Clear Sky Science · en
A proteomics and redox proteomics approach to understanding ARDS heterogeneity
Why this matters for people in intensive care
Acute Respiratory Distress Syndrome (ARDS) is a life-threatening form of lung failure that can strike people with severe infections, trauma, or other critical illnesses. Many patients with ARDS look similar at the bedside, yet some recover while others do not, and no drug has been proven to specifically reverse the syndrome. This study asks a simple but important question: if we look closely at the proteins and chemical reactions inside the lungs and blood of ARDS patients, can we uncover hidden biological "types" of the disease that explain why patients fare so differently?
Looking inside the lungs, not just the bloodstream
To explore this, researchers followed 16 adults in an intensive care unit who had ARDS and were on breathing machines. Within the first three days after diagnosis, they collected blood and a special sample called bronchoalveolar lavage (BAL) fluid, which is a gentle wash of the air sacs in the lungs. They repeated these collections over several days when possible. Using high-end mass spectrometry, they measured hundreds of proteins in each sample (proteomics) and, crucially, also examined how oxidized some of these proteins were (redox proteomics), which reflects how much they have been damaged or regulated by reactive oxygen species, the chemically aggressive by-products of inflammation.
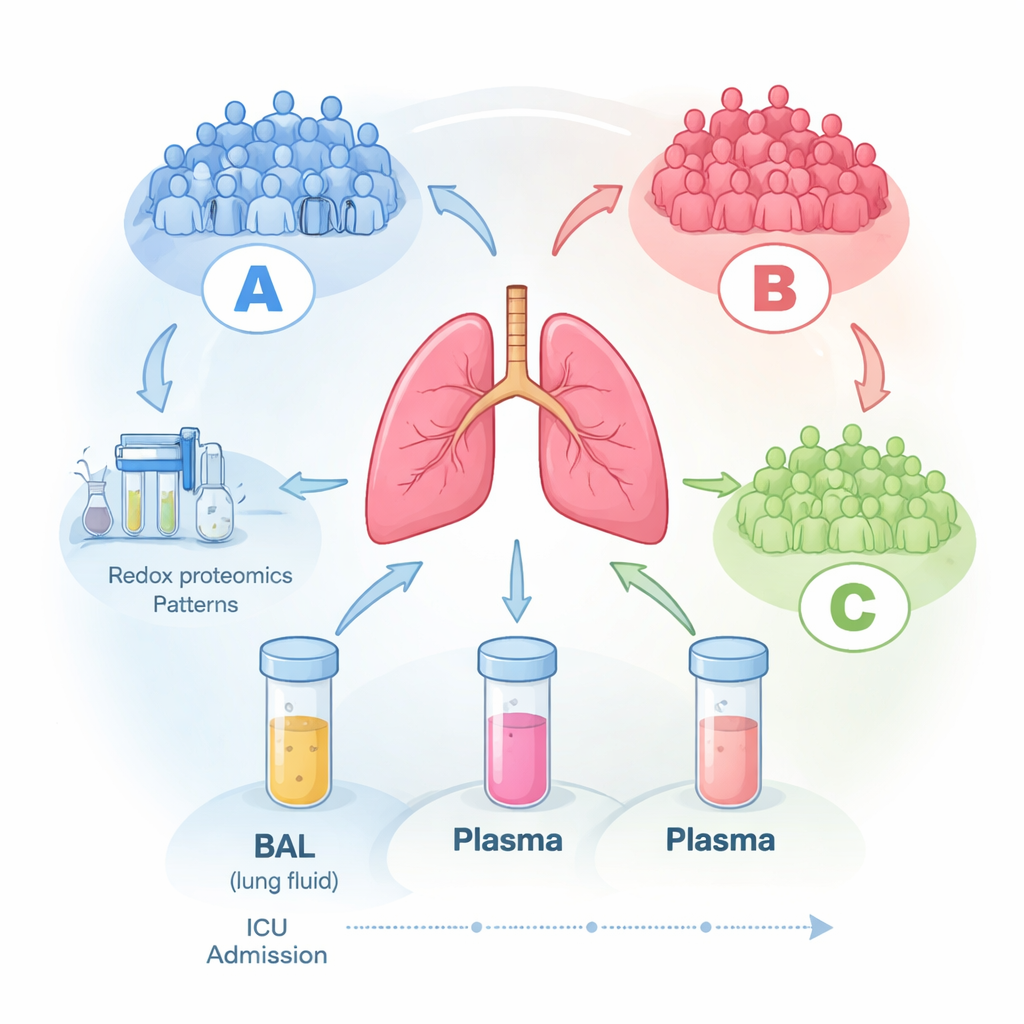
Three hidden patient groups emerge
When the team let the data group itself without telling the computer anything about the patients, a clear pattern appeared in the lung fluid samples: three distinct molecular groups, labeled A, B, and C. Group A patients tended to be more severely ill at enrollment, while Groups B and C had milder disease. Strikingly, these molecular signatures stayed largely stable for up to six days in the ICU, suggesting that each patient falls into a relatively consistent biological pattern rather than randomly fluctuating from day to day. Standard bedside measures—such as oxygen levels, length of stay, or general illness scores—did not track well with these molecular groupings, hinting that the usual clinical tools miss key biology happening inside the lungs.
