Clear Sky Science · en
Rational redesign of high-activity G-quadruplex DNAzyme through flanking and looping of nucleobases
Small DNA Machines With Big Potential
Imagine replacing fragile protein enzymes with tiny strands of DNA that can survive heat, chemicals, and rough handling, yet still carry out useful chemistry. This study explores exactly that idea. The researchers tweak special DNA structures so they behave like miniature cleaning enzymes, able to use hydrogen peroxide to generate a strong signal. These tougher, faster DNA “machines” could make future medical tests, environmental sensors, and portable diagnostics cheaper, more reliable, and easier to use outside of the lab.
Turning DNA Into a Tiny Chemical Tool
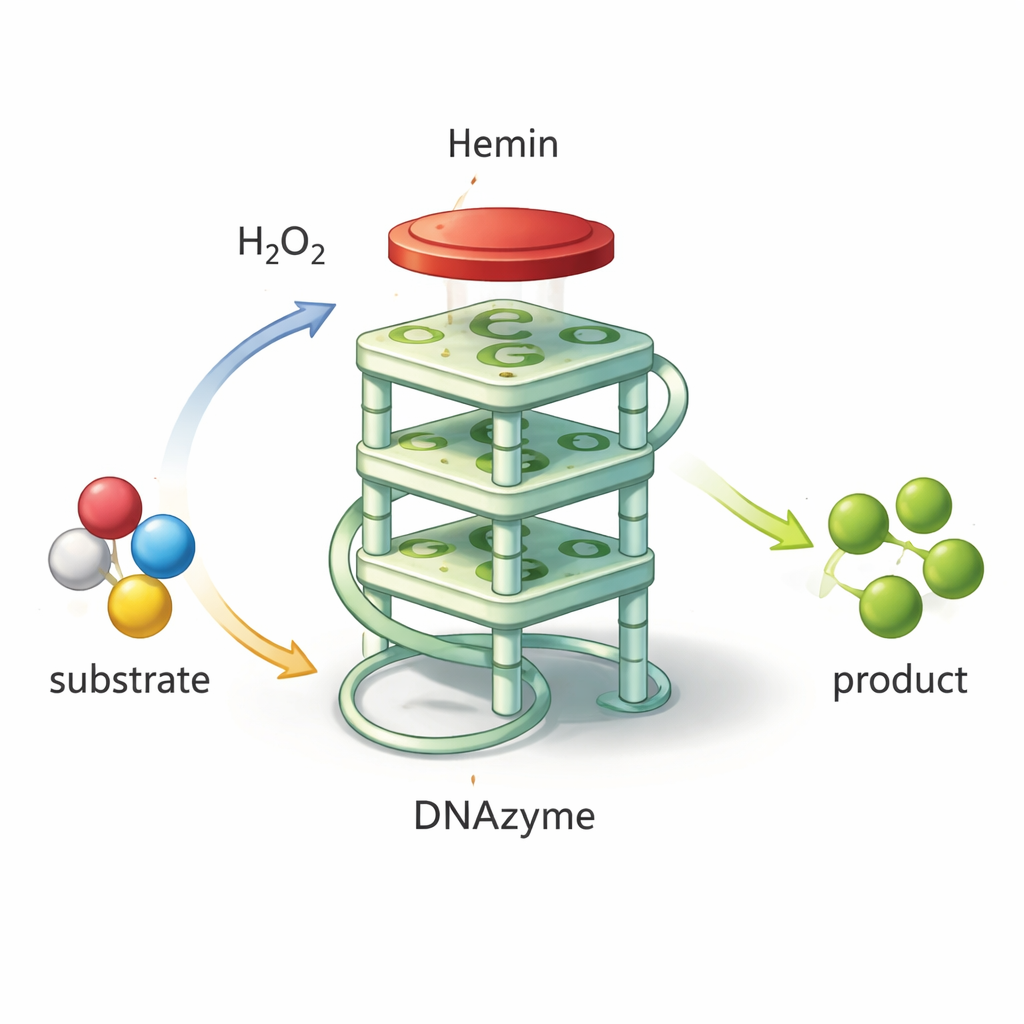
Not all DNA is just a passive carrier of genetic information. Certain short sequences can fold into unusual shapes that grab specific molecules or even speed up chemical reactions. One such shape is the G‑quadruplex, in which guanine-rich DNA folds into a compact, four‑layered stack. When a small iron-containing molecule called hemin sits on top of this stack, the pair acts like a “DNAzyme”: a DNA-based catalyst that mimics natural peroxidase enzymes. It can use hydrogen peroxide to oxidize a color‑forming chemical, producing a strong green signal that is easy to measure. Because these DNAzymes are cheap to make, highly stable, and easy to redesign, they are promising building blocks for biosensors that detect pathogens, toxins, or disease markers.
Why Current DNA Enzymes Need an Upgrade
Despite their promise, most DNAzymes are still slower and less efficient than protein enzymes found in nature. Existing biosensors often must amplify the target using techniques like PCR or add extra helper chemicals, which raises cost and complexity. Previous attempts to improve DNAzymes have included linking two DNA units together, attaching hemin permanently, or surrounding the reactive site with extra chemical groups. These tricks sometimes help, but they can also introduce bulk that gets in the way or demand elaborate chemistry. A key open question has been how simple changes to the nearby DNA bases—especially those that do not break the core G‑quadruplex shape—might tune activity in a predictable, “designable” way.
