Clear Sky Science · en
A method for magnetic resonance splenomegaly image segmentation based on large-kernel multi-scale attention mechanism
Why doctors care about an enlarged spleen
The spleen is a fist‑sized organ tucked under the left rib cage, and it quietly filters blood, fights infections, and manages some blood cells. When it becomes enlarged — a condition called splenomegaly — it can signal serious problems, from liver disease to blood cancers. Modern hospital scanners can capture detailed images of the spleen, but turning those images into reliable measurements still often depends on time‑consuming, error‑prone manual work by specialists. This study presents a new artificial‑intelligence method that automatically outlines enlarged spleens in MRI scans with very high accuracy, potentially giving doctors a faster and more precise tool for diagnosis and monitoring.
The challenge of seeing the spleen clearly
On MRI images, the spleen does not stand out as sharply as many people might expect: its gray tone often looks very similar to nearby organs and tissues. To make matters harder, spleens differ dramatically in size and shape from person to person, especially when they are enlarged by disease. Some patients have only mildly increased spleen volume, while others show organs that are many times the normal size. Collecting high‑quality images of these extreme cases is also difficult in practice, so researchers must work with relatively small datasets. All of this means that traditional computer programs, and even earlier deep‑learning methods, struggle to draw clean, accurate borders around the spleen on MRI slices.
A smarter network for tricky medical images
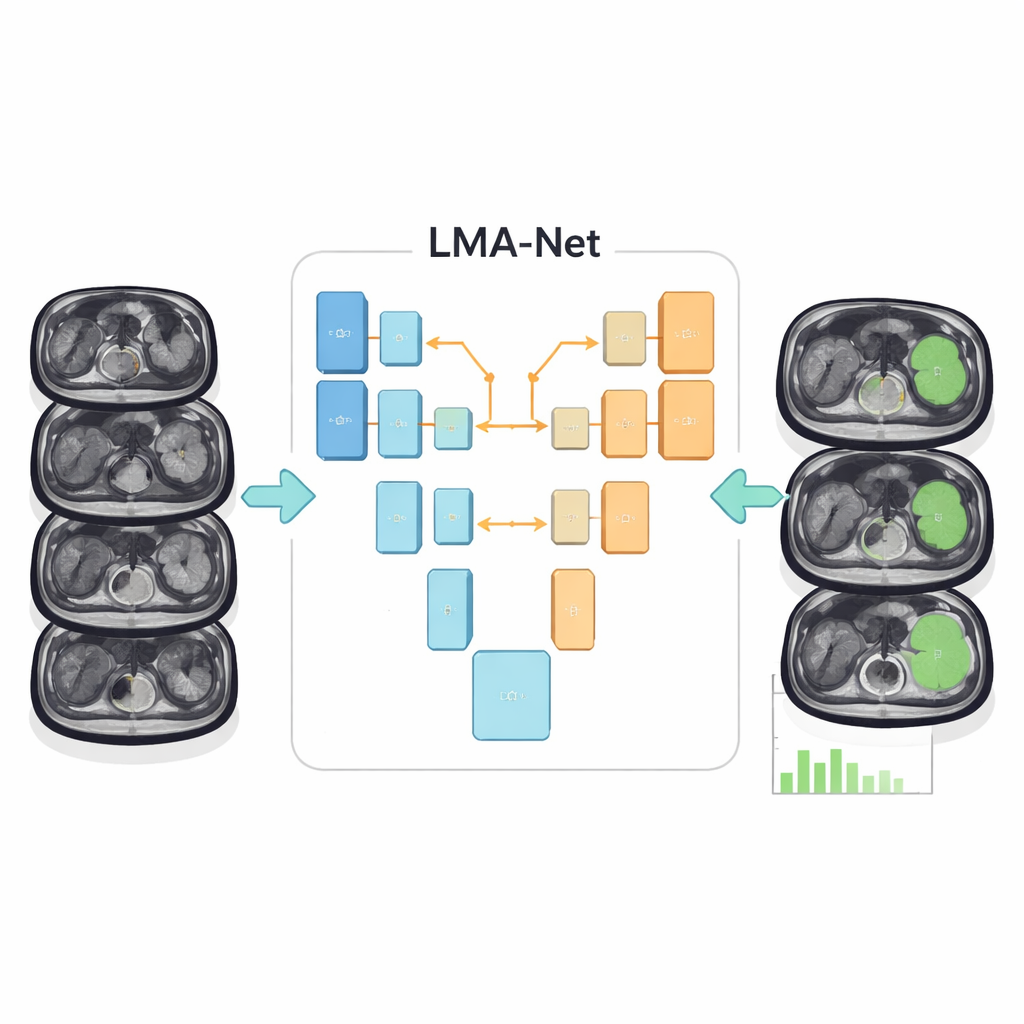
The authors introduce a new deep‑learning architecture called LMA‑Net (Large‑kernel Multi‑scale Attention Net) designed specifically for this problem. It follows a U‑shaped layout that has become standard in medical image analysis: one side of the “U” gradually compresses the image into abstract features (the encoder), while the other side reconstructs a detailed segmentation map (the decoder). LMA‑Net uses a hybrid encoder that combines two powerful ideas. First, a conventional ResNet‑50 convolutional network picks up fine‑grained local details. Then, a Transformer module, borrowed from modern language and vision models, captures broader patterns across the entire image so the algorithm develops a global sense of where the spleen sits and how it tends to look.
