Clear Sky Science · en
A lysine-free REC tag system for proximity-biotinylation applications
Why tiny tags on proteins matter
Inside every cell, proteins rarely work alone—they team up in shifting partnerships that drive health and disease. To study these partnerships, scientists often attach small “name tags” to proteins so they can be tracked, pulled out of a mixture, or seen under a microscope. A popular method called proximity biotinylation lets researchers map who is standing near whom inside the cell, but it comes with a hidden problem: many of these name tags become chemically modified and effectively disappear just when they are needed most. This study introduces a new kind of tag, called the REC tag, designed to stay visible under these demanding conditions.
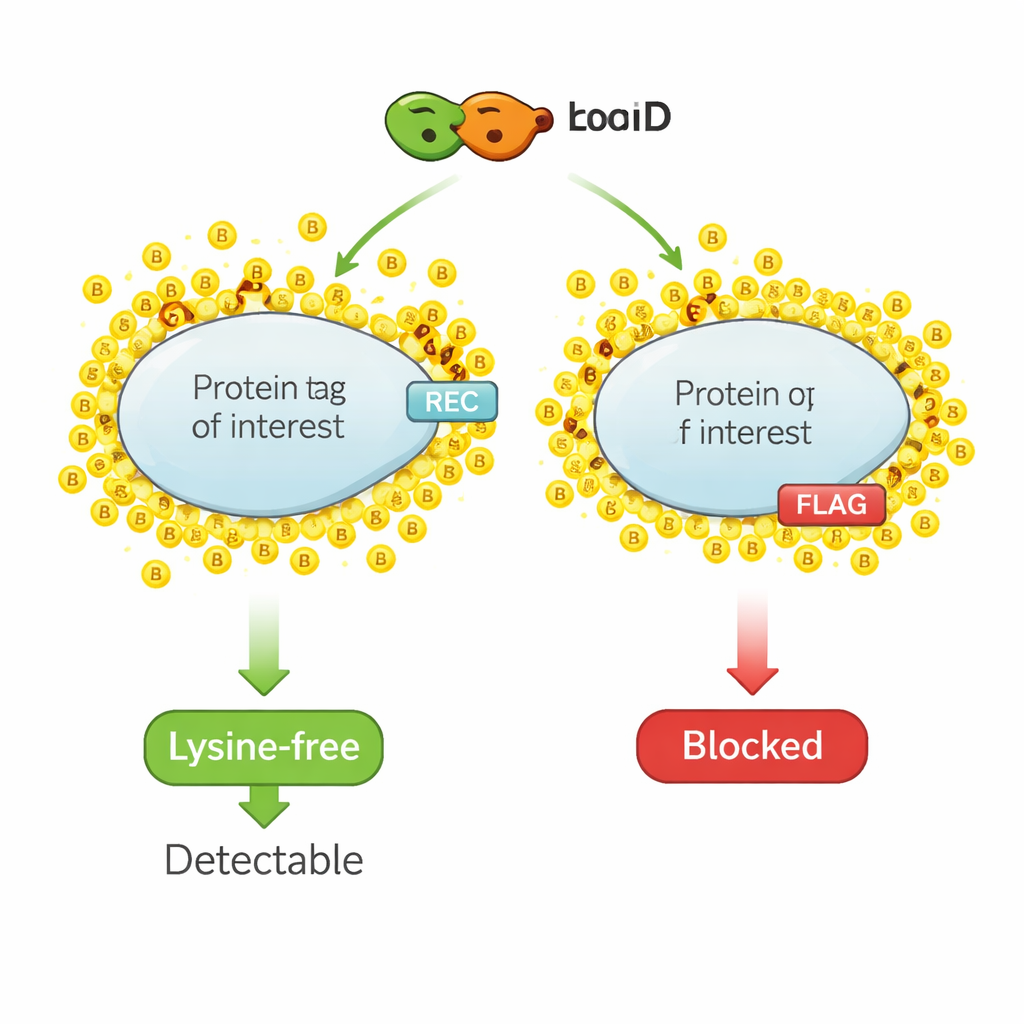
A chemical highlighter for protein neighbors
Proximity biotinylation works like a molecular highlighter pen. An engineered enzyme, such as TurboID, is fused to a protein of interest. When biotin—a small vitamin—is added, TurboID rapidly attaches biotin molecules to nearby proteins. Researchers then fish out all the biotin-marked proteins and identify them, gaining a snapshot of the protein’s local neighborhood. However, the same chemistry that labels nearby proteins also targets the small epitope tags commonly used for detection, because those tags usually contain lysine, an amino acid that is the main landing site for biotin. When lysines in the tag are covered with biotin, antibodies can no longer recognize the tag, and the signal in standard detection methods drops or vanishes.
Designing a tag that refuses modification
To solve this, the authors set out to create a new tag completely free of lysine. They started from a fragment of a malaria parasite protein called PfRipr5, which has little similarity to human proteins, reducing the risk of unwanted cross-reactions. Using a microchip-based screening method in rabbits, they generated 22 monoclonal antibodies against PfRipr5 and identified one, dubbed clone No. 6, with especially strong and specific binding. By systematically trimming and mutating the PfRipr5 fragment, they narrowed the antibody’s target down to an 11–amino acid stretch. This minimal sequence contained only one lysine, which they replaced with a similar amino acid, arginine. Remarkably, the antibody still bound just as well. This optimized, lysine-free sequence was named the REC tag, and the matched antibody the REC antibody.
