Clear Sky Science · en
Integrative analysis of proteomics and metabolomics reveals amino acid metabolism disorder in adriamycin-resistant acute myeloid leukemia cells
Why some leukemia drugs stop working
Chemotherapy has transformed the treatment of acute myeloid leukemia (AML), a fast‑growing blood cancer. Yet many patients see their disease return because the cancer cells learn to survive the very drugs designed to kill them. This study asks a simple but crucial question: what changes inside leukemia cells when they become resistant to adriamycin, one of the standard chemotherapy drugs, and could those changes point to new ways to restore the drug’s power?
Peeking inside leukemia cells
To explore this, the researchers compared a common human AML cell line, called HL60, with a sister line that had been made resistant to adriamycin (HL60/R). Rather than looking at just one gene or protein at a time, they used two broad, complementary approaches. Proteomics measured thousands of different proteins—the workhorse molecules that carry out most tasks in a cell. Metabolomics measured hundreds of small molecules that make up the cell’s metabolism, including fats, sugars, and amino acids. By combining these “omics” layers, the team built a detailed picture of how resistant cells differ from their still‑sensitive cousins.
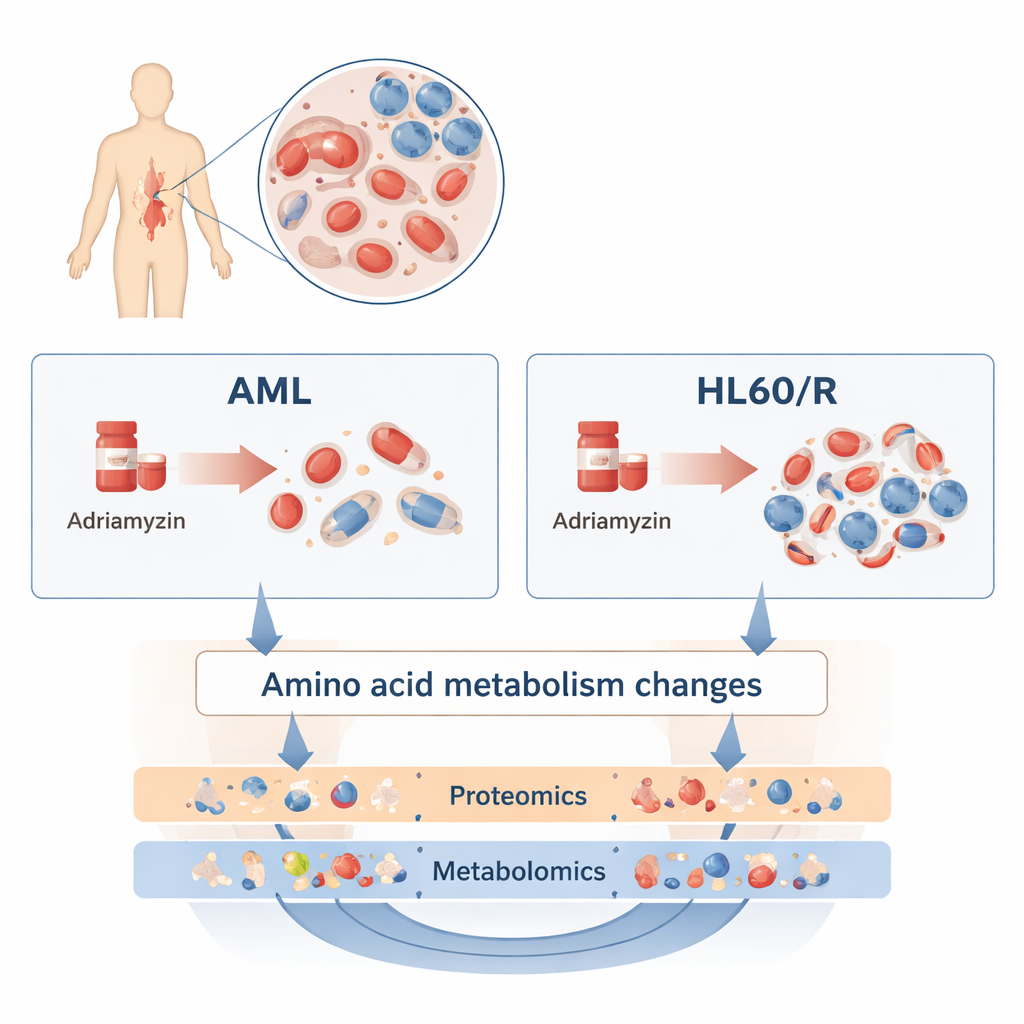
Massive rewiring of cell machinery
The proteomics survey revealed sweeping changes: more than 3,200 proteins were present at higher or lower levels in adriamycin‑resistant cells compared with normal HL60 cells. Many of these proteins clustered in pathways that control how cells use energy and respond to stress. In particular, proteins in the cAMP signaling pathway, the HIF‑1 pathway (which helps cells cope with low oxygen), and oxidative phosphorylation (a major energy‑producing process in mitochondria) were altered. These shifts suggest resistant leukemia cells reprogram how they breathe, grow, and signal to one another in order to withstand chemotherapy.
Metabolism shifts toward amino acids
The metabolomics analysis told a complementary story. Out of about 1,400 detected metabolites, 260 changed significantly in the resistant cells. Statistical models cleanly separated resistant from non‑resistant cells, indicating a consistent metabolic signature of resistance. When the altered metabolites were mapped onto known biochemical routes, several pathways stood out. Many of the most strongly affected routes were tied to amino acids, the building blocks of proteins—especially those involving alanine, aspartate, glutamate, cysteine, methionine, and glutathione. Other networks, such as purine and pyrimidine metabolism (important for DNA and RNA) and the handling of certain fats, were also disturbed, underscoring that resistance is linked to broad metabolic remodeling.
