Clear Sky Science · en
In silico design of a multi-epitope vaccine targeting DENV-1 and DENV-3
Why dengue vaccines still matter
Dengue fever is no longer a rare tropical illness; it now threatens billions of people across more than 100 countries and regularly overwhelms hospitals in parts of Asia, Latin America, and beyond. Yet even with two licensed vaccines on the market, protection is uneven, especially for people who have never had dengue before and in regions where several viral types circulate at once. This study asks a timely question: can computer‑guided design help us build a safer, more precise vaccine, aimed specifically at dangerous co‑infections with two dengue types that often strike together?
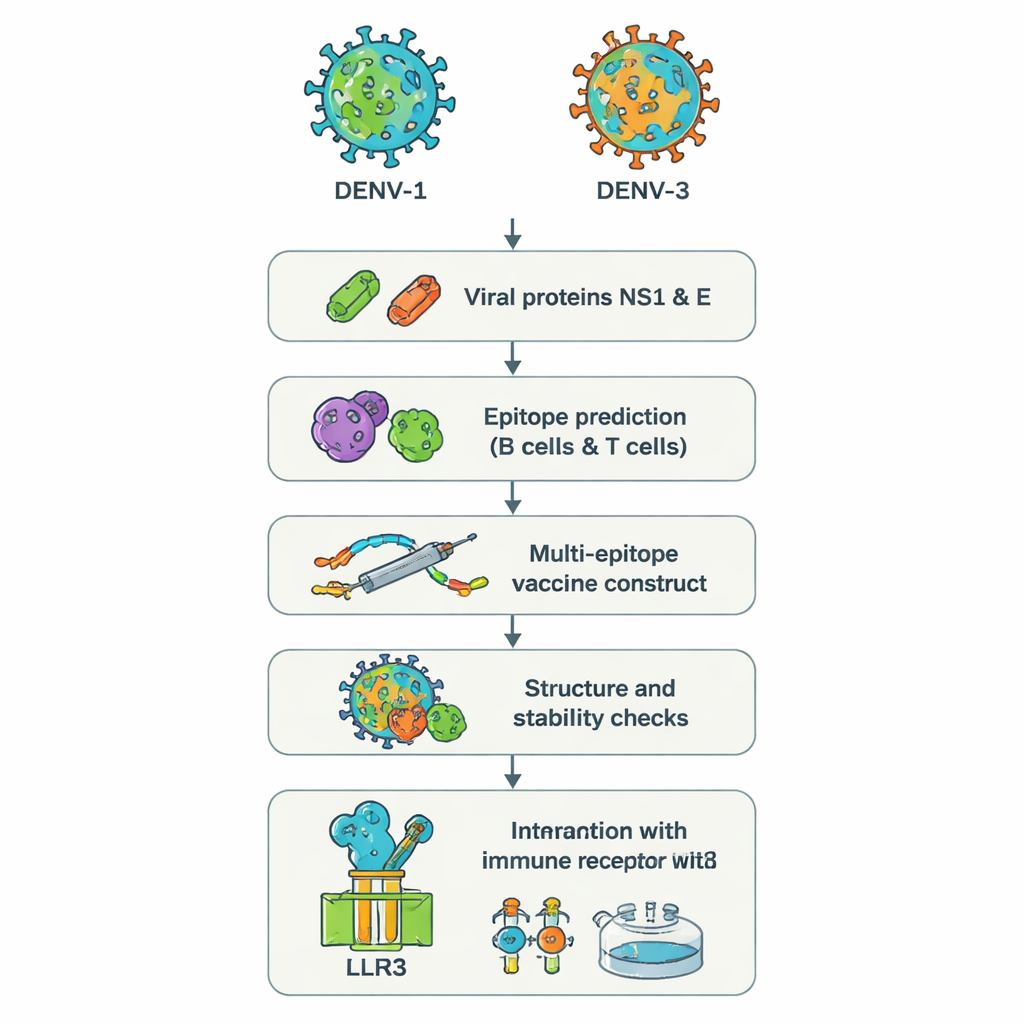
Two sneaky viral types and why they are a problem
Dengue virus comes in four versions, called serotypes DENV‑1 through DENV‑4. Infection with any one can cause high fever, severe pain, and in some cases life‑threatening bleeding or shock. Worryingly, getting sick a second time with a different serotype can sometimes make disease worse rather than better, because existing antibodies may help the new virus enter cells more easily, a process called antibody‑dependent enhancement. In recent outbreaks, doctors have reported patients infected at the same time with DENV‑1 and DENV‑3, a combination linked to more severe illness and confusing test results. Current vaccines do not reliably protect all age groups and all serotypes, especially in people with no prior exposure, leaving a dangerous gap in defenses.
Designing a vaccine from the molecule up
Instead of growing whole viruses in the lab, the researchers used an approach sometimes called “reverse vaccinology.” They started from the genetic sequences of two dengue proteins, NS1 and E, from DENV‑1 and DENV‑3. These proteins are key to how the virus invades cells and how the immune system sees it. Powerful web‑based tools scanned the protein sequences to locate short stretches—epitopes—that are most likely to be recognized by human B cells (which make antibodies) and T cells (which kill infected cells and coordinate responses). From hundreds of candidates, the team selected a small set that were predicted to be strongly visible to the immune system, shared between the two serotypes, and capable of triggering helpful antiviral signals such as interferon‑gamma.
