Clear Sky Science · en
Compact deep learning models for colon histopathology focusing performance and generalization challenges
Why this research matters for patients and doctors
Colon cancer is one of the world’s deadliest cancers, yet diagnosing it still depends on specialists carefully inspecting microscope images of tissue, a task that is slow and prone to disagreement. This study explores whether very small, efficient artificial intelligence (AI) models can help flag cancerous colon tissue accurately enough to be useful in everyday clinics, including those with limited computing power. It also uncovers a hidden weakness: models that look almost perfect during development can still stumble badly on new, real‑world data.
Teaching computers to read microscope pictures
When a colon biopsy is taken, pathologists examine thin, stained slices of tissue under a microscope. Cancerous tissue shows distorted glands, irregular cell shapes, and invasion into surrounding structures, while healthy tissue has orderly, regular patterns. The authors used a public collection of 24,000 digital images of such slices, evenly split between cancer (colon adenocarcinoma) and benign tissue. They resized all images to a standard small format and applied realistic tweaks—small rotations, flips, zooms, and gentle color changes—to mimic the natural variation in how slides are cut, stained, and scanned. This careful preparation helps AI models focus on meaningful tissue patterns instead of superficial details like exact orientation or brightness.
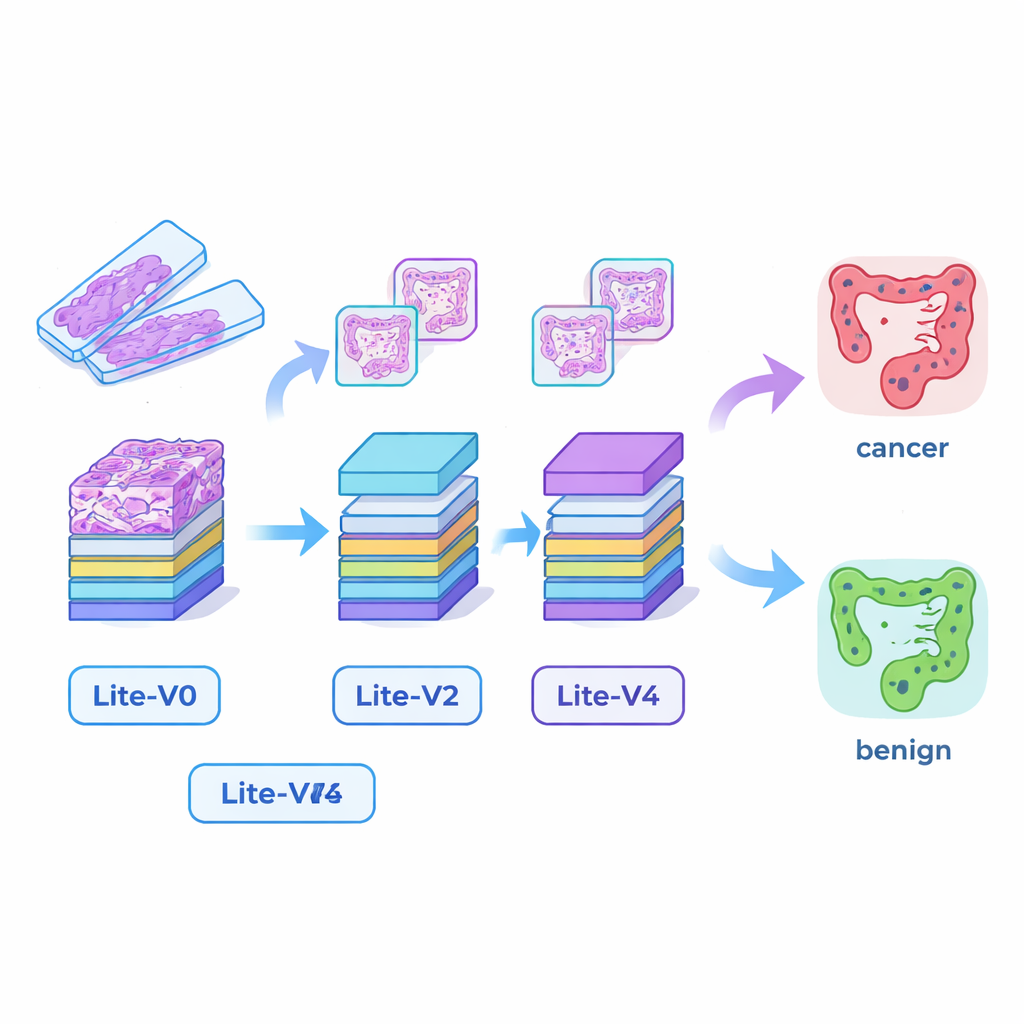
Building tiny but capable AI “eyes”
Many successful medical AI systems rely on very large deep‑learning models that demand powerful graphics cards and lots of memory, making them hard to deploy in smaller hospitals or at the bedside. To bridge this gap, the researchers designed four compact convolutional neural networks—Lite‑V0, Lite‑V1, Lite‑V2, and Lite‑V4. Each one looks at the same input image patches, but they differ in how many layers and filters they use to detect visual features such as edges, textures, and gland shapes. All four share a simple, transparent design: repeated blocks of standard convolution, normalization, and pooling, followed by a small “decision head” that outputs the probability of cancer or benign tissue. The goal was to see how much accuracy could be squeezed out of models small enough to fit comfortably on basic clinical hardware.
