Clear Sky Science · en
Exploration of potential biomarkers in salivary duct carcinoma based on bioinformatics analysis
Why a rare salivary cancer matters
Most people have never heard of salivary duct carcinoma, yet this rare cancer of the salivary glands is one of the most aggressive head and neck tumors known. Patients often face early spread of disease and limited treatment choices beyond surgery and radiation. This study uses modern gene-scanning and data-mining tools to search for molecular "fingerprints" in these tumors. Finding such biomarkers could open the door to earlier diagnosis and, crucially, new targeted and immune-based therapies for patients who currently have few options.
Hunting for clues in tumor DNA
The researchers began by collecting tumor samples and nearby normal tissue from patients with salivary duct carcinoma in two hospitals in China. They used a high-throughput technique called SNP microarray, which scans many locations across the genome at once, to look for genes that were turned up or down in the cancer compared with normal gland tissue. To strengthen their results, they combined these new data with an existing public dataset of salivary duct carcinoma from an international gene-expression database. By intersecting the two sources, they focused only on genes that were consistently different between cancer and normal tissue in both sets of patients.
Narrowing down to a core set of genes
From dozens of altered genes in their own samples and more than three thousand in the public dataset, the team found 13 genes that overlapped between the two. Using software that maps how proteins interact with one another, they built a network of relationships among these genes and then applied ranking algorithms to pinpoint the ones most central to the network. This process yielded 10 "hub" genes that appear to sit at important control points in salivary duct carcinoma cells. Most were more active in tumor tissue than in healthy salivary gland, suggesting they may help drive cancer behavior, while one gene, COL11A1, was less active in tumors than in normal tissue.
What the key genes might be doing
To understand what these genes actually influence, the scientists ran enrichment analyses, which group genes by the cellular jobs they perform. The core genes clustered around processes such as moving calcium ions in and out of cells, powering molecular pumps using ATP, and working through transporter molecules that shuttle substances across cell membranes. These functions are deeply tied to how cells grow, move, and respond to their surroundings—processes that often go awry in cancer. One gene in particular, PIK3R5, stood out because it belongs to a well-known immune and growth signaling pathway and was flagged as both a core gene and an immune-related gene, hinting that it might link tumor behavior to the body’s immune response.
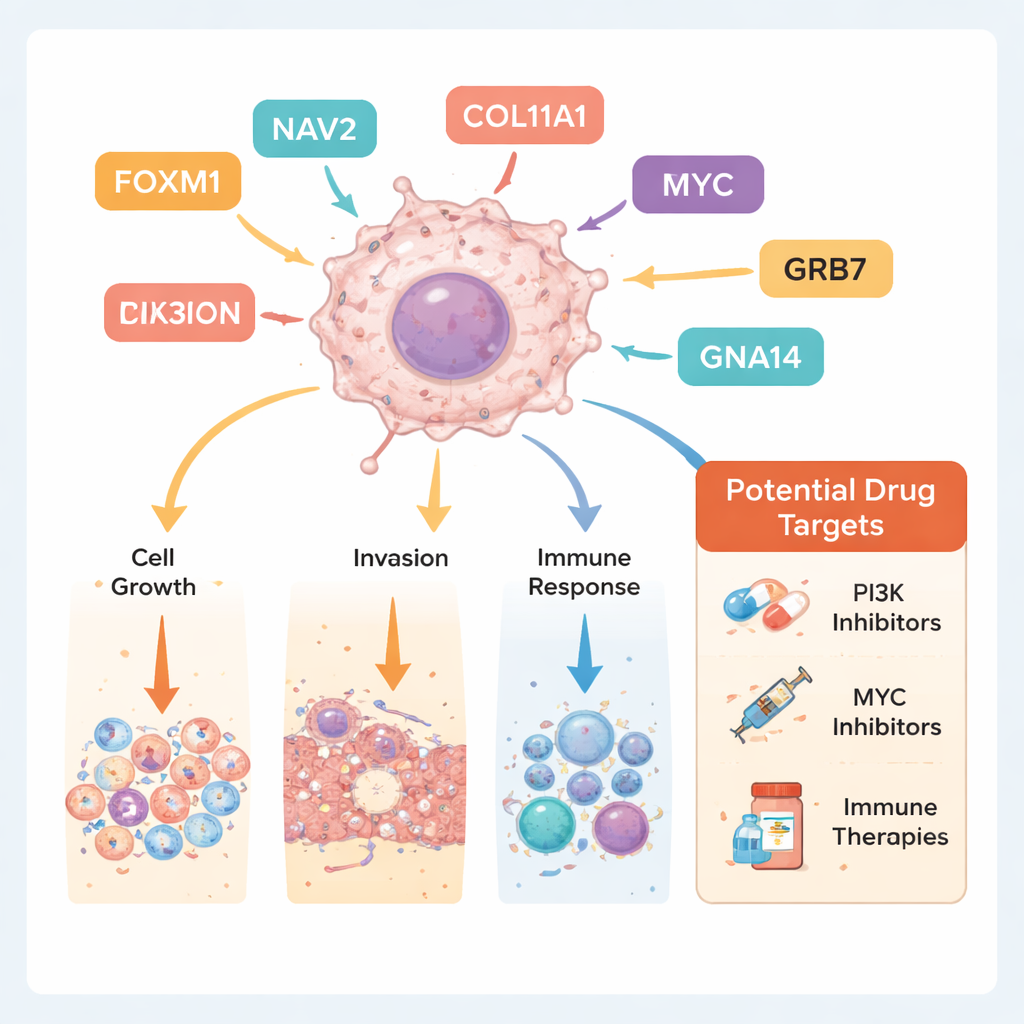
Connecting genes to many cancers and real tissue samples
The team then checked how their 10 hub genes behave across 34 different cancer types using a large-scale cancer database. Many of the genes, including FOXM1 and NAV2, were also more active in other tumors such as breast, colon, and liver cancers, suggesting that salivary duct carcinoma shares molecular features with more common cancers. Finally, they confirmed the altered activity of several genes directly in patients’ tissues using immunohistochemistry, a staining method that makes specific proteins visible under the microscope. Tumor samples showed stronger signals for FOXM1, NAV2, and LILRA2, and weaker signals for COL11A1, supporting the computer-based findings.
What this means for future treatment
Taken together, the work shows that salivary duct carcinoma has a distinct pattern of gene activity involving cell growth control, cell movement, and immune-related pathways. The highlighted genes—especially FOXM1, NAV2, COL11A1, and PIK3R5—could serve as biomarkers to help diagnose or classify this cancer and may eventually guide targeted drugs or immune therapies. While more laboratory and clinical studies are needed, these molecular signposts provide a crucial starting map for turning a poorly understood, hard-to-treat disease into one where therapies can be tailored to the tumor’s inner wiring.
Citation: Zhang, R., Zhu, X., Ma, H. et al. Exploration of potential biomarkers in salivary duct carcinoma based on bioinformatics analysis. Sci Rep 16, 5525 (2026). https://doi.org/10.1038/s41598-026-35239-5
Keywords: salivary duct carcinoma, cancer biomarkers, gene expression, targeted therapy, immunotherapy