Clear Sky Science · en
Comparative genomics of colistin-nonsusceptible multidrug-resistant Pseudomonas aeruginosa reveals emerging lineages in Thailand
Why this hospital germ matters to everyone
Pseudomonas aeruginosa is a hospital germ that preys on people when they are weakest—after surgery, on breathing machines, or with serious burns or lung disease. For years, doctors have relied on a powerful “last‑resort” antibiotic called colistin when other drugs fail. This study looks at strains of Pseudomonas from hospitals across Thailand that no longer respond to colistin or many other antibiotics. By reading the complete DNA of these bacteria, the researchers show how new, highly drug‑resistant lineages are spreading, and why that should concern patients, clinicians, and health systems worldwide.
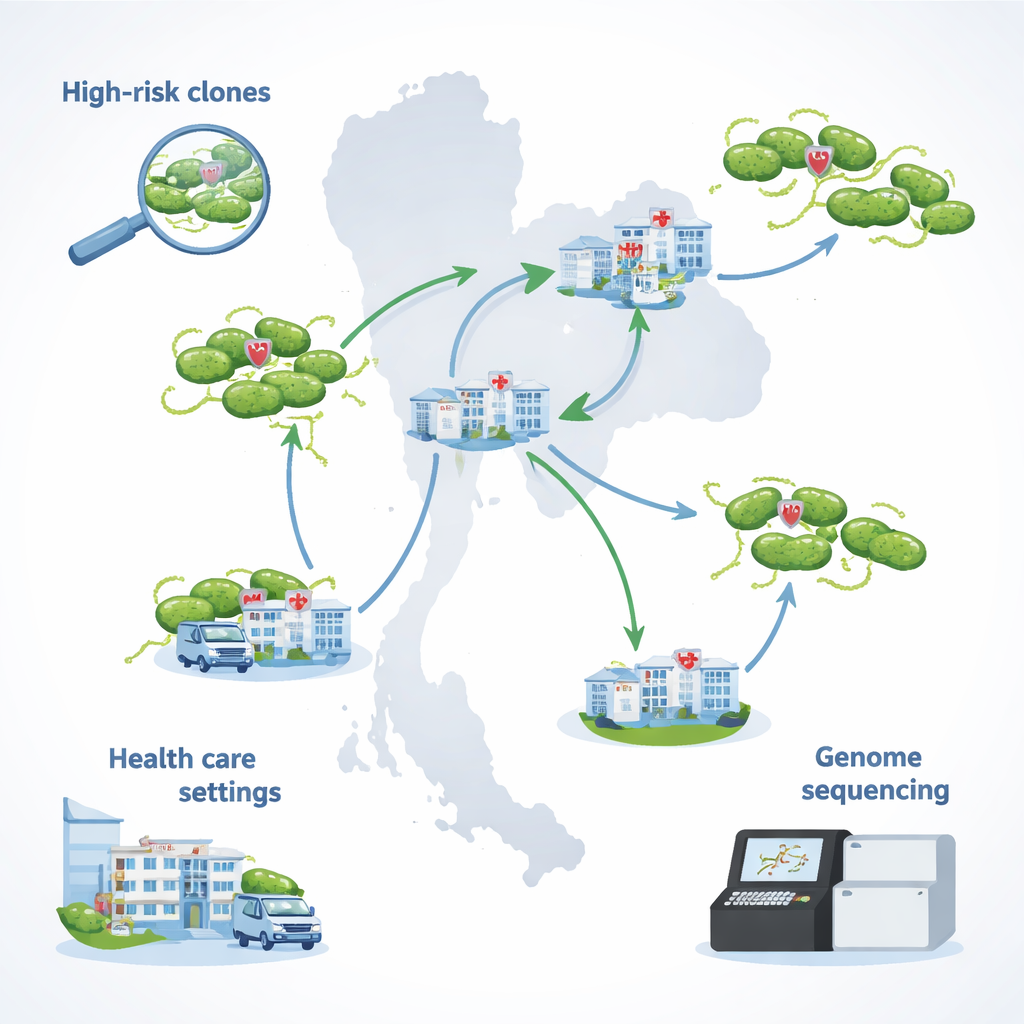
Tracking a hard‑to‑treat infection across Thailand
The team focused on 29 strains of Pseudomonas aeruginosa collected in 2021–2022 from hospitals that take part in Thailand’s national antibiotic‑resistance surveillance program. All of these strains were multidrug‑resistant: they could withstand several major classes of antibiotics, including drugs usually used for serious infections. Crucially, they were also not fully susceptible to colistin, the drug often kept in reserve for life‑threatening cases. Most samples came from urine, but others were taken from blood, sputum, pus, and surgical drain fluids—reflecting the many types of infections this germ can cause in hospitalized patients.
Reading the bacteria’s genetic “fingerprints”
Using a combination of short‑read and long‑read DNA sequencing, the researchers assembled high‑quality genomes for each strain. They then compared these genomes to classify the bacteria into genetic families, known as sequence types. Nine distinct sequence types were found, revealing substantial diversity. One, labeled ST5340, had never been described before. It turned out to be closely related to a known international high‑risk clone called ST357, differing at only one of seven standard housekeeping genes. Despite this close relationship, ST5340 stood out because every one of its isolates resisted all antibiotics tested, marking it as an especially worrisome lineage.
Emerging high‑risk lineages and their spread
By lining up tiny DNA differences called single‑nucleotide polymorphisms across 108 Thai Pseudomonas genomes (the 29 new ones plus 79 from public databases), the team built a family tree of strains circulating in the country. This analysis highlighted several dominant clusters centered on ST5340, ST357, ST654, and ST235—lineages already known, or now emerging, as “high‑risk” because they frequently resist multiple drugs and cause hospital outbreaks. ST5340 in particular appeared in multiple provinces and regions, hinting that it is spreading widely rather than being confined to a single hospital. Other high‑risk global clones, such as ST654 and ST235, were also present, while some globally important lineages like ST244 were absent, likely because the study included only colistin‑nonsusceptible strains.