Learning to focus on the right details
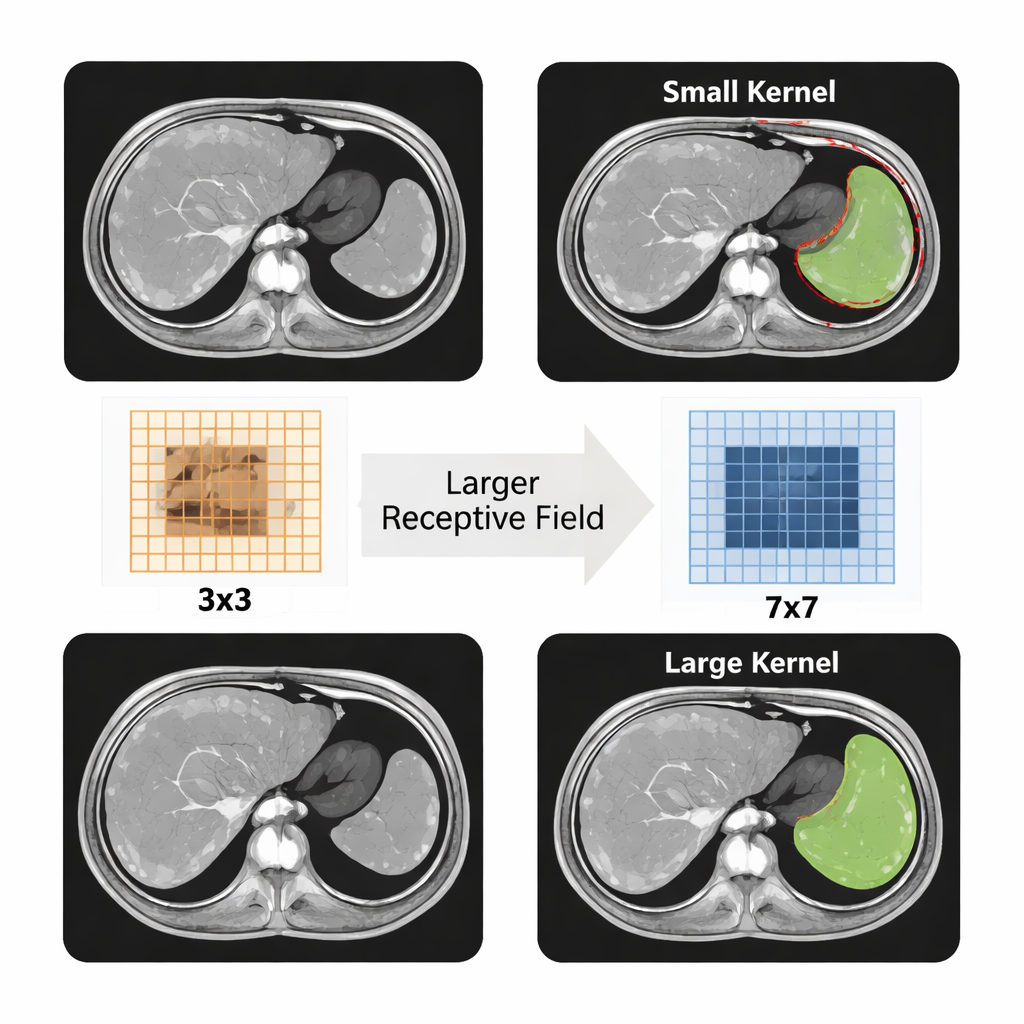
Between the encoder and decoder, LMA‑Net adds a specialized attention block that looks at the image at several scales at once. It uses unusually large convolutional filters, along with an efficient grouping strategy, to expand its field of view without becoming too slow or heavy. These large filters help the network see the whole outline of the spleen rather than just tiny patches, which is crucial when boundaries are fuzzy. The model then learns to assign higher weights to the most informative channels and locations, effectively “paying attention” to the regions and textures that most likely belong to the spleen. In the decoder, a lightweight fusion module and a boundary‑refinement block further sharpen the organ’s edges, aiming for smooth, realistic contours while keeping the computation modest enough for clinical use.
How well the system works in practice
To test their approach, the researchers trained and evaluated LMA‑Net on two different collections of medical images. One dataset contained MRI scans from 51 patients with varying degrees of splenomegaly, with precise outlines drawn by expert radiologists. The other came from the public Medical Segmentation Decathlon and consisted of CT scans focused on the spleen. Using widely accepted accuracy measures that compare the overlap between predicted and expert‑drawn regions, LMA‑Net outperformed several popular segmentation networks, including U‑Net and newer attention‑based and Transformer‑based models. On the MRI splenomegaly data, it correctly overlapped with the expert labels for more than 96% of the spleen area on average, a noticeable improvement over competing methods.
What this could mean for patients and clinics
For non‑specialists, the key takeaway is that this new AI method can automatically and very precisely outline enlarged spleens on routine MRI scans, even when the organ’s shape is unusual or its edges are hard to see. That means doctors could obtain accurate spleen volumes and shapes more quickly, track changes over time, and better evaluate how patients respond to treatments for liver disease, blood disorders, or cancers that affect the spleen. While further validation and integration into hospital systems are still needed, LMA‑Net points toward a future in which detailed, quantitative measurements from medical images become a standard, automated part of care rather than a manual chore.
Citation: Fu, Z., Zhao, C., Yan, H. et al. A method for magnetic resonance splenomegaly image segmentation based on large-kernel multi-scale attention mechanism. Sci Rep 16, 5227 (2026). https://doi.org/10.1038/s41598-026-35122-3
Keywords: splenomegaly, MRI segmentation, deep learning, medical imaging, attention networks