Building a single multi‑purpose vaccine molecule
The chosen epitopes were then digitally stitched together into one long artificial protein, a “multi‑epitope” vaccine. Short amino‑acid linkers act as flexible spacers so each epitope keeps its shape and remains accessible to immune cells. An extra piece, based on a natural human antimicrobial peptide called beta‑defensin, was added as an adjuvant to boost the overall response. Computer analysis predicted that the final construct, containing 575 amino acids, would be stable, water‑loving (and therefore easier to dissolve), and unlikely to act as an allergen. Further structure‑prediction tools generated a three‑dimensional model and checked that most building blocks sat in realistic positions, similar to those seen in known protein structures.
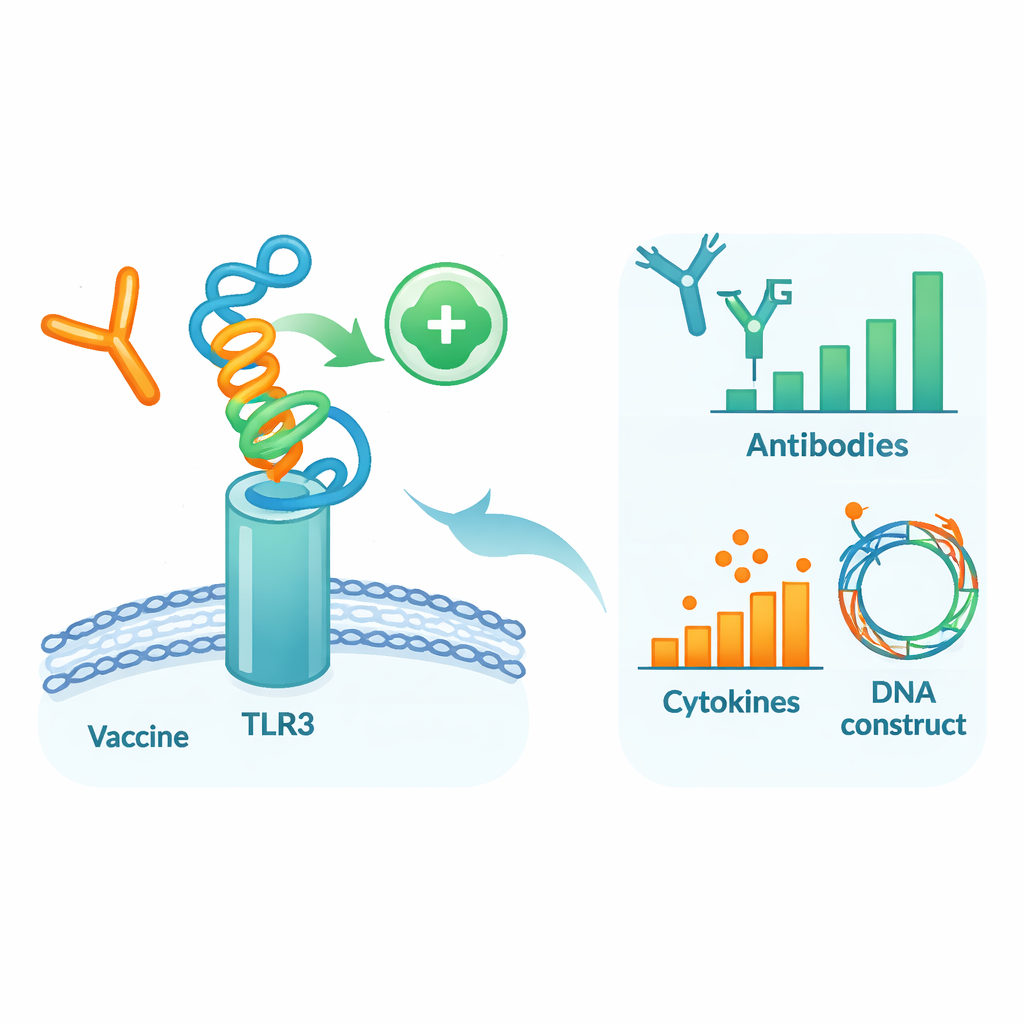
Testing the candidate inside a virtual body
To see how this designed protein might behave in a human‑like setting, the authors ran a series of detailed computer simulations. First, they docked the vaccine model onto TLR3, a sensor protein on immune cells that detects viral material and helps launch early defenses. Molecular dynamics simulations—virtual movies of atoms in motion—suggested that the vaccine and TLR3 form a stable complex, supported by favorable binding energies and numerous hydrogen bonds. Additional analyses of motion and energy pointed to specific regions of both molecules that act as contact “hotspots.” Next, an immune‑system simulator was used to mimic three vaccine doses over several months. The virtual immune system produced strong waves of protective IgG antibodies, long‑lived B and T cell memory, and signaling molecules consistent with a robust antiviral response.
From computer model to lab‑ready blueprint
Finally, the researchers adapted the genetic code of the vaccine for efficient production in common laboratory microbes and successfully placed this optimized DNA sequence into a standard expression plasmid, ready for future experimental testing. In simple terms, their work delivers a detailed blueprint for a new dengue vaccine that targets carefully chosen pieces of DENV‑1 and DENV‑3, is predicted to be stable and safe, and should strongly engage both arms of the immune system. While these results are purely computational and must be confirmed in cells, animals, and eventually people, they show how modern bioinformatics can rapidly generate tailored vaccine candidates for complex problems like dengue co‑infection.
Citation: Ishwar, D., Padavu, S., Kumar, M. et al. In silico design of a multi-epitope vaccine targeting DENV-1 and DENV-3. Sci Rep 16, 5308 (2026). https://doi.org/10.1038/s41598-026-35678-0
Keywords: dengue vaccine, multi-epitope, DENV-1, DENV-3, immunoinformatics