Clear Sky Science · en
Alzheimer’s disease prediction using deep learning and XAI based interpretable feature selection from blood gene expression data
Why this research matters
Alzheimer’s disease slowly robs people of their memory and independence, yet today’s most accurate tests often require brain scans or spinal taps that are expensive, invasive, and hard to repeat. This study explores a less painful alternative: using a simple blood draw and advanced computer analysis to spot patterns in gene activity that signal Alzheimer’s, potentially paving the way for earlier and more accessible diagnosis.
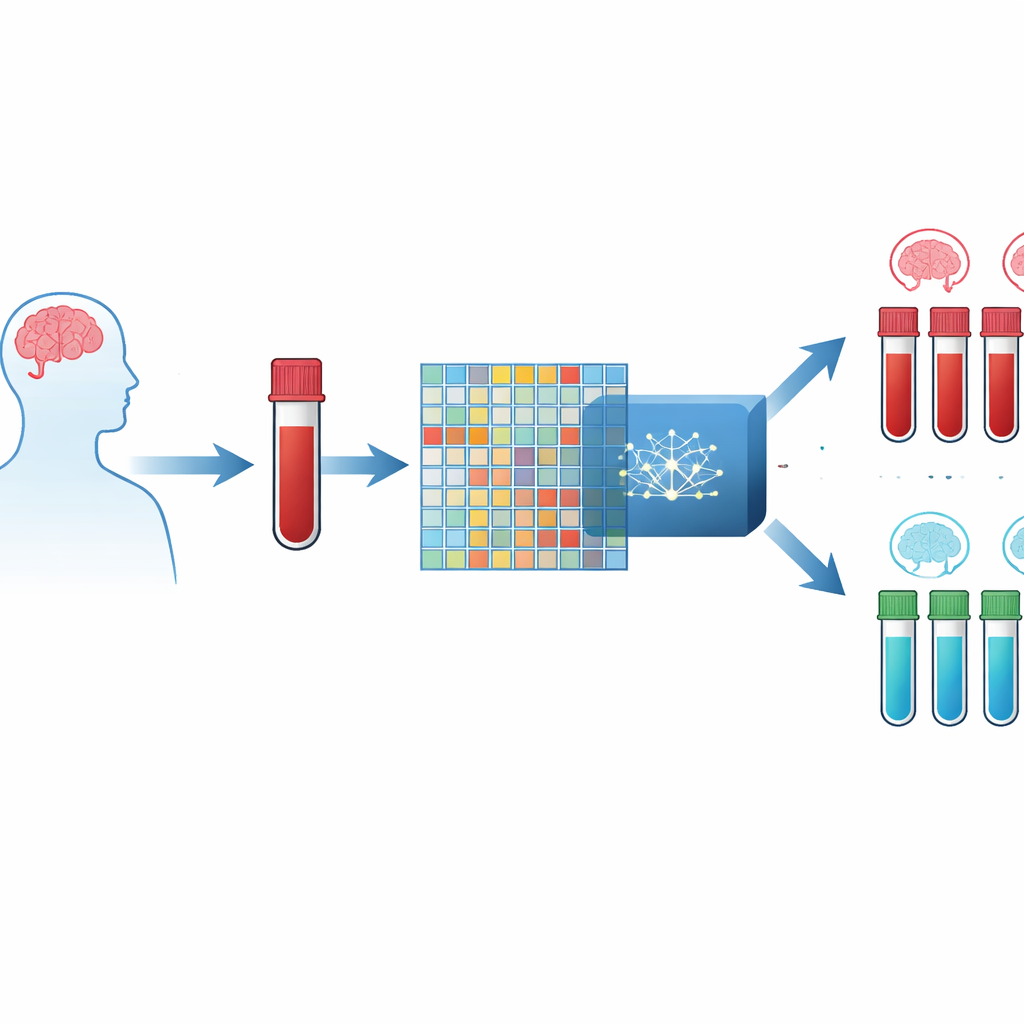
A blood test instead of a brain scan
The authors focus on tiny changes in how genes are switched on or off in blood cells. Modern lab chips can measure the activity of thousands of genes at once, producing a massive table of numbers for each person. The challenge is that there are far more gene measurements than patients, which can easily mislead computer models. To get around this, the researchers combined three large public datasets of blood samples from people with Alzheimer’s and from healthy volunteers, creating an integrated resource with over twelve thousand shared genes measured in hundreds of individuals.
Teaching computers to pick out key warning signs
Instead of asking an algorithm to digest all twelve thousand genes, the team first taught it to select a much smaller set of especially informative ones. They compared several ways of doing this, including simple statistical tests, methods that remove less useful genes step by step, and approaches that build the selection directly into the model. These “feature selection” tools narrowed the list to hundreds or a little over a thousand genes that best distinguished patients from healthy controls. The reduced gene sets helped keep the models from memorizing noise and improved their performance on unseen data.