Redesigning a High‑Performance DNA Enzyme
The team focused on a particularly active DNAzyme known as B730, which already ranks among the best unmodified G‑quadruplex catalysts. They systematically altered the DNA just outside its core by adding or repositioning common bases such as adenine, thymine, and cytosine in the loop and tail regions. One redesigned version, called B730‑1.2, combined added adenines in the loops with a short thymine–cytosine pair at one end of the strand. Under moderate hydrogen peroxide conditions, this variant tripled the initial reaction speed and roughly quadrupled the total amount of colored product compared with the original B730. It also clearly outperformed two other well‑known DNAzymes, AS1411 and CatG4, when tested side by side.
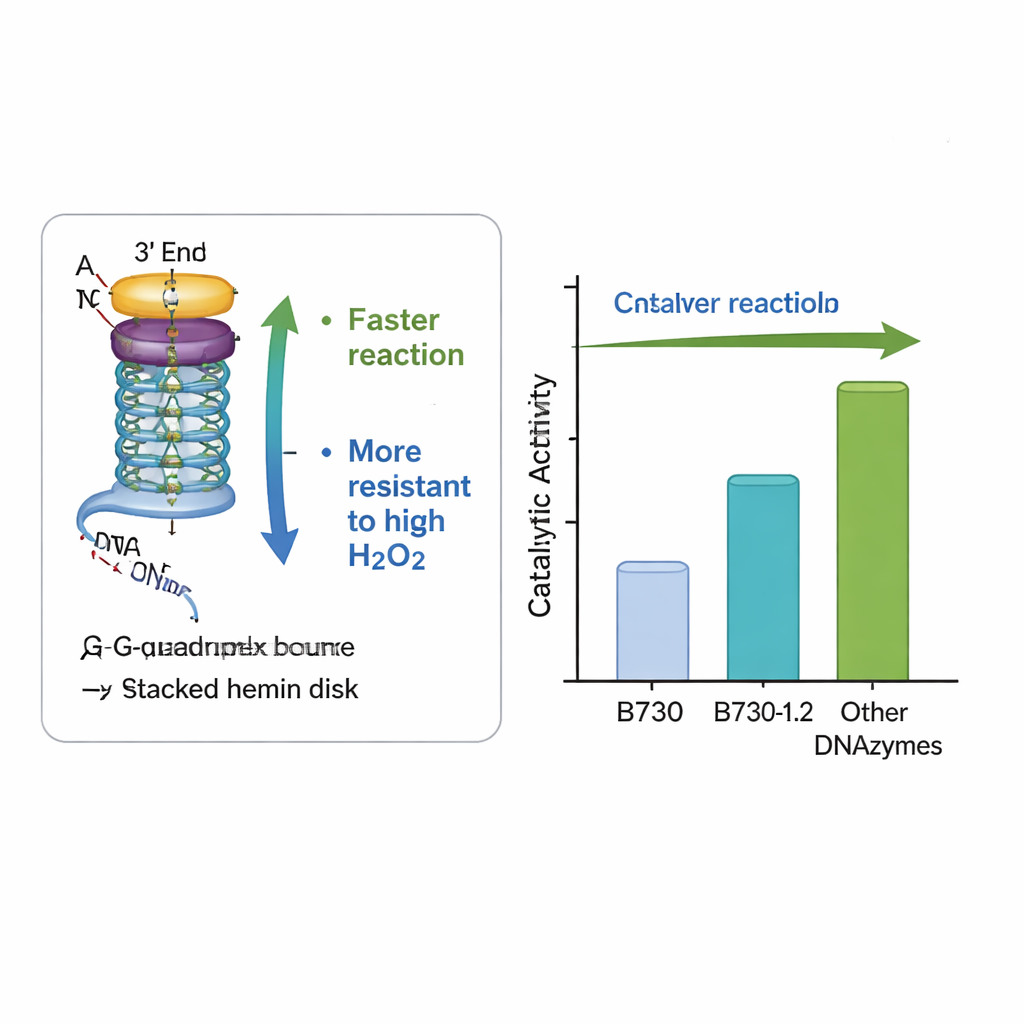
Built to Withstand Harsh Conditions
An important practical hurdle for both natural and artificial peroxidases is that high levels of hydrogen peroxide, the very ingredient that drives the reaction, can destroy the enzyme and shut the process down. The redesigned B730‑1.2 DNAzyme showed remarkable resilience: it maintained and even increased its activity at peroxide levels that typically inactivate similar systems. Measurements of light absorption confirmed that the modified DNA helped form the key reactive intermediate—the so‑called Compound I—more quickly, without disturbing the overall G‑quadruplex shape. In other words, subtle changes in the surrounding bases created a more favorable local environment for chemistry, speeding up the useful steps while helping to protect the catalytic center from self‑destruction.
What This Means for Future Sensors
To a non‑specialist, the message is straightforward: by carefully adjusting just a few “letters” on either side of an already good DNA enzyme, the authors produced a version that works faster and keeps going under harsher conditions. Their strategy of tweaking flanking and loop bases offers a simple, low‑cost recipe for building more powerful DNA‑based catalysts without resorting to complex chemical modifications. Such rugged, efficient DNAzymes could sit at the heart of next‑generation test strips and portable devices that quickly turn invisible biological signals—like traces of virus or pollutants—into easy‑to‑read color changes.
Citation: Adeoye, R.I., Babbudas, N., Birchenough, M. et al. Rational redesign of high-activity G-quadruplex DNAzyme through flanking and looping of nucleobases. Sci Rep 16, 5060 (2026). https://doi.org/10.1038/s41598-026-35686-0
Keywords: G-quadruplex DNAzyme, peroxidase mimic, biosensing, aptamer engineering, hydrogen peroxide catalysis