Clear Sky Science · en
White blood cell classification using custom deep neural network and visualizing features of the images using heatmaps
Why Smarter Blood Tests Matter
A routine blood test can reveal early signs of infection, allergy, or even cancer—but today, much of that insight still depends on experts carefully inspecting cells under a microscope. This process is slow, expensive, and not always available in smaller clinics or rural hospitals. The paper described here presents a compact artificial intelligence system that can accurately recognize different kinds of white blood cells from microscope images, potentially bringing faster and more reliable blood analysis to many more patients.
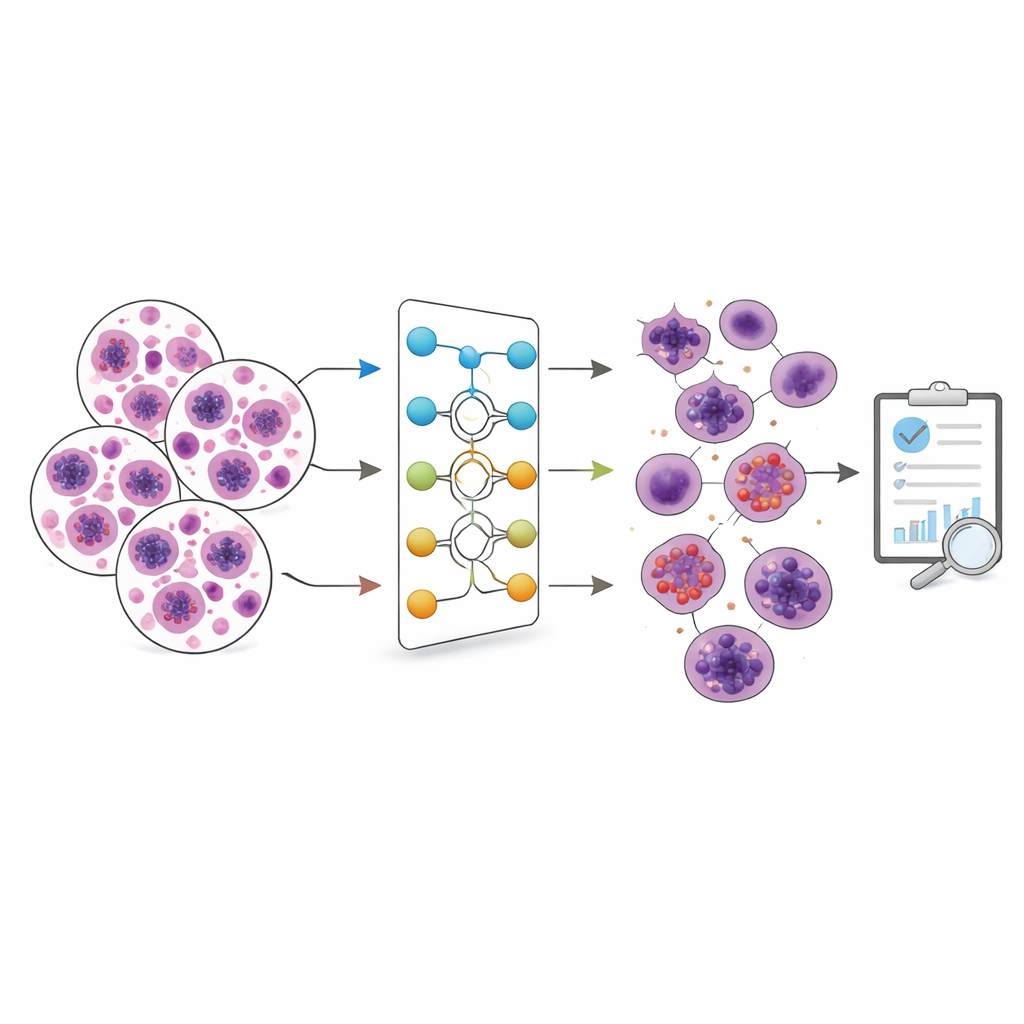
The Body’s Tiny Defenders
White blood cells are the body’s front-line defenders against germs and other threats. They come in several main varieties, each with a different job: some attack bacteria, others fight parasites, respond to allergies, or help coordinate long-term immunity. Doctors often look at both the number and type of these cells to diagnose illnesses and monitor treatments. Today, this is commonly done either by manual counting under a microscope—a skilled but time-consuming task—or by using large automated machines that many smaller laboratories cannot afford.
From Manual Counting to Digital Eyes
Over the last decade, researchers have turned to computer vision and machine learning to help automate white blood cell identification. Basic programs can measure shapes and colors, while more advanced systems use deep learning, which can learn complex patterns directly from images. However, many powerful deep learning models are huge, needing substantial computing power and memory, and they can suffer from training problems such as “vanishing” signals inside the network. This limits their use in small clinics, mobile health devices, or hospitals with modest computing resources.
A Lean yet Powerful Digital Microscope
The authors introduce a streamlined deep learning model they call a custom deep neural network, or CDNN. It is designed specifically to recognize white blood cells in microscope images. The model was trained and tested on two popular datasets of blood cell images: one large and somewhat imbalanced set (Raabin WBC) with five types of white blood cells, and another smaller but evenly balanced set (BCCD) with four types. Before training, all images were resized, their brightness levels normalized, and then lightly altered—rotated, reflected, and sheared—to mimic natural variation and reduce overfitting, which happens when a model memorizes training examples instead of learning general patterns.
