Clear Sky Science · en
Genetic characterization and whole-genome sequencing-based genetic analysis of influenza B in Shandong Province during 2015–2024
Why local flu viruses matter to you
Seasonal flu is often brushed off as a minor illness, but it quietly infects hundreds of millions of people every year and can be deadly, especially for children and older adults. This study tracks how one branch of the flu family—influenza B—has changed over nearly a decade in Shandong Province, a populous coastal region of China. By reading the genetic code of local viruses, the researchers show how well current vaccines match what is actually circulating, which age groups are hit hardest, and where signs of emerging drug resistance may be appearing.
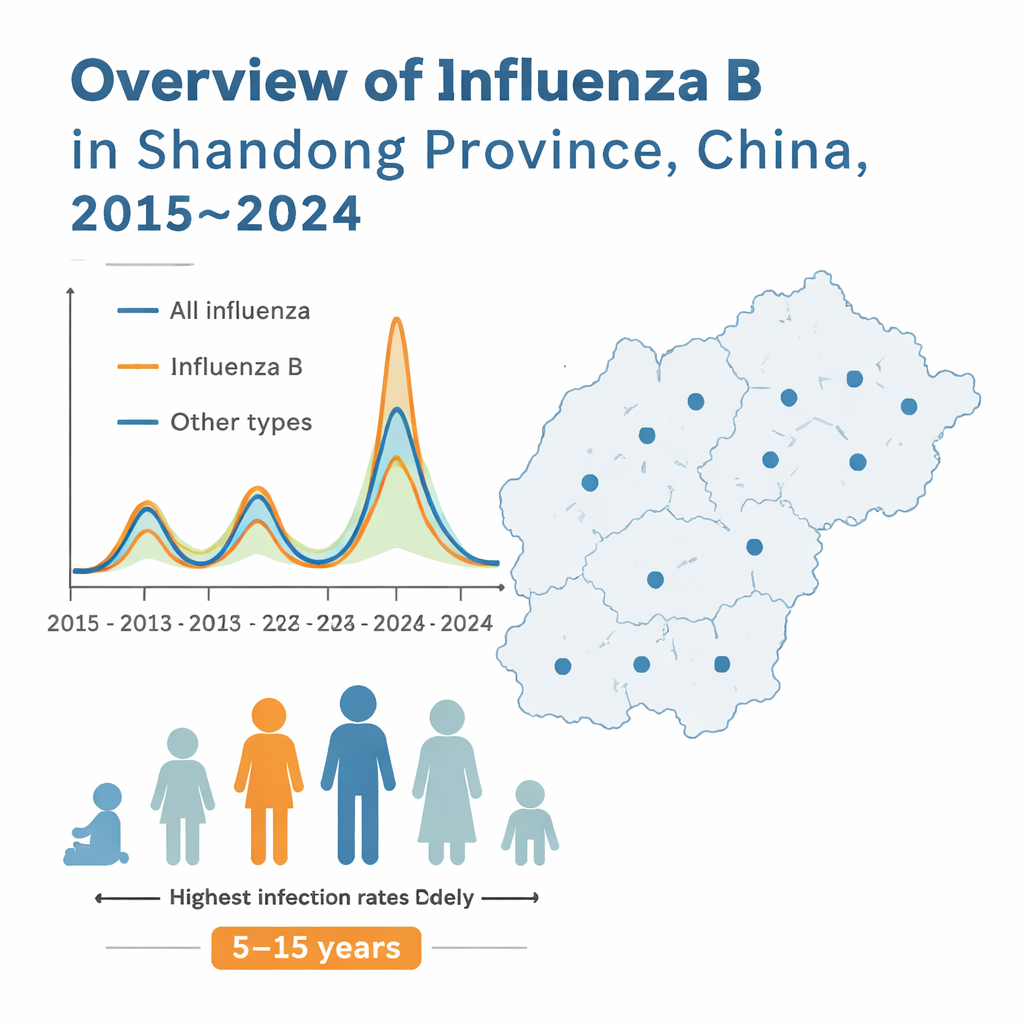
Watching flu waves roll through a province
Between 2015 and early 2024, hospitals across Shandong collected throat and nose swabs from almost half a million people with “influenza-like illness” (fever plus cough or sore throat). About one in eight patients actually had an influenza virus, and roughly one third of these infections were caused by influenza B. Flu activity usually peaked in the cold months, but the pattern was disrupted during the height of the COVID-19 pandemic, when mask-wearing and other precautions drove flu detections close to zero. When influenza did surge, school-age children between 5 and 15 years old were the most likely to test positive, while older adults over 60 had the lowest detection rates, highlighting children as key drivers of community spread.
Two branches of influenza B and changing dominance
Influenza B comes in two long-standing branches, named B/Victoria and B/Yamagata. Both circulated in Shandong early in the study, but the Yamagata branch gradually disappeared after 2018, mirroring trends seen worldwide. From then on, nearly all influenza B detections in the province were of the Victoria branch. The team selected 109 representative viruses—76 Victoria and 33 Yamagata—and read the full genetic sequence of all eight segments of their RNA genomes. These were then compared to the World Health Organization’s recommended vaccine strains for each year, as well as to reference viruses from around the globe, to see how closely local viruses tracked global evolution.
Small genetic changes with big consequences
Most Shandong influenza B viruses remained closely related to the vaccine strains, but year by year they accumulated a series of small mutations, especially in two surface proteins called hemagglutinin (HA) and neuraminidase (NA). These proteins sit on the virus’s outer shell and are the main targets of our immune system and of current antiviral drugs. The study found repeated changes in key “hot spots” on HA that antibodies normally recognize. In several time periods, clusters of viruses carried four or more changes in these sites, enough to qualify as potential “antigenic drift” variants that could blunt vaccine protection. One mutation at position 197 of HA, affecting both an immune target and the receptor-binding region, became especially common and also removed a sugar attachment (“glycosylation”) that may alter how the protein is seen by the immune system.