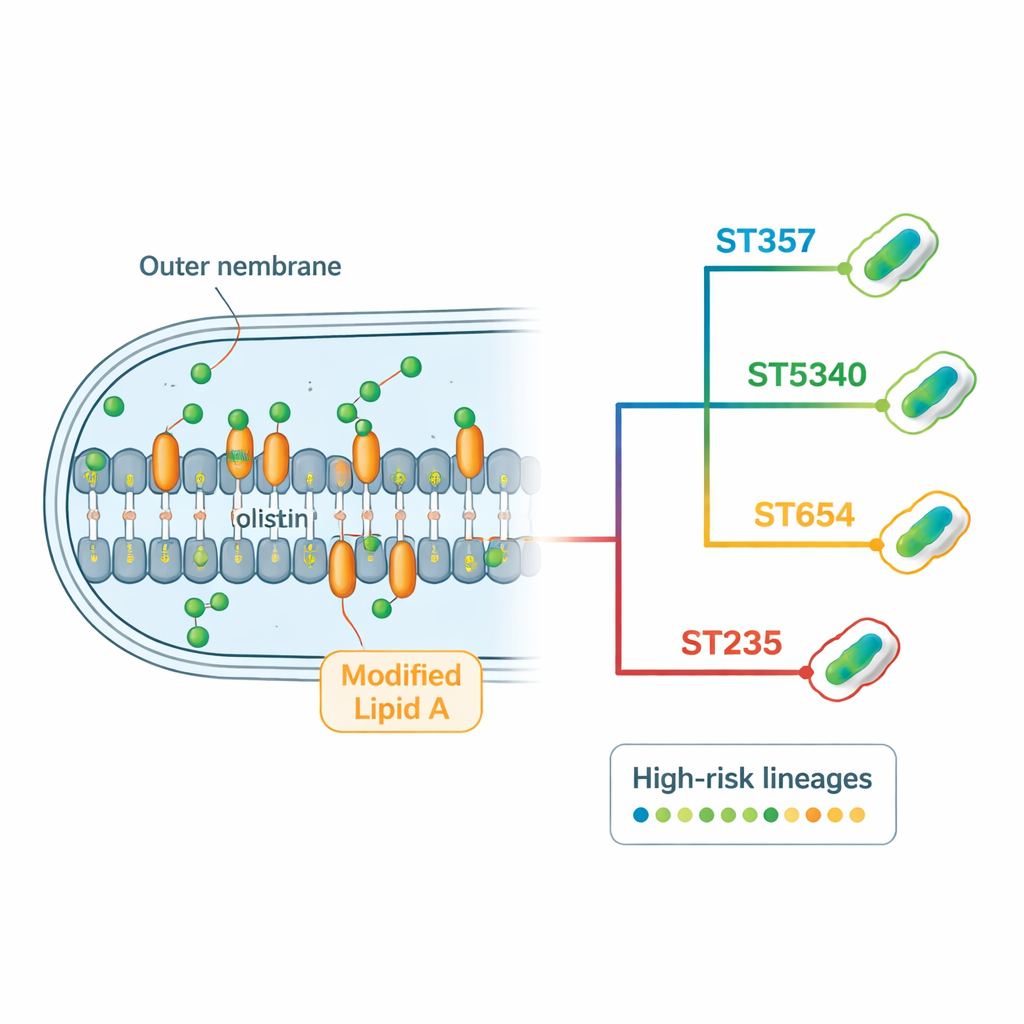
How these bacteria outsmart antibiotics
The genomic analysis revealed a packed “resistome”—the full set of resistance genes and mutations each strain carries. Many isolates encoded several kinds of beta‑lactamases, enzymes that break down common antibiotics such as penicillins, cephalosporins, and carbapenems. The carbapenemase gene blaNDM‑1, associated with resistance to some of the most powerful hospital drugs, appeared in nearly all strains, sometimes in multiple copies. The bacteria also carried genes that chemically modify aminoglycoside antibiotics, as well as powerful efflux pumps that work like molecular pumps to eject drugs from the cell. For colistin specifically, the team did not find mobile resistance genes but instead identified recurring changes in chromosomal genes involved in the outer cell membrane and its regulation. Certain mutations in regulatory proteins and in lipid A–building enzymes were strongly linked to colistin resistance, especially in the dominant ST357 and ST5340 lineages.
What this means for patients and hospitals
By combining national surveillance with modern genome sequencing, this study shows that Thailand’s hospitals are facing a growing threat from a newly recognized high‑risk clone, ST5340, alongside established global problem strains. These bacteria are not only resistant to colistin but also to many other key drugs, sharply narrowing treatment options when patients develop serious infections. For lay readers, the message is clear: antibiotic resistance is not an abstract future risk but a present‑day reality that can directly affect outcomes of surgery, intensive‑care treatment, and cancer therapy. The authors argue that continued genomic surveillance, stricter infection control, and careful antibiotic use are urgently needed to prevent these highly resistant lineages from becoming even more widespread and harder to contain.
Citation: Wankaew, N., Arigul, T., Kruasuwan, W. et al. Comparative genomics of colistin-nonsusceptible multidrug-resistant Pseudomonas aeruginosa reveals emerging lineages in Thailand. Sci Rep 16, 5968 (2026). https://doi.org/10.1038/s41598-026-35520-7
Keywords: Pseudomonas aeruginosa, antibiotic resistance, colistin, genomic surveillance, hospital infections