Clear Sky Science · en
Immunoassays for the detection and differentiation of Paenibacillus larvae, the etiological agent of American foulbrood (AFB) in honey bees
Why bee brood disease matters to all of us
Honey bees do far more than make honey. By pollinating crops and wild plants, they help secure our food supply and keep ecosystems running smoothly. One of the most destructive threats to bee colonies is a bacterial disease of the young, called American foulbrood. Once it takes hold in a hive, it can wipe out an entire colony and spread quickly to neighbors. This study describes new rapid tests that make it easier to spot the disease early and to understand which form of the bacterium is involved, giving beekeepers and veterinarians a better chance to stop outbreaks before they spiral.
A deadly childhood disease of bees
American foulbrood targets honey bee larvae, the developing brood that will grow into adult workers and queens. The culprit is a spore-forming bacterium, Paenibacillus larvae. Its spores can survive for years in old combs and dried larval remains, and only a few spores swallowed by a young larva can trigger infection. As the bacteria multiply, the larva collapses into a sticky mass that later dries into a dark scale tightly glued to the cell. These scales are packed with millions of spores and act as long-lasting reservoirs of infection that foragers and beekeepers can unwittingly spread between colonies and apiaries.
Two flavors of the same killer
Not all P. larvae are equally dangerous in the same way. Around the world, two main genetic types, known as ERIC I and ERIC II, are responsible for nearly all current outbreaks. Both are lethal, but they use different tools to break through the larval gut and invade the body. All virulent strains secrete a powerful chitin-degrading enzyme called PlCBP49, which helps them chew through the protective lining of the gut. ERIC I strains also make classic toxins that directly damage gut cells, whereas ERIC II strains instead rely on a surface protein called SplA that helps them stick to and then destroy the gut barrier by a still-unclear route. Because ERIC I and ERIC II differ in how fast they kill larvae and how an outbreak progresses, knowing which one is present can influence control decisions.
Turning bacterial weapons into diagnostic targets
The authors realized that these virulence tools—PlCBP49 and SplA—could be exploited as highly specific markers. If a test could detect PlCBP49, it would reveal infection by any dangerous P. larvae strain. If it could also detect SplA, it would flag specifically the ERIC II type. To do this, the team produced purified versions of both proteins, then used them to raise sets of monoclonal antibodies: laboratory-made proteins that bind only to a single molecular target. They screened these antibodies carefully, using dot blots and western blots, against multiple strains of ERIC I and ERIC II and against more than 20 other bacterial species commonly found in honey and brood comb. One antibody to PlCBP49 and one to SplA proved especially selective, recognizing all the right P. larvae strains while ignoring look‑alike bacteria from the hive environment.
From laboratory plates to a hive-side strip test
With suitable antibodies in hand, the researchers built two lab-based sandwich ELISA kits and one strip-style lateral flow assay, similar in concept to an at-home pregnancy or COVID-19 test. In the ELISAs, one antibody captures the target protein from a homogenized larva, and a second labeled antibody reveals its presence as a color change in a plastic microplate. Tests on experimentally infected larvae showed that the PlCBP49 ELISA detected 89% of infected individuals with no false positives, while the SplA ELISA detected 94% of ERIC II-infected larvae and correctly distinguished ERIC II from ERIC I with 97% accuracy. Building on the same antibody pairs, the team designed a duplex lateral flow strip with two test lines: one for PlCBP49 (all P. larvae) and one for SplA (ERIC II only). When larval samples from both lab infections and real outbreaks were run across the strips, the test correctly identified P. larvae in 95–99% of infected larvae and showed 96–100% specificity, with about 9 out of 10 genotype calls (ERIC I versus ERIC II) classified correctly.