Mixing, matching, and resisting medicines
Beyond gradual mutation, the study spotted signs of genetic shuffling between virus lineages. One B/Victoria virus carried an internal gene from the B/Yamagata branch, clear evidence of reassortment—a process in which two viruses co-infect a cell and swap genome segments, potentially creating new variants. The researchers also scanned for mutations linked to drug resistance. Among 109 viruses, they found one with a change called H273Y in the NA protein, known to reduce sensitivity to commonly used neuraminidase-blocking drugs such as oseltamivir; lab testing confirmed reduced drug susceptibility. Other internal genes showed mutations that may influence how efficiently the virus replicates or escapes early immune responses, although these did not yet point to resistance against newer antiviral pills that target the polymerase machinery.
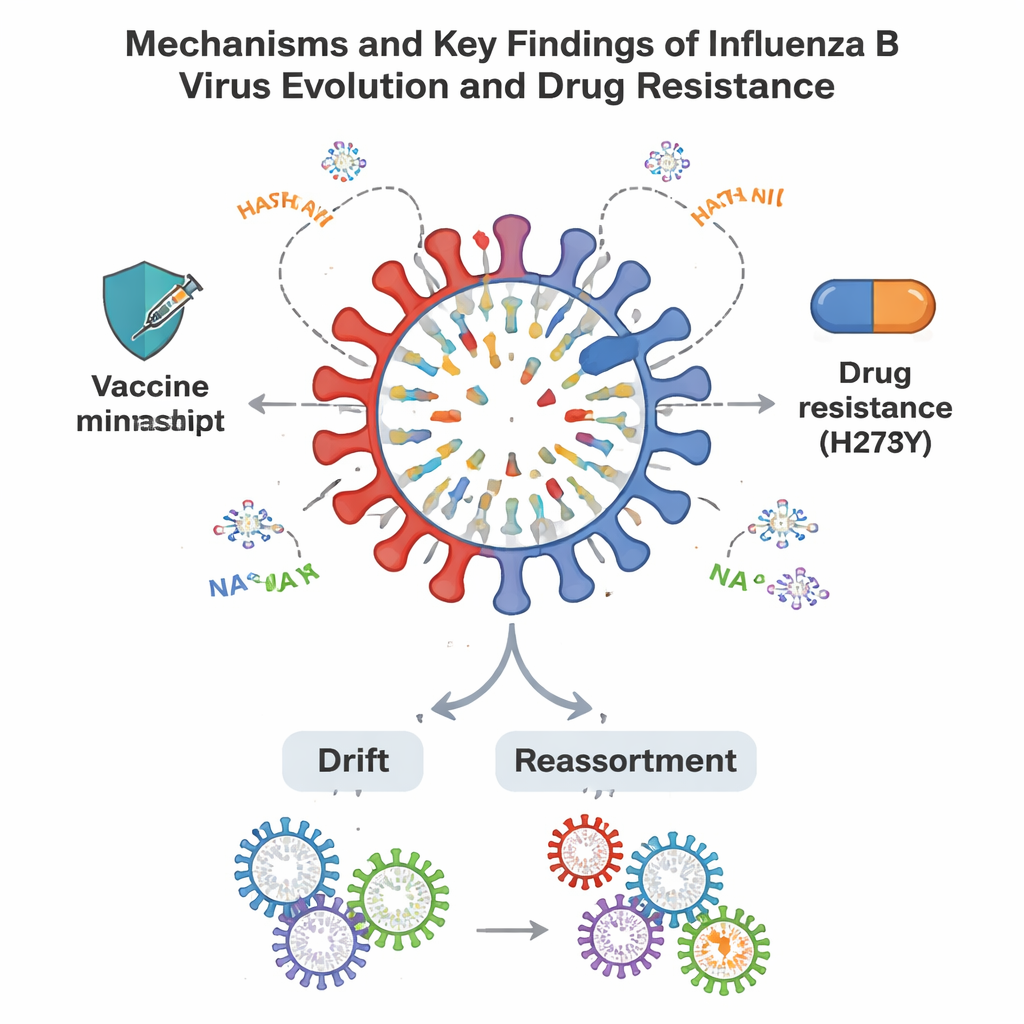
What this means for vaccines and prevention
Overall, the influenza B viruses circulating in Shandong from 2015 to 2024 still resembled the strains chosen for seasonal vaccines, supporting the value of annual flu shots. Yet the steady build-up of changes in key immune-targeted regions, the disappearance of one major lineage, occasional gene swapping, and the appearance of a drug-resistant virus all signal that influenza B is far from static. For the public, the takeaway is straightforward: vaccination remains the best shield, especially for school-age children who drive much of the spread. For health authorities, the study underscores the need for continuous genetic monitoring of local flu viruses, regular updates to vaccine composition, and vigilant testing for antiviral resistance so that treatments and prevention strategies keep pace with an ever-evolving foe.
Citation: Wu, J., He, Y., Sun, L. et al. Genetic characterization and whole-genome sequencing-based genetic analysis of influenza B in Shandong Province during 2015–2024. Sci Rep 16, 5229 (2026). https://doi.org/10.1038/s41598-026-35291-1
Keywords: influenza B, genomic surveillance, antigenic drift, antiviral resistance, vaccine effectiveness