Clear Sky Science · en
Development of a global screening system for detecting protein–protein interactions by luminescence complementation in fission yeast
Lighting Up the Hidden Life of Proteins
Every heartbeat, thought, and breath in our bodies depends on proteins teaming up inside cells. These tiny partnerships, called protein–protein interactions, control everything from how genes are read to how cells respond to stress. Yet many of these partnerships are fleeting or delicate, making them hard to detect with current tools. This study describes a new, highly sensitive "light-based" screening system built in a simple model organism, fission yeast, to reveal thousands of such hidden protein connections at once.
Why Protein Partnerships Are So Hard to See
Proteins rarely work alone. They latch onto different partners at different times, sometimes only for moments, and their behavior can be altered by chemical tweaks or by where they sit in the cell. Traditional methods to study these interactions either pull proteins out of their natural environment or rely on genetic tricks that do not catch every pairing. As a result, existing maps of who interacts with whom in the cell are incomplete. The authors set out to improve this picture by creating a more flexible, sensitive system that could survey nearly every protein in fission yeast in living cells.
Turning Touch into Light
At the heart of the new approach is a split enzyme that gives off light. One part, called LgBiT, is attached to a "bait" protein the researchers want to study. The other part, SmBiT, is attached to many possible "prey" partners, one per yeast strain. On their own, these two enzyme pieces barely interact. But when bait and prey proteins bind each other inside the cell, the two pieces snap together, re-forming a working enzyme that glows in the presence of a chemical fuel. By measuring how much light each yeast strain produces, the team can tell which proteins are actually meeting inside the cell and how strongly they interact. To avoid missing partners because the tag blocks normal function, the bait protein is made in two versions, with LgBiT attached to either end, greatly increasing the odds that at least one orientation will work. 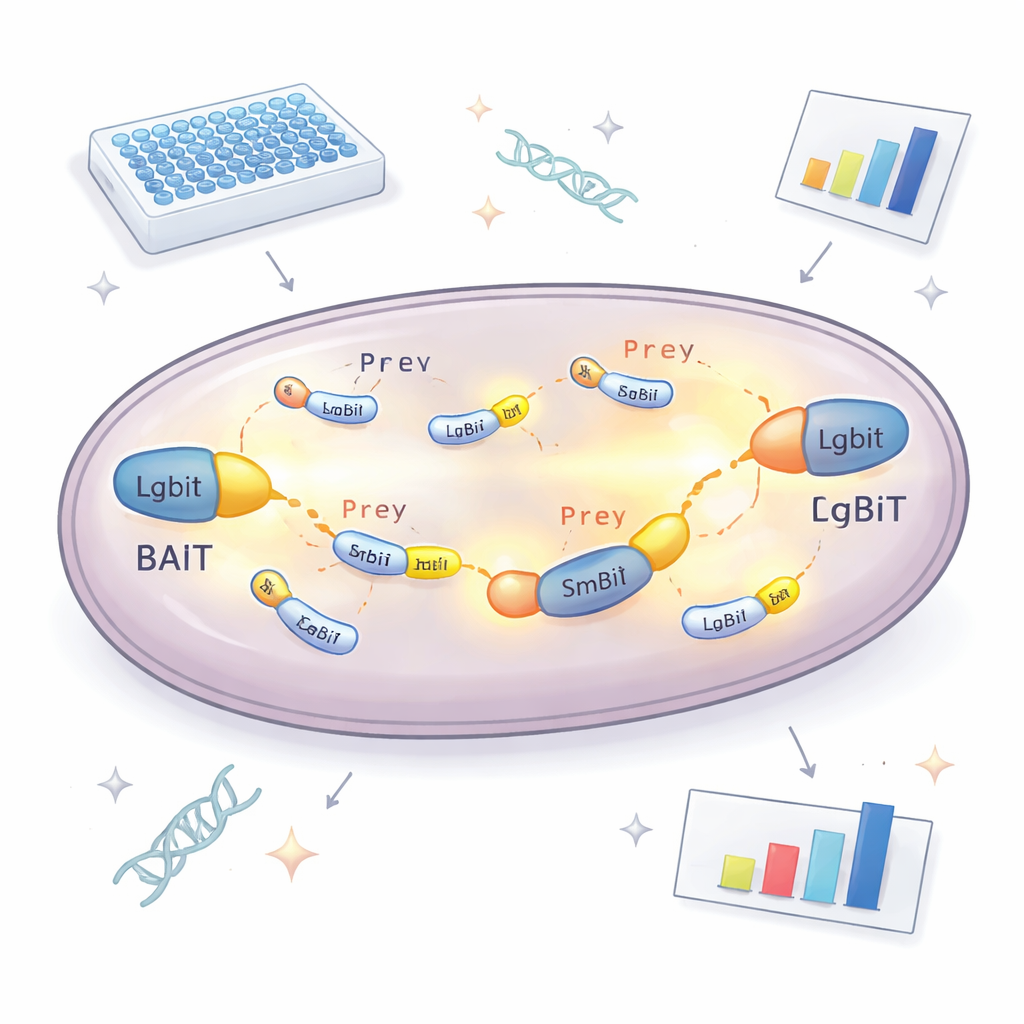
Engineering a Yeast Library and Strain System
Building this platform required more than a clever enzyme. The researchers also had to introduce thousands of prey proteins into yeast in a controlled, single-copy manner. They designed a special DNA vector for prey proteins that inserts into a defined spot in the yeast genome and carries a red fluorescent marker. The brightness of this red signal reflects how many copies are inserted, allowing the team to easily select colonies with exactly one copy, which is essential for reliable comparisons. They also created dedicated bait vectors that place LgBiT at the beginning or end of the bait protein, separated by flexible linkers so the protein can still fold and move normally. Carefully engineered yeast strains were then built to host two bait constructs and one prey construct at once, using nutritional markers to ensure only correctly combined cells survive.