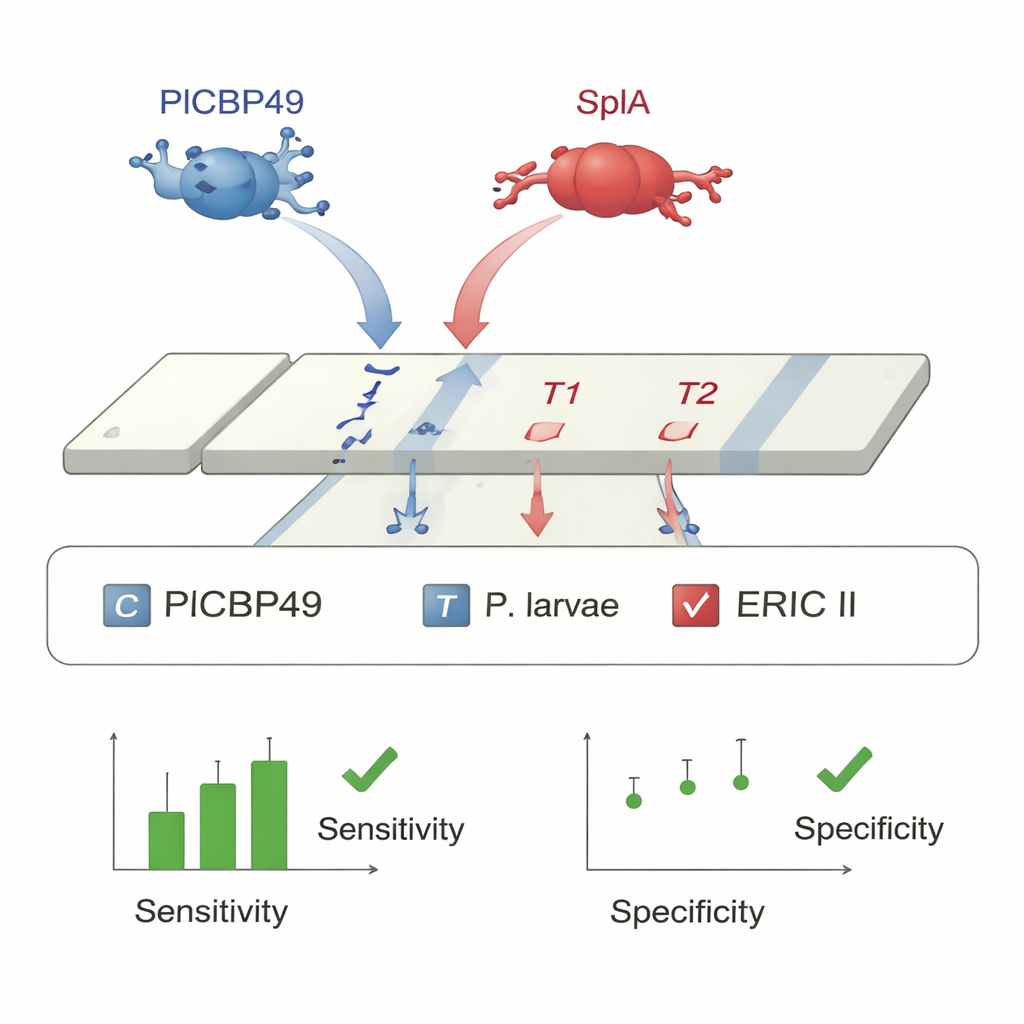
What this means for bees and beekeepers
Today, confirming American foulbrood often requires shipping suspicious combs or larvae to a specialized laboratory for culture or DNA testing, a process that can take days to weeks while the disease continues to spread. The new ELISA kits give laboratories a faster, automatable way to screen many samples, while the duplex lateral flow strip can be used directly at the hive as a point-of-care test. A beekeeper or inspector can sample a few suspect larvae, run the test in minutes, and learn not only whether P. larvae is present, but also whether the more rapidly acting ERIC II type is involved. This combination of speed, accuracy, and on-site usability has the potential to transform foulbrood control: earlier detection means earlier intervention, fewer colonies lost, and better protection of the pollination services on which agriculture and natural ecosystems depend.
Citation: Reinecke, A., Göbel, J. & Genersch, E. Immunoassays for the detection and differentiation of Paenibacillus larvae, the etiological agent of American foulbrood (AFB) in honey bees. Sci Rep 16, 2635 (2026). https://doi.org/10.1038/s41598-026-35590-7
Keywords: honey bee disease, American foulbrood, Paenibacillus larvae, rapid diagnostic test, lateral flow assay