Inside the Model’s “Thought Process”
CDNN is built from a series of building blocks that help it learn effectively while staying compact. These blocks include “residual” connections that allow information to bypass some layers, preventing the training signals from fading as they pass through the network. The model gradually increases the number of internal filters as images move through it, capturing both fine details and broader shapes of cell nuclei and surrounding material. Despite having far fewer adjustable settings than well-known image models such as VGG16 or ResNet-50, CDNN achieves very high accuracy: about 98% on the Raabin dataset and nearly 99.6% on the BCCD dataset, outperforming many larger state-of-the-art methods.
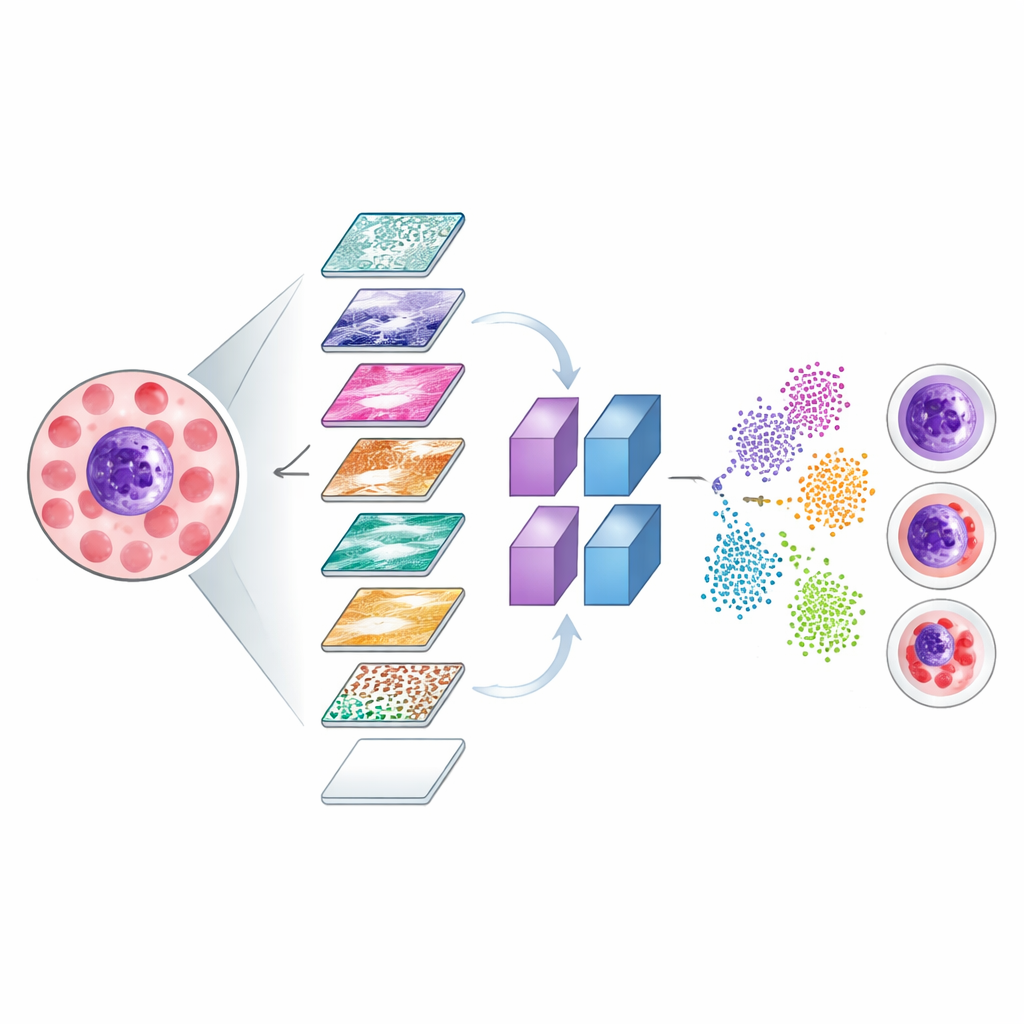
Making Invisible Decisions Visible
To ensure that the system focuses on medically meaningful parts of the images, the researchers used two visualization tools, known as Grad-CAM and LIME. These methods generate color heatmaps that highlight which regions of a cell image most strongly influenced the model’s decisions. The bright zones in these maps tended to align with crucial structures such as the cell’s nucleus and surrounding cytoplasm, rather than irrelevant background. The team also explored how the network internally separates different cell types by projecting its internal signals into a two-dimensional map, where correctly classified cells form clear clusters and misclassified cases can be inspected to understand where the model struggles.
What This Could Mean for Patients
In simple terms, this work shows that a carefully designed, relatively small AI model can sort white blood cells from microscope images with accuracy on par with or better than many large, complex systems. Because the model is compact and efficient, it is more suitable for use in smaller laboratories, point-of-care devices, or even portable equipment, potentially bringing high-quality blood analysis closer to patients. While further testing in real clinical settings is still needed, this approach points toward faster, more accessible, and more transparent tools that can assist pathologists in diagnosing disease and catching problems earlier.
Citation: Karaddi, S.H., Bitra, H., Bairaboina, S.S.R. et al. White blood cell classification using custom deep neural network and visualizing features of the images using heatmaps. Sci Rep 16, 9311 (2026). https://doi.org/10.1038/s41598-026-35138-9
Keywords: white blood cells, medical imaging, deep learning, blood diagnostics, neural networks