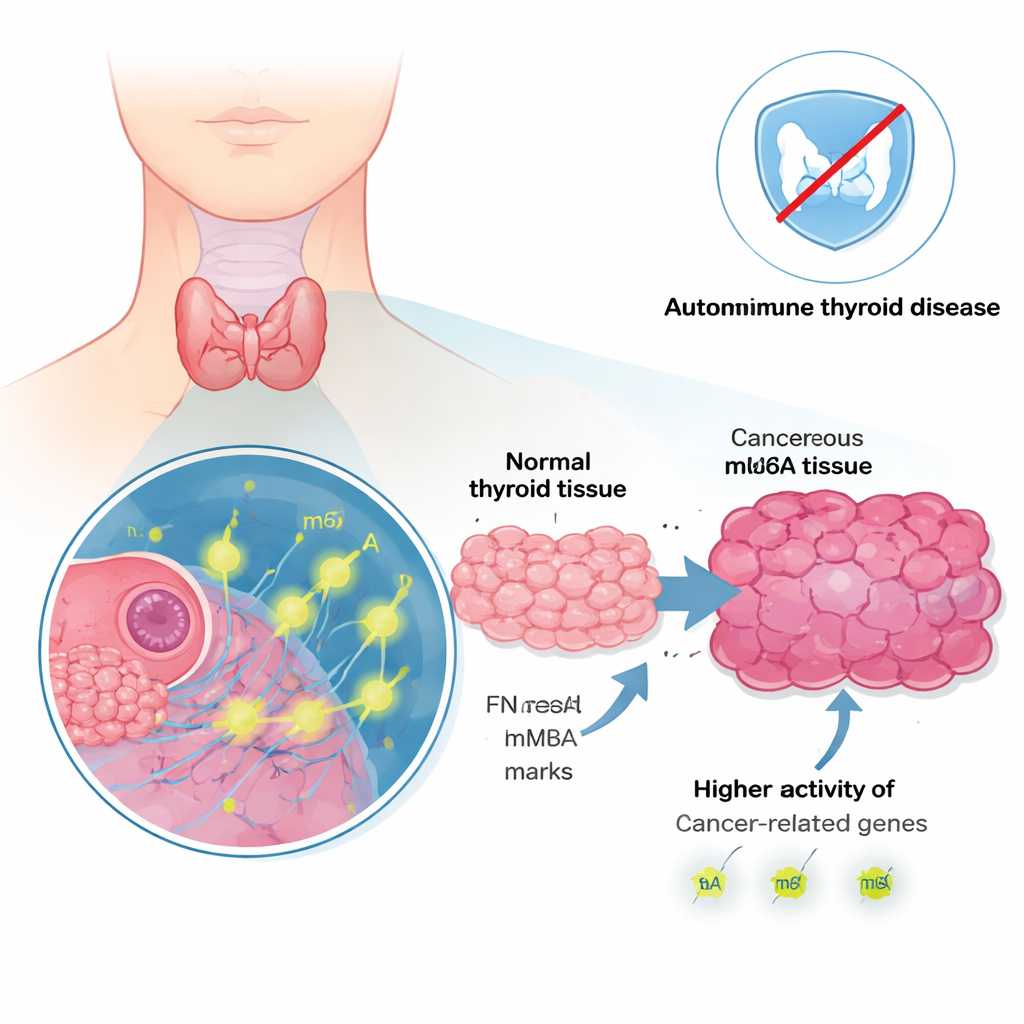
A surge of cancer-linked genes and pathways
When the researchers compared tumor tissue with nearby healthy tissue, they found 486 genes that were more active and 39 that were less active in the cancer. Many of the most strongly increased genes have already been implicated in cancer growth, spread, or treatment resistance. These include genes such as LAMB3, FN1, and NMU, which help cells interact with their surrounding support structure and with neighboring cells. Computer analysis showed that the boosted genes clustered in pathways already known to fuel cancer, including signaling networks related to cell division, stress responses, and communication between cells and the immune system. In other words, the tumors had clearly reprogrammed their gene activity toward growth and invasion.
RNA tags and a key “reader” are turned up
The next question was whether the m6A tag on RNA was altered in these cancers. Using a biochemical assay, the team found that overall m6A levels were higher in tumor tissue than in the matched normal tissue. They then examined the genes that control this tagging system: “writers” that add m6A, “erasers” that remove it, and “readers” that bind to tagged RNA and influence what happens next. Only one stood out in the tumors: a reader protein called IGF2BP2 was produced at significantly higher levels. Earlier work has shown that IGF2BP2 can latch onto m6A-marked RNAs and make them more stable and more likely to be translated into protein. In papillary thyroid cancer, IGF2BP2 has already been linked to more aggressive behavior, poorer outcomes, and greater ability to spread to lymph nodes.