Putting the System to the Test
To check whether their design really improved detection, the scientists first examined proteins with well-known partners. They confirmed that signal strength depended strongly on where the light-emitting tag was attached: some interactions only showed up when LgBiT was on one end of the bait, and not the other. Expressing both versions together boosted the light output, proving that the dual-bait strategy makes weak or orientation-sensitive interactions easier to catch. The system also detected contacts involving membrane-bound proteins, a class that is notoriously difficult for traditional yeast two-hybrid methods. With this validation in hand, the team scaled up to a genome-wide screen, using a transcription-related protein called Tfs1 as bait. Screening more than 4,600 prey proteins, they found 31 strong candidates, including several already known partners and many new ones, some located outside the nucleus in areas like mitochondria and the cell surface. 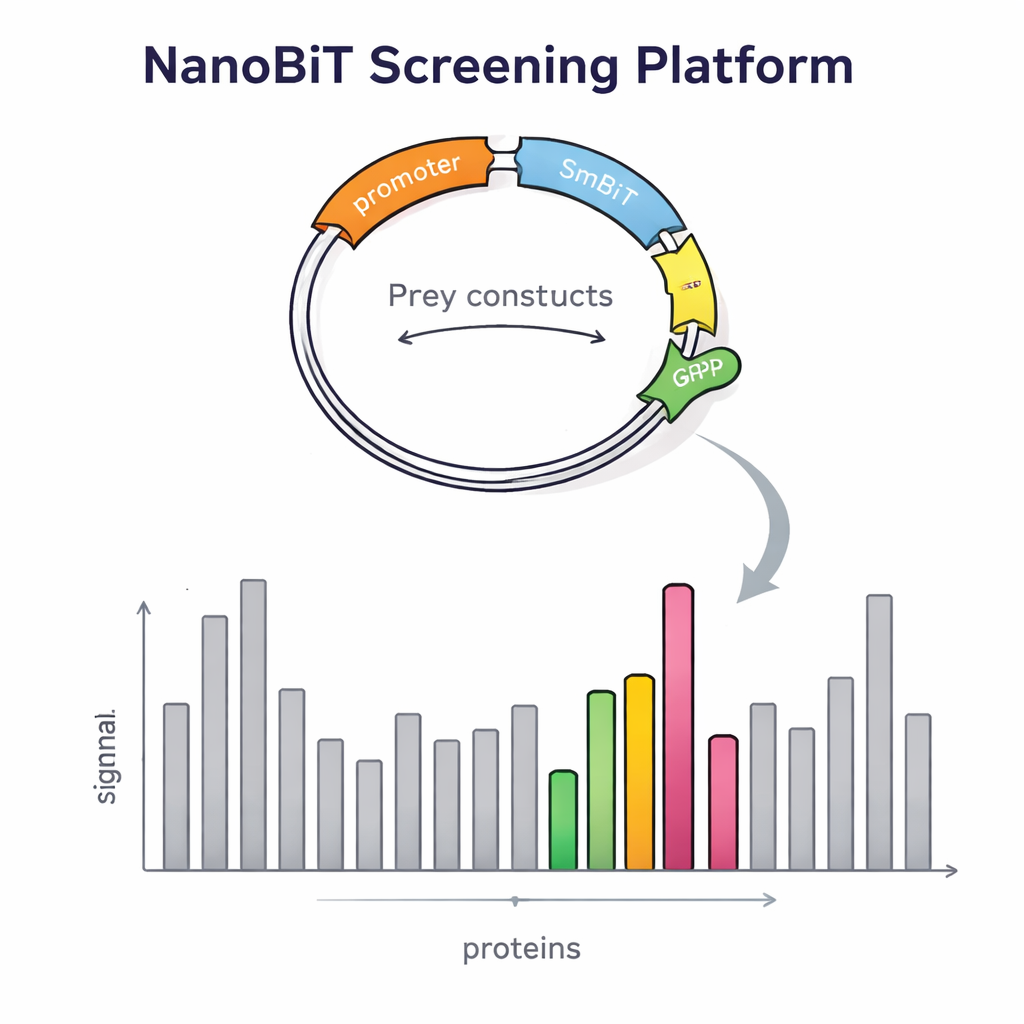
What This Means for Biology and Drug Discovery
To a non-specialist, the key message is that the authors have built a more sensitive "interaction radar" inside yeast cells. By converting brief protein handshakes into measurable flashes of light, and by cleverly arranging the genetics so three tagged proteins can be expressed together, their system uncovers both known and previously invisible protein partners. While further checks with other techniques are still needed to confirm each new pairing, this platform fills in important gaps in our maps of cellular networks. Because the strains were also engineered to be more sensitive to small molecules, the same setup could be adapted to search for drugs that either disrupt harmful protein partnerships or stabilize beneficial ones, offering a powerful new tool for both basic biology and early-stage drug discovery.
Citation: Azadeh, F., Hashimoto, A., Nishimura, S. et al. Development of a global screening system for detecting protein–protein interactions by luminescence complementation in fission yeast. Sci Rep 16, 5034 (2026). https://doi.org/10.1038/s41598-026-35430-8
Keywords: protein interactions, fission yeast, luminescence assay, high-throughput screening, drug discovery tools