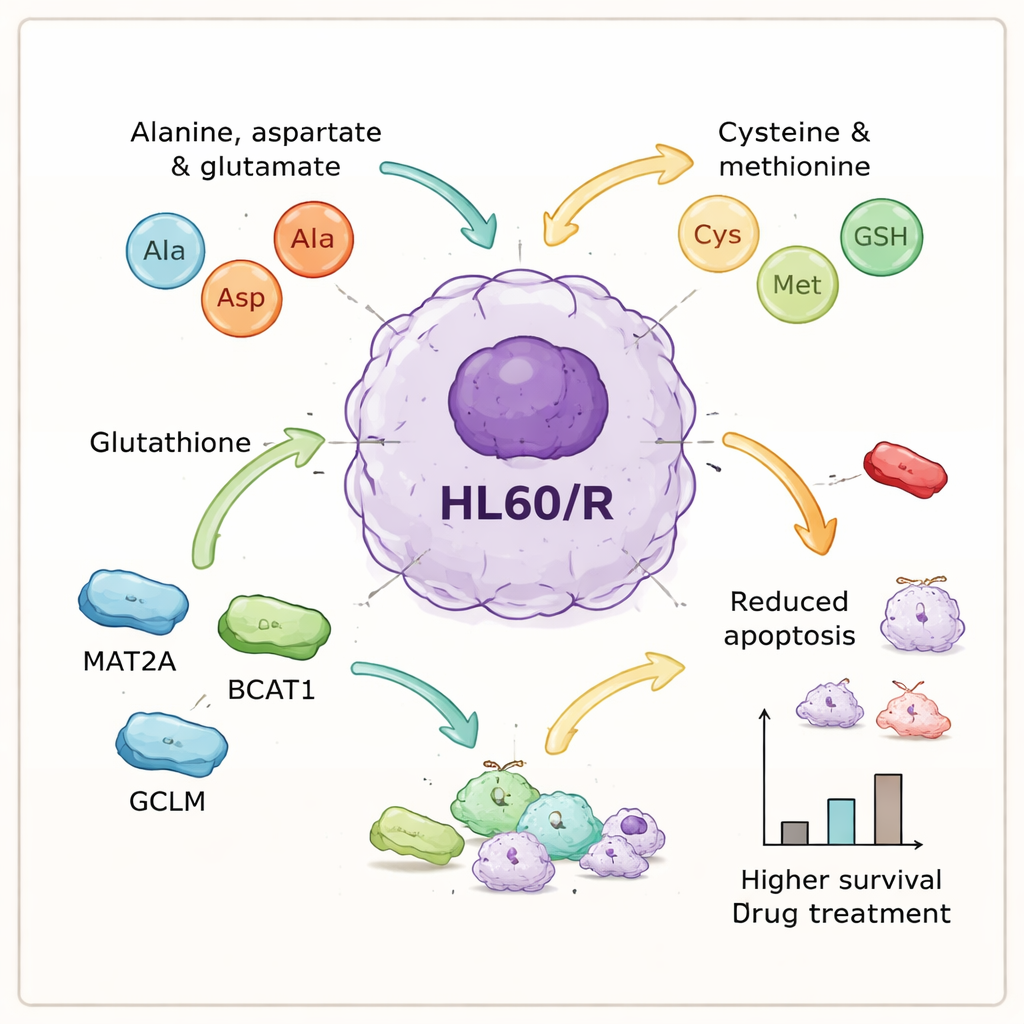
Key molecular players that help cancer cells endure
By integrating the protein and metabolite data, the authors homed in on a set of amino acid‑related processes that seemed central to resistance. They selected six proteins tied to these pathways—GOT1, GPX1, AHCY, MAT2A, BCAT1, and GCLM—for closer inspection. Laboratory tests confirmed that five of them were more abundant in resistant cells, consistent with the idea that amino acid processing is turned up, while one antioxidant enzyme, GPX1, was reduced. To probe cause and effect, the team used small interfering RNAs to dial down three of the up‑regulated proteins—MAT2A, BCAT1, and GCLM—in the resistant cells. When these proteins were knocked down, the cells underwent significantly more programmed cell death after treatment, meaning they lost much of their hard‑won protection against adriamycin.
What this means for future treatments
Taken together, these findings suggest that adriamycin‑resistant AML cells survive not just because of single mutations, but because they reshape their internal chemistry, with amino acid metabolism as a central hub. By diverting more resources into particular amino acid and glutathione pathways, the cells appear better able to manage stress, repair damage, and avoid death when exposed to chemotherapy. For non‑specialists, the key message is that resistance is not random: it follows identifiable patterns that can be measured and, potentially, targeted. In the long run, drugs that interfere with specific amino acid‑processing proteins such as MAT2A, BCAT1, or GCLM could be paired with adriamycin or similar agents, giving clinicians new tools to prevent or overcome resistance in patients with acute myeloid leukemia.
Citation: Li, C., Liang, X., Gong, S. et al. Integrative analysis of proteomics and metabolomics reveals amino acid metabolism disorder in adriamycin-resistant acute myeloid leukemia cells. Sci Rep 16, 4902 (2026). https://doi.org/10.1038/s41598-026-35675-3
Keywords: acute myeloid leukemia, drug resistance, adriamycin, amino acid metabolism, multi-omics