Clear Sky Science · en
Immunoinformatics-based design and evaluation of a multi-epitope vaccine against Vibrio fluvialis
Why a New Vaccine Matters for Seafood Lovers
Vibrio fluvialis is a lesser-known cousin of cholera-causing bacteria that lives in warm coastal waters and can contaminate seafood and drinking water. It causes severe diarrhea and gut infections and is becoming more common as oceans warm and antibiotics lose their power. Yet there is currently no vaccine for people. This study uses advanced computer methods to design a new kind of experimental vaccine, built from carefully chosen tiny fragments of the bacterium, with the long-term goal of protecting people who are exposed through food, water, or climate‑driven floods.

From Hidden Threat to Clear Target
Vibrio fluvialis has often been mistaken for other Vibrio species, which means its role in foodborne disease has likely been underestimated. Outbreaks have been linked to contaminated seafood and polluted water, and recent storms and floods have brought these bacteria inland, sickening people far from the coast. Many patients require hospital care, and a worrying number of strains now resist multiple antibiotics. Because the microbe’s surface looks similar to that of related bacteria, doctors can misdiagnose infections, delaying proper treatment. All of this creates a strong need for prevention rather than relying only on drugs.
Designing a Vaccine Inside a Computer
Instead of growing whole bacteria in the lab, the researchers turned to “immunoinformatics” — using software to predict which small pieces of Vibrio fluvialis are most likely to be seen and attacked by the human immune system. They focused on two membrane proteins that sit on the bacterium’s surface and are important for its survival and ability to cause disease. From these proteins, the team extracted ten short segments, called epitopes, that can be recognized by both T cells and B cells, the main soldiers of our immune defenses. They then stitched these pieces together into a single 246–amino acid chain, adding short spacer segments so the immune system can process each piece properly, and an adjuvant fragment to boost the overall response.
Checking Strength, Safety, and Global Reach
Once the virtual vaccine was built, the team ran a series of tests entirely in silico, meaning on computers rather than in animals or people. Programs predicted that the combined construct would be strongly “visible” to the immune system but unlikely to trigger allergy or toxicity. The chosen epitopes matched common immune gene variants from around the world, suggesting that about 99.97 percent of the global population should be able to respond, including people in heavily affected regions such as South and East Asia. The vaccine protein also appeared stable, water‑loving enough to dissolve well, and suitable for efficient production in standard laboratory bacteria, making eventual manufacturing more practical.
How the Vaccine May Spark Protection
The researchers next asked whether the designed protein would plausibly engage with a real immune sensor. Using modern structure‑prediction tools, they built a three‑dimensional model of the vaccine and then simulated how it might dock onto Toll‑like receptor 2 (TLR2), a molecule on immune cells that detects bacterial components. Computer docking suggested a tight fit between the vaccine and TLR2, supported by many stabilizing molecular contacts. Long, detailed motion simulations over 100 nanoseconds indicated that the vaccine‑receptor pair remained stable and compact over time. Additional immune‑system simulations predicted strong waves of antibodies and T‑cell activity, along with the formation of memory cells that, in principle, could provide lasting protection against Vibrio fluvialis.
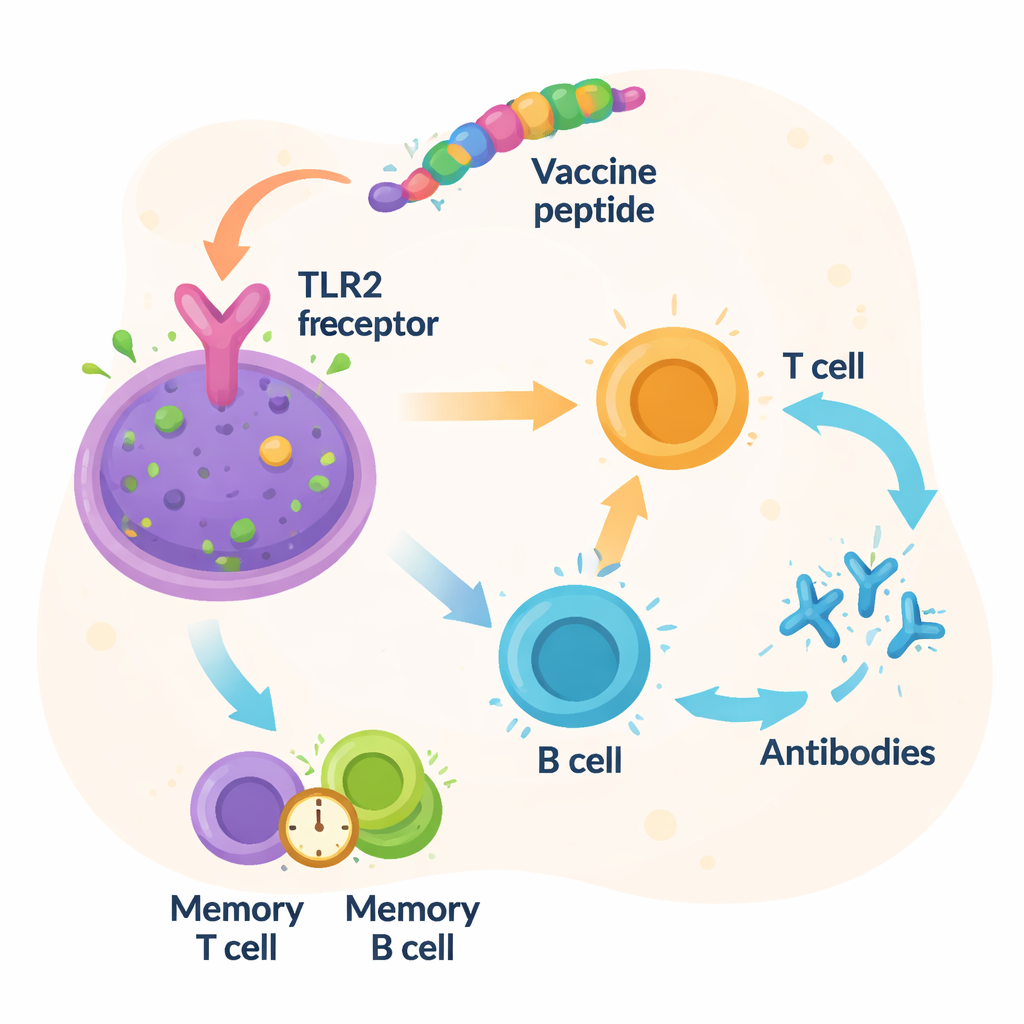
From Digital Blueprint to Real‑World Shield
In everyday terms, this work offers a digital “blueprint” for a future Vibrio fluvialis vaccine built from just the most important slivers of the bacterium, arranged to be widely effective and potentially safer than whole‑cell vaccines. The study shows that this construct should be recognizable to most human immune systems, bind well to a key immune sensor, and be manufacturable using standard tools. However, all of these results come from computer models. The next steps will require producing the vaccine in the lab, testing it in cells and animals, and eventually in human trials to confirm that it truly protects against infection without causing harm. If those tests succeed, such multi‑epitope vaccines could become powerful new tools against foodborne bacteria in a warming, increasingly crowded world.
Citation: Naveed, M., Husnain, M., Aziz, T. et al. Immunoinformatics-based design and evaluation of a multi-epitope vaccine against Vibrio fluvialis. Sci Rep 16, 4100 (2026). https://doi.org/10.1038/s41598-026-35434-4
Keywords: Vibrio fluvialis, multi-epitope vaccine, foodborne infection, immunoinformatics, antibiotic resistance