Clear Sky Science · en
Automated identification of contextually relevant biomedical entities with grounded LLMs
Why smarter tagging of medical papers matters
Every year, thousands of biomedical studies appear, each packed with details about genes, cell types, diseases, and treatments. Yet most of this information stays locked away in long PDFs, making it hard for other scientists to find the exact data they need. This article explores how modern artificial intelligence—large language models, or LLMs—can automatically pull out these key biomedical terms from research papers, helping to turn scattered publications into well-organized, searchable resources.
From messy papers to searchable building blocks
Biomedical research centers, like Germany’s Collaborative Research Centers, depend on clear, structured data to make studies reusable for years to come. Traditionally, researchers had to manually tag their datasets with important entities such as organisms, cell lines, and genes—a tedious and time-consuming task. LLMs can read full papers and understand context, making them promising tools for automating this tagging. But there is a catch: deciding which terms are truly relevant depends on the scientific question and how the data will be reused. The authors work within a carefully designed metadata scheme from the nephrology-focused CRC “NephGen,” which tells the AI what kinds of entities to look for and how they should be organized.
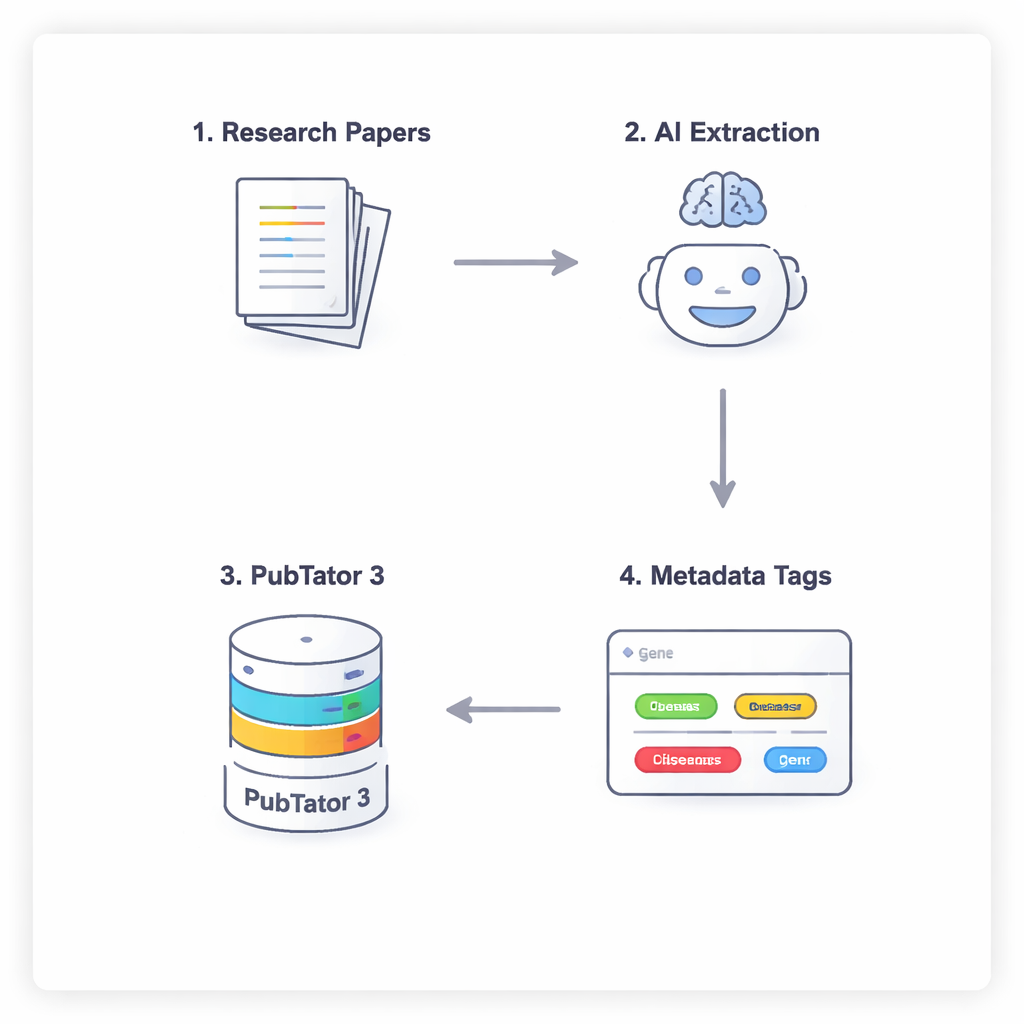
A four-step conversation between AI and a biology database
To keep the AI from simply guessing or “hallucinating” biomedical facts, the researchers use a four-step process that forces the models to reason carefully and double-check themselves. First, the model scans the full text of a paper (ignoring the discussion and references) to suggest potentially relevant entities. Second, it must consult an external tool, PubTator 3, a large biomedical database, to confirm that each suggested term actually exists and has a recognized identifier. Third, the AI assigns each confirmed entity to a slot in the NephGen metadata scheme, which groups entities in a hierarchical, human-designed structure. Finally, the model consolidates all of this into a structured JSON output, essentially a tidy machine-readable summary of key biomedical entities in the article.
Testing eight AI models with real kidney research
The team implemented this workflow using APIs for 14 different LLMs and found that only eight could reliably follow the strict requirements, such as returning valid JSON and correctly using tools. They then applied these eight models to six nephrology research articles and asked each paper’s author to review the AI’s final list of entities in a short, face-to-face interview. Because there is no fixed “correct” number of entities to extract, the authors focused on precision: what fraction of the suggested entities the scientists judged to be correct. Using statistical meta-analysis methods adapted for proportions close to 100%, they estimated precision for each model while accounting for variation between papers.