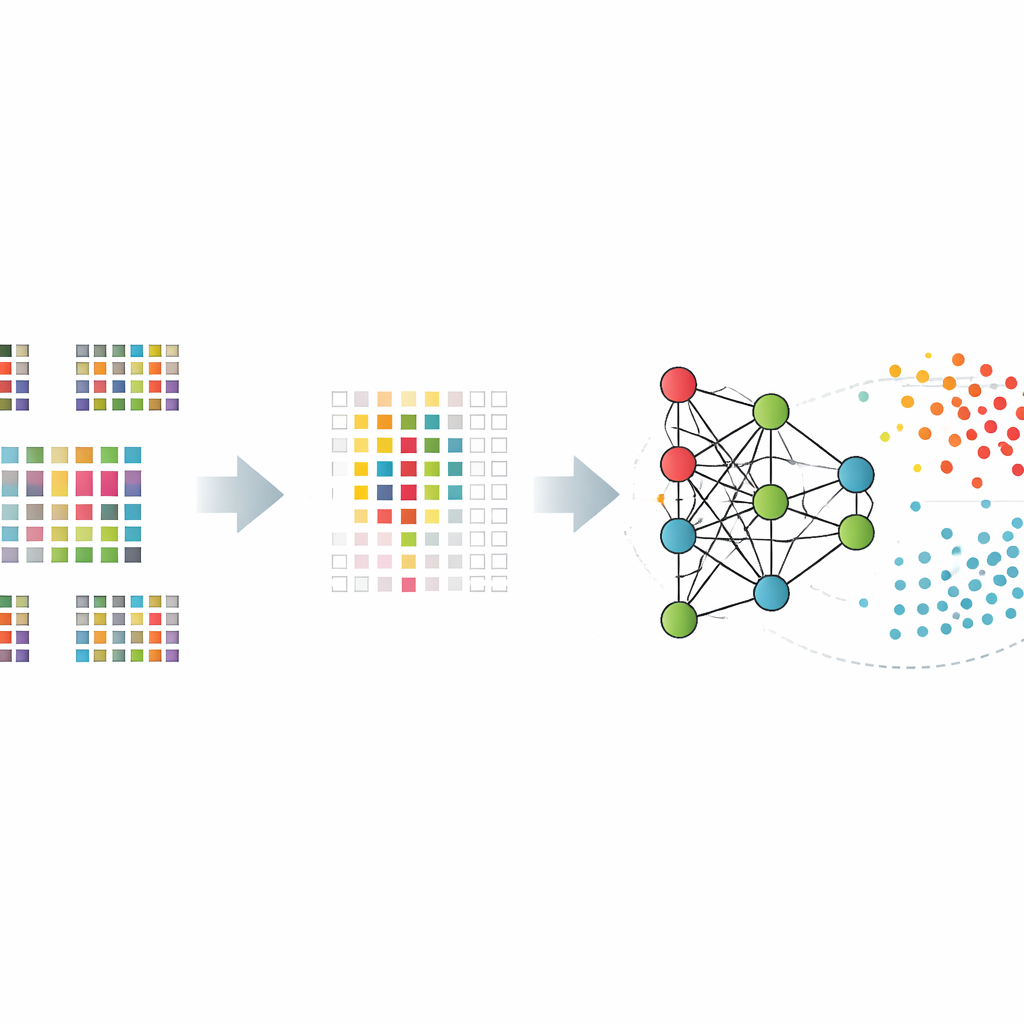
Making sense of a black box
To avoid blind trust in a black-box prediction, the researchers used explainable artificial intelligence techniques to understand which genes mattered most and how they influenced each decision. A method called SHAP, borrowed from game theory, scores each gene’s contribution to the final outcome for each person. By applying it to their best-performing models, the authors highlighted a core group of genes whose activity patterns consistently tipped the scales toward an Alzheimer’s or healthy classification. Many of these genes have already been linked to brain health or immune function, lending biological credibility to the model’s inner workings.
Boosting power with synthetic patients
Even after merging datasets, the number of real blood samples remained modest. To strengthen their models, the authors trained a specialized type of neural network, known as a generative adversarial network, to create realistic synthetic gene profiles that resemble those of actual patients. These artificial samples were added only to the training data, never to the test data, so that performance checks stayed honest. With this augmented training pool and carefully chosen genes, a deep neural network was able to identify Alzheimer’s cases with about 91% overall accuracy and 95% precision, meaning very few healthy people were incorrectly flagged as having the disease.
What the findings mean for patients
This work suggests that a future blood-based test for Alzheimer’s, powered by smart algorithms that both select and explain key gene signals, could complement or even reduce reliance on costly scans and invasive procedures. While more validation is needed on independent groups of patients, and differences between lab methods must be better controlled, the study shows that combining multiple datasets, trimming away unhelpful information, and opening up the “black box” of AI can bring us closer to a practical, interpretable blood test for earlier and more comfortable Alzheimer’s detection.
Citation: Hariharan, J., Jothi, R. Alzheimer’s disease prediction using deep learning and XAI based interpretable feature selection from blood gene expression data. Sci Rep 16, 8022 (2026). https://doi.org/10.1038/s41598-026-35260-8
Keywords: Alzheimer’s diagnosis, blood biomarkers, gene expression, deep learning, explainable AI