Which RNAs carry extra tags—and why that matters
By combining m6A mapping with gene-activity measurements in the same samples, the researchers identified 367 RNAs that were more heavily tagged in tumors and 12 that were less tagged. Strikingly, most of the top hyper-tagged RNAs belonged to the same list of cancer-boosting genes that were more active overall. For 147 genes, tumors showed both higher m6A levels and higher expression, including FN1, LAMB3, NMU, and CDKN2B. Network analysis suggested these genes sit at the heart of systems that control how tumor cells stick to their surroundings, respond to inflammatory signals, and remodel the tissue around them—key steps for invasion and metastasis. The team also found dozens of tagged long noncoding RNAs, including ones linked to known cancer pathways, hinting that RNA marking may shape not only protein-coding messages but also regulatory RNAs that fine-tune gene behavior.
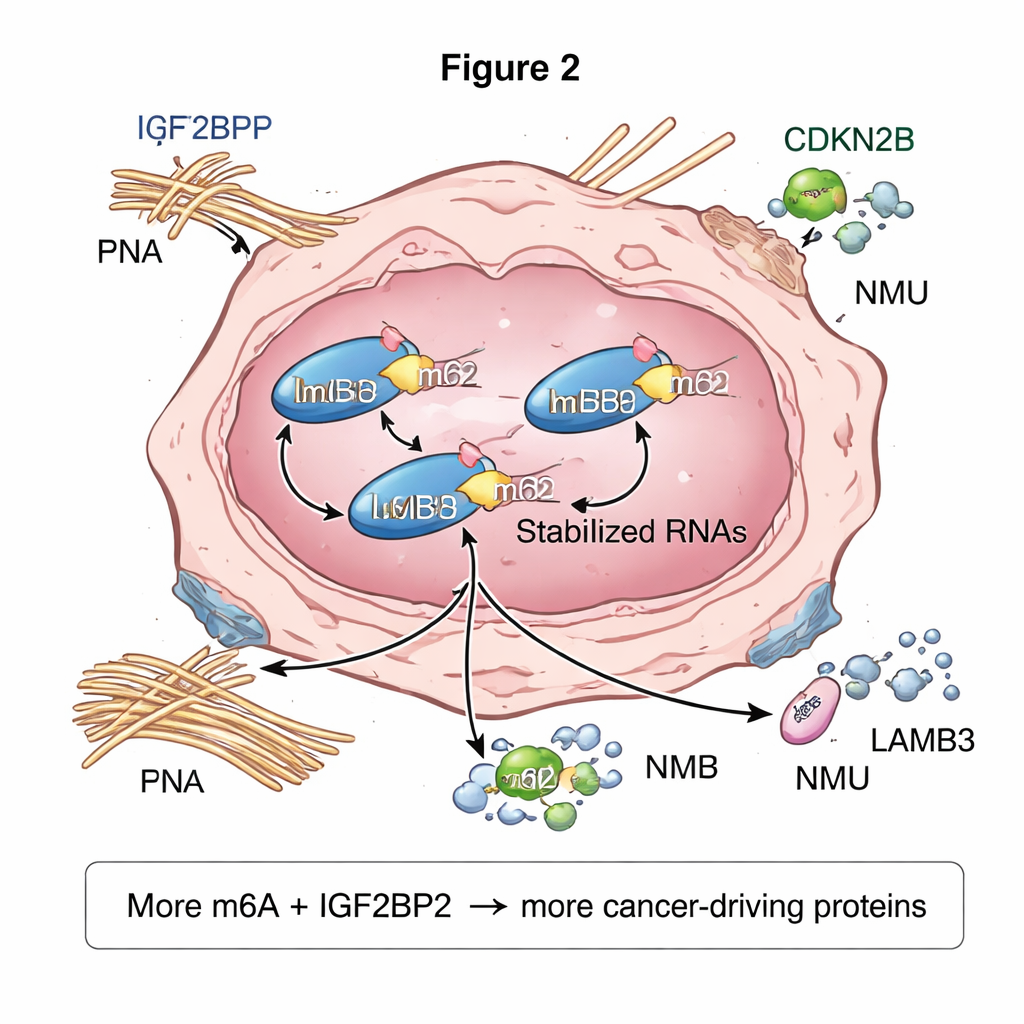
What this could mean for patients
For non-specialists, the core message is that papillary thyroid cancers—even in patients without autoimmune thyroid disease—appear to exploit a chemical marking system on RNA to amplify cancer-driving networks. Tumor cells show higher global m6A levels and increased amounts of the reader protein IGF2BP2, which together stabilize and boost the production of key genes involved in growth and spread. While this work is mainly based on advanced computational and laboratory analyses in a small group of patients, it highlights specific players—such as IGF2BP2 and the m6A-marked genes FN1, LAMB3, and NMU—as promising candidates for new tests or targeted therapies. Future studies will need to confirm these findings in larger groups and in experimental models, but the results suggest that “epigenetic” marks on RNA are an important, and potentially druggable, layer of thyroid cancer biology.
Citation: Jiang, Z., Luo, S., Lin, Y. et al. Correlations of m6A methylation-related mRNAs with thyroid cancer. Sci Rep 16, 5688 (2026). https://doi.org/10.1038/s41598-026-35712-1
Keywords: papillary thyroid cancer, RNA methylation, m6A modification, IGF2BP2, cancer biomarkers