High accuracy, but trade-offs in effort, cost, and speed
Across all models, the AI systems achieved an overall precision of about 91%, meaning that the great majority of suggested entities were judged correct. GPT-4.1, GPT-4o Mini, and Gemini 2.0 Flash had the highest precision—roughly 94% to 98%—though their differences were not statistically clear. Gemini models tended to propose more entities overall, leading to more correct tags but also more for humans to check. Some smaller or cheaper models, such as GPT-4.1 Nano, were faster and inexpensive but substantially less accurate. The authors visualized these tensions using Pareto frontiers, identifying combinations of models that balanced precision, number of correct entities, cost, and processing time: for example, GPT-4o Mini emerged as particularly attractive when both accuracy and low cost are priorities.
Why humans still belong in the loop
Despite strong performance, the study highlights important limitations. The models sometimes mixed up information about the published article with details that were not truly relevant to the underlying dataset that future users might want to reuse. This confusion reflects a broader challenge in automated text mining: scientific papers discuss far more than what ends up in a shared dataset. The authors therefore recommend that human experts continue to review AI-generated annotations before they are published. They also note that their evaluation covers only six nephrology papers, so broader testing across fields is needed. Over time, a routine “human-in-the-loop” workflow could build a consensus reference set, making it possible to measure not just precision but also how many entities the AI missed.
What this means for future biomedical data sharing
The study shows that, when carefully guided and grounded in trusted databases, modern LLMs can reliably help annotate biomedical papers, greatly reducing the manual burden on researchers. The best models approach expert-level precision while offering a range of trade-offs between thoroughness, cost, and speed. For now, human review remains essential to ensure that annotations truly match the datasets and research context. But as tools and open-source models continue to mature, workflows like this could become a standard backbone for turning today’s flood of medical papers into tomorrow’s well-organized, reusable data commons.
Citation: Watter, M., Giuliani, C., Benadi, G. et al. Automated identification of contextually relevant biomedical entities with grounded LLMs. Sci Rep 16, 1952 (2026). https://doi.org/10.1038/s41598-026-35492-8
Keywords: biomedical text mining, large language models, metadata annotation, grounded AI, nephrology research