Impressive scores inside the lab
The team trained and compared all four models on a fixed split of the dataset, using widely accepted measures: accuracy, a balanced F1‑score that weighs errors in both classes equally, confusion matrices, and diagnostic plots such as ROC and precision–recall curves. A mid‑sized model, Lite‑V2, emerged as the star performer. Despite being only about 1.5 megabytes in size and having roughly 128,000 trainable parameters, it achieved almost flawless performance on the internal validation set, with a macro F1‑score around 0.999 and near‑perfect sensitivity and specificity. In other words, within this carefully prepared environment, Lite‑V2 could almost always distinguish cancerous from benign colon tissue, while remaining fast and light enough for use on modest computers.
When real‑world variation breaks the spell
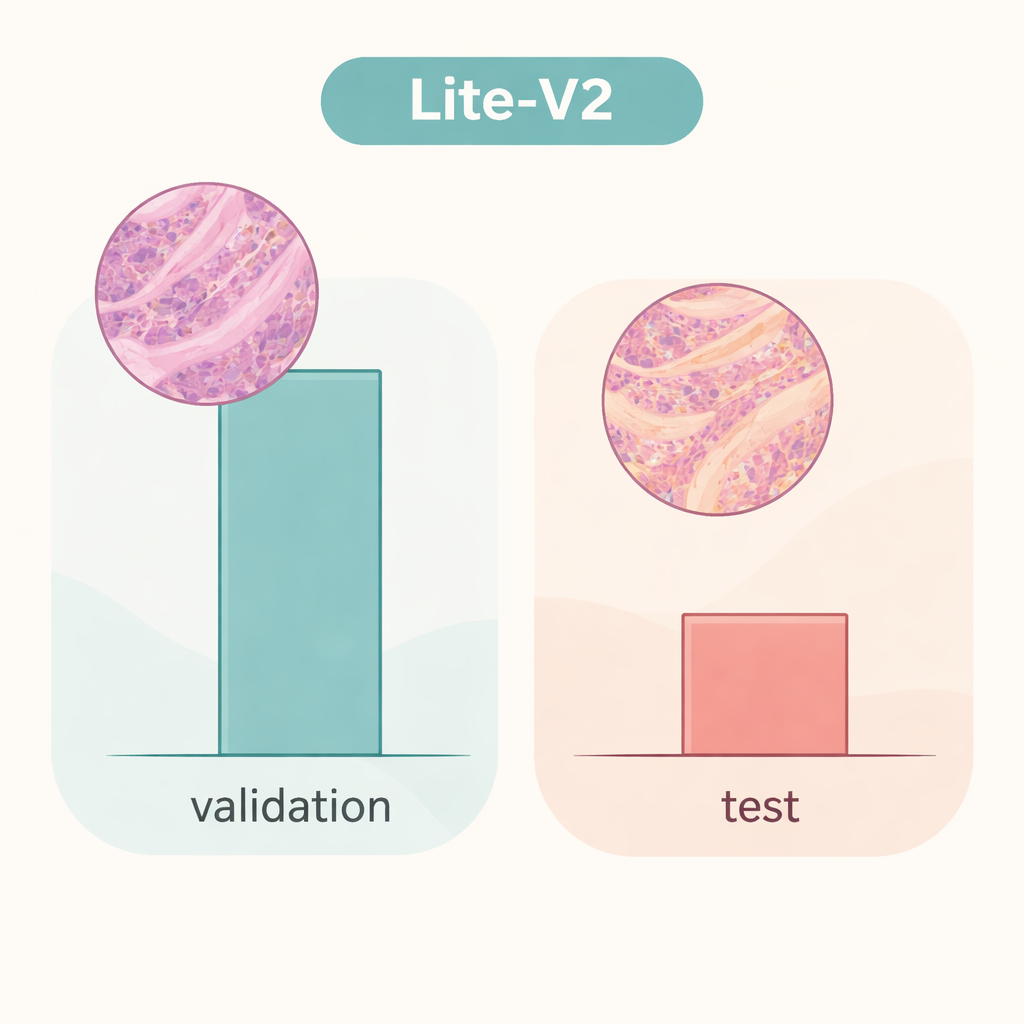
However, the story changes dramatically when the same Lite‑V2 model is tested on an independent set of images that differ subtly in ways that mimic slides from another laboratory—what researchers call a “domain shift.” On this unseen test set, overall accuracy dropped to about 50%, and the balanced F1‑score fell to roughly 0.33. The model continued to recognize many cancer samples but struggled badly with benign tissue, mislabeling a large fraction as malignant. This shows that the network had learned details that were tightly tied to the original data source—such as staining style or scanner characteristics—rather than robust, portable signatures of disease. The work highlights that glowing results on internal validation can give a false sense of security if models are not challenged with truly different data.
What this means for future AI diagnosis tools
For a general reader, the takeaway is twofold. First, compact AI systems really can reach expert‑level performance on colon tissue images while remaining small and efficient enough for wide deployment, opening the door to faster screening and support for overburdened pathologists. Second, and just as important, a model that looks “perfect” on its home dataset may falter badly when it meets images from a new hospital. The authors argue that future work must focus on making these lightweight models robust to changes in staining, scanners, and patient populations—using strategies like stain‑robust training, domain adaptation, and broader multi‑center datasets. Until then, AI should be seen as a promising assistant rather than a stand‑alone decision‑maker in cancer diagnosis.
Citation: Hanif, F., Raza, A. & Mohammed, H.A. Compact deep learning models for colon histopathology focusing performance and generalization challenges. Sci Rep 16, 5489 (2026). https://doi.org/10.1038/s41598-026-35119-y
Keywords: colon cancer, histopathology, deep learning, lightweight CNN, domain shift