Oxidative stress and the body’s defenses
A major theme running through the data was the balance between damaging reactive oxygen species and the body’s antioxidant defenses. In Group A, the sickest group, many lung proteins involved in energy production and cellular housekeeping were present at lower levels. Even more telling, important antioxidant enzymes such as peroxiredoxins, glutathione-related proteins, thioredoxin, and catalase were significantly reduced. In contrast, Groups B and especially C had higher levels of these protective proteins, suggesting that their lungs were better equipped to detoxify harmful oxidants and limit collateral damage from inflammation. Patterns in the oxidation state of proteins added another layer, showing different subsets of oxidized proteins enriched in the most and least severe groups.
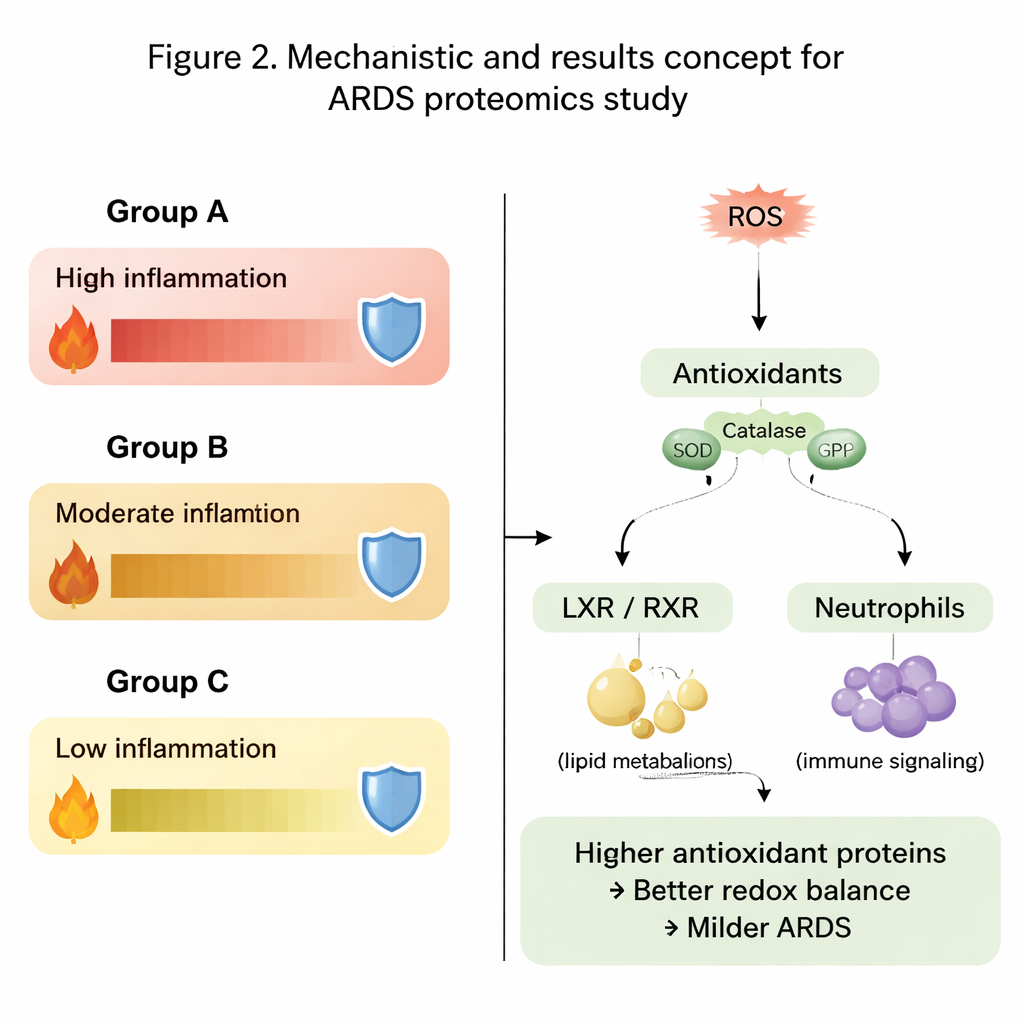
Signaling pathways that shape inflammation
Beyond single proteins, the authors examined entire biological pathways. They found that pathways tied to inflammation, cholesterol and fat handling, and immune cell activity behaved differently across the three groups. Signals linked to the liver X receptor–retinoid X receptor (LXR/RXR) pathway and an enzyme called DHCR24, which together help regulate both lipid metabolism and immune responses, were especially notable. In the most severe group, these pathways were predicted to be more active, paired with reduced signaling of the immune messenger interleukin-12. At the same time, pathways involving reactive oxygen species, their detoxification, and neutrophil (white blood cell) activity showed contrasting patterns between lung fluid and blood, underscoring that what happens locally in the lung may differ from what is seen in the circulation.
What this could mean for future care
This small, exploratory study does not change treatment today, and its findings need to be confirmed in much larger groups of patients. However, it demonstrates that by closely profiling proteins and their redox state directly in lung fluid—rather than relying only on blood tests or bedside scores—we can distinguish biologically meaningful subtypes of ARDS. These subtypes differ in oxidative stress, antioxidant capacity, metabolism, and immune signaling, and they remain relatively stable over the first critical days of illness. In the future, such molecular fingerprints could help doctors sort ARDS patients into more precise groups, choose targeted therapies tailored to each pattern, and design smarter clinical trials with a better chance of finding treatments that truly work.
Citation: Forshaw, T.E., Shukla, K., Wu, H. et al. A proteomics and redox proteomics approach to understanding ARDS heterogeneity. Sci Rep 16, 6034 (2026). https://doi.org/10.1038/s41598-026-35606-2
Keywords: acute respiratory distress syndrome, proteomics, oxidative stress, bronchoalveolar lavage, critical care