Putting the REC tag through its paces
The team then tested whether the REC tag could stand in for widely used tags such as FLAG in common lab techniques. When fused to fluorescent or signaling proteins and produced in cell-free systems and mammalian cells, REC-tagged proteins were readily detected by immunoblotting, a high-sensitivity light-based plate assay called AlphaScreen, and immunostaining in cells. Detection strength was similar to that of the FLAG tag and to another lysine-free tag system (AGIA), and the REC antibody showed very little background noise across a variety of human and monkey cell lines. Importantly, the tag worked whether placed at the beginning or end of a protein, and on proteins located in diverse cellular regions, including the nucleus, mitochondria, endoplasmic reticulum, and cell membrane.
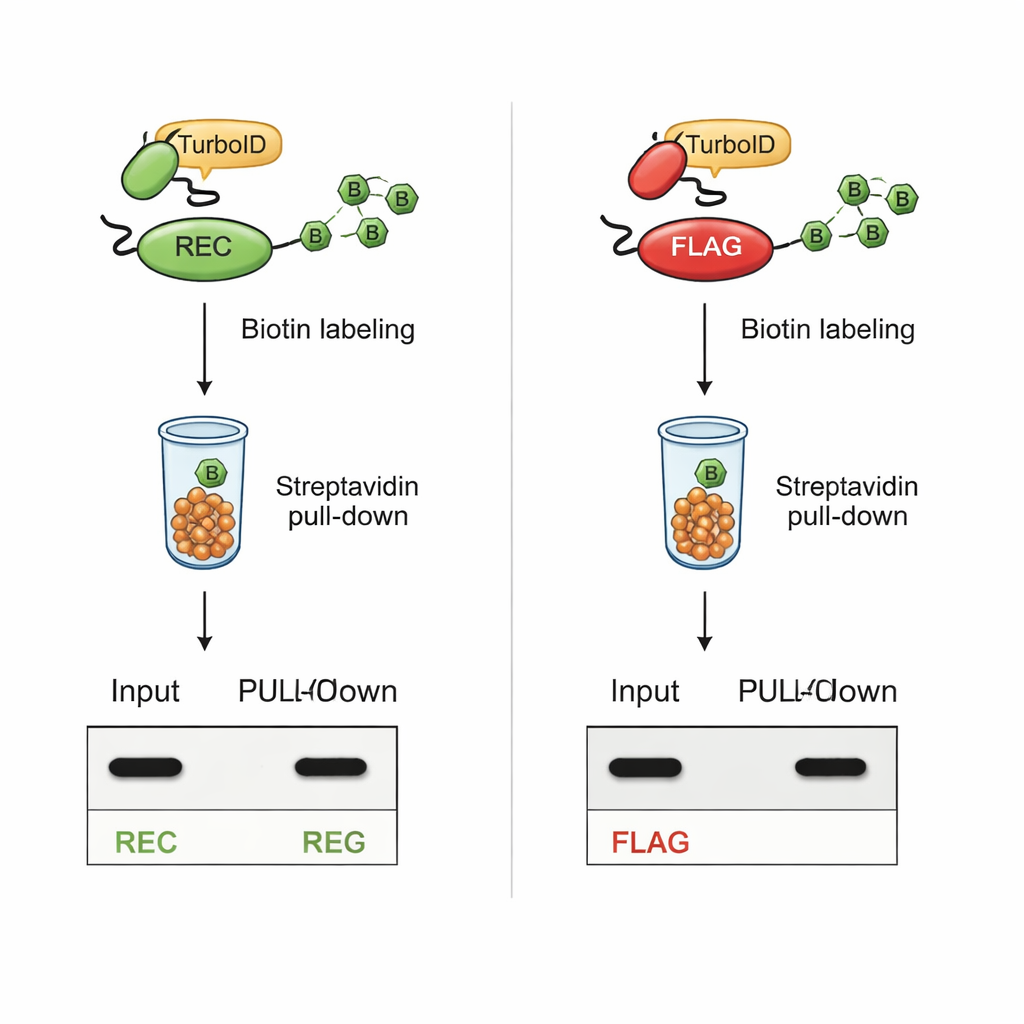
Staying visible during intense labeling
The critical test was whether REC would remain detectable after TurboID-based biotinylation. In cells where TurboID was fused to known interaction partners, proteins carrying the REC tag stayed clearly visible in both whole-cell extracts and in the purified, biotin-labeled fraction. In contrast, proteins carrying the classic FLAG tag lost their FLAG signal once TurboID was active, even though the proteins themselves were still present. Additional experiments with synthetic peptides showed that TurboID directly biotinylates the lysines in the FLAG sequence, physically blocking antibody access. The REC tag, lacking lysine altogether, was untouched by TurboID and remained easy to detect.
What this means for future experiments
For researchers exploring protein networks inside living cells, the REC tag offers a practical new tool: it behaves like familiar tags in everyday assays but continues to work even when powerful proximity-labeling enzymes are active. This means scientists can more reliably confirm that their tagged proteins are present and correctly captured, and can study other lysine-based modifications—such as ubiquitination and acetylation—without the tag itself interfering. In short, REC is a small design change with a big impact, helping make complex protein-mapping experiments clearer and more trustworthy.
Citation: Tokunaga, S., Nagaoka, H., Ozawa, T. et al. A lysine-free REC tag system for proximity-biotinylation applications. Sci Rep 16, 4846 (2026). https://doi.org/10.1038/s41598-026-35323-w
Keywords: protein tagging, proximity biotinylation, TurboID, protein interactions, epitope tags