Clear Sky Science · en
Deubiquitination-related genes define immune subtypes of colorectal cancer and are associated with prognosis and immunotherapy-related signatures
Why this study matters for people
Colorectal cancer is one of the most common and deadly cancers worldwide, yet people with tumors that look similar under the microscope can have very different outcomes and responses to modern treatments such as immunotherapy. This study asks a new question: can we sort colorectal cancers into clearer groups by looking at how tumor cells handle protein "recycling" and how this links to the body’s immune defenses? The answer could help doctors better predict prognosis and design smarter combinations of therapies in the future.
Protein cleanup and cancer behavior
Inside every cell, worn-out or damaged proteins are tagged and recycled so they do not build up and cause trouble. One arm of this system, called deubiquitination, removes those tags and helps fine-tune which proteins are destroyed and which are spared. The authors gathered large public datasets of bowel tumors and healthy tissue and scanned thousands of genes to find those tied both to this protein cleanup system and to patient survival. They narrowed the list to 17 key genes that were strongly linked to cell division control, repair of DNA damage, and the structure of the material that surrounds cells. These genes became the backbone for a new way of grouping colorectal cancers.
Two main tumor types emerge
Using patterns of activity in the 17 genes, the researchers split tumors into two main subtypes. People in one group tended to live longer and had earlier-stage disease. The other group had a worse outlook. When the team looked more broadly at which genes were switched on or off in each subtype, they saw that the poor-outcome group showed strong signals of rapid cell growth, stress responses to damaged DNA, and heavy remodeling of the tissue scaffold surrounding the tumor. In contrast, the better-outcome group showed less of this aggressive remodeling and a more balanced pattern of cell growth and repair.
The tumor’s neighborhood and the immune response
Cancer does not grow in isolation; it lives within a neighborhood of immune cells, support cells, and scar-like tissue. The study used computational tools to estimate which immune cells were present in each tumor. The two subtypes showed strikingly different immune landscapes. The poor-outcome subtype was enriched for dense collagen and fibrous tissue, forming a physical and chemical barrier that can keep cancer-fighting T cells out. It also carried signals of immune suppression and higher scores on measures that predict resistance to immunotherapy. The better-outcome subtype had weaker matrix buildup, more favorable mixes of immune cells such as killer T cells and helper T cells, and scores suggesting greater potential sensitivity to immune-based treatments.
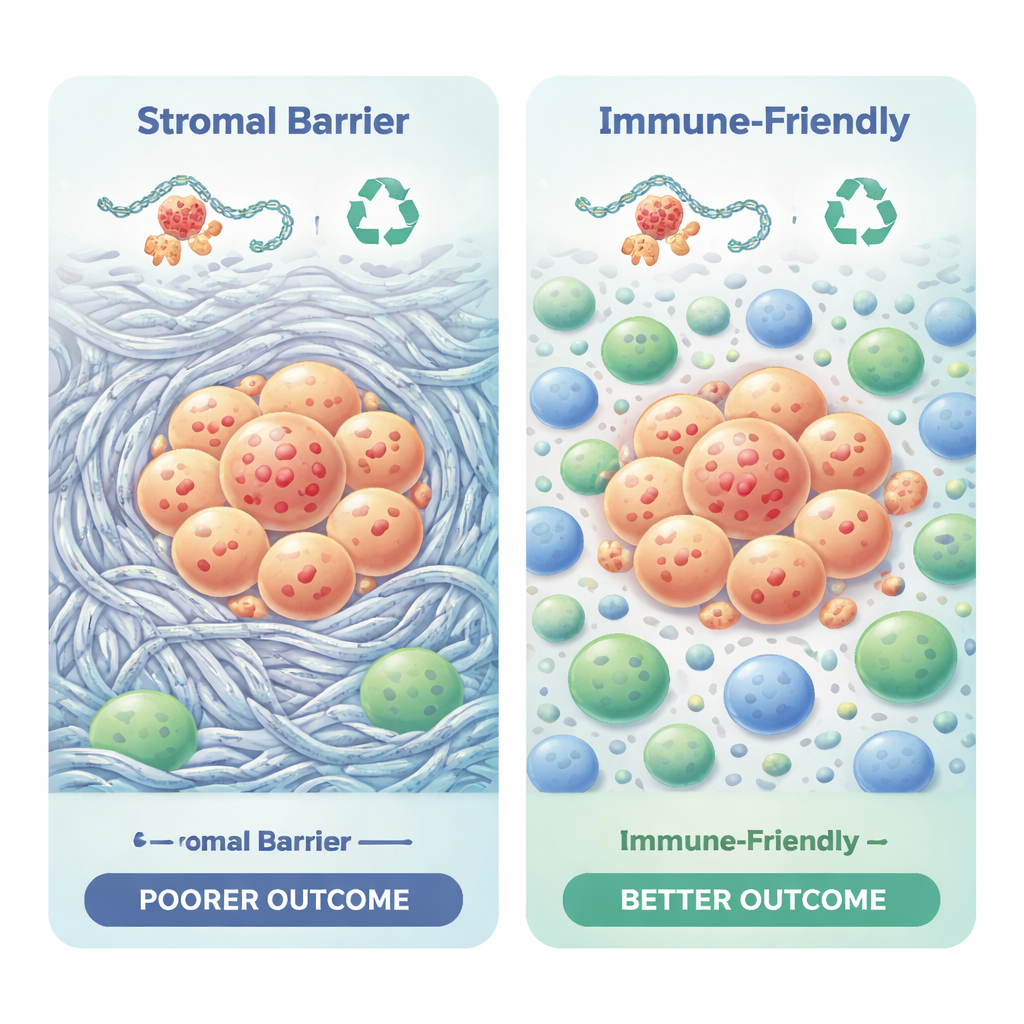
Key structural genes as warning beacons
To move from broad patterns to practical markers, the authors built networks of interacting proteins and searched for central "hub" genes. They identified nine genes largely involved in building and remodeling the tumor’s surrounding scaffold, including several collagens and molecules like fibronectin and periostin. High activity of some of these genes, especially BGN, FN1, and POSTN, consistently signaled worse survival in two independent patient groups. These hub genes sit at the crossroads of mechanical stiffness, chemical signaling, and immune cell recruitment, making them attractive candidates for future tests that might help predict risk or guide therapy choices.
What this means going forward
This work does not yet change how patients are treated, because it is based on computer analysis of existing data rather than new clinical trials. Still, it offers a clear big-picture takeaway for non-specialists: how a colorectal tumor manages protein recycling and reshapes its local environment appears to help decide whether the immune system can reach and attack it. Tumors with highly disrupted protein control and a thick, fibrous shell tend to fend off immune cells and are linked to poorer outcomes, while tumors with less scarring and more open access for immune cells fare better. In the future, combining drugs that target DNA repair or the fibrous matrix with immunotherapy may prove especially helpful for the high-risk group defined in this study.
Citation: Xu, Y., Mo, Z., Jiang, Q. et al. Deubiquitination-related genes define immune subtypes of colorectal cancer and are associated with prognosis and immunotherapy-related signatures. Sci Rep 16, 4862 (2026). https://doi.org/10.1038/s41598-026-35271-5
Keywords: colorectal cancer, tumor microenvironment, immune subtypes, protein degradation, immunotherapy