Clear Sky Science · en
DrugBank mining with machine learning reveals novel candidates for BCL-2 inhibition
Why finding smarter cancer drugs matters
Cancer cells often refuse to die when they should. Many tumors survive by overusing a family of “bodyguard” proteins called BCL-2 that block the cell’s built‑in suicide program. Drugs that target BCL-2 already exist, but they can cause side effects and do not work for every patient. This study explores how modern machine learning can sift through thousands of existing drugs to find new, safer candidates that may disarm BCL-2 and help cancer cells self‑destruct.
How cells choose between life and death
Healthy tissues constantly remove damaged or unneeded cells through a controlled self‑destruct process known as apoptosis, or programmed cell death. A group of proteins called the BCL-2 family acts as a central switch for this decision. Some members push cells toward survival, while others push them toward death. In many cancers, the survival‑promoting members, including BCL-2 and its close relative BCL-XL, are produced in excess. This extra protection allows cancer cells to ignore death signals and resist chemotherapy. Because of this, blocking BCL-2 has become an attractive strategy in cancer treatment, but current drugs often hit related proteins as well, leading to side effects such as dangerous drops in platelet counts.
Teaching computers to recognize promising molecules
Instead of searching for new compounds from scratch, the researchers turned to databases of molecules that have already been studied or used as drugs. They began with a large public resource called ChEMBL, which contains experimental measurements of how strongly different chemicals bind to BCL-2. After carefully cleaning this information—removing duplicates, uncertain measurements, and overly large or unusual molecules—they ended up with 601 well‑characterized compounds. Each molecule was translated into a kind of digital fingerprint capturing its structural features. These fingerprints were used to train and compare seven different machine‑learning models on the task of deciding whether a new molecule is likely to be a strong BCL-2 blocker or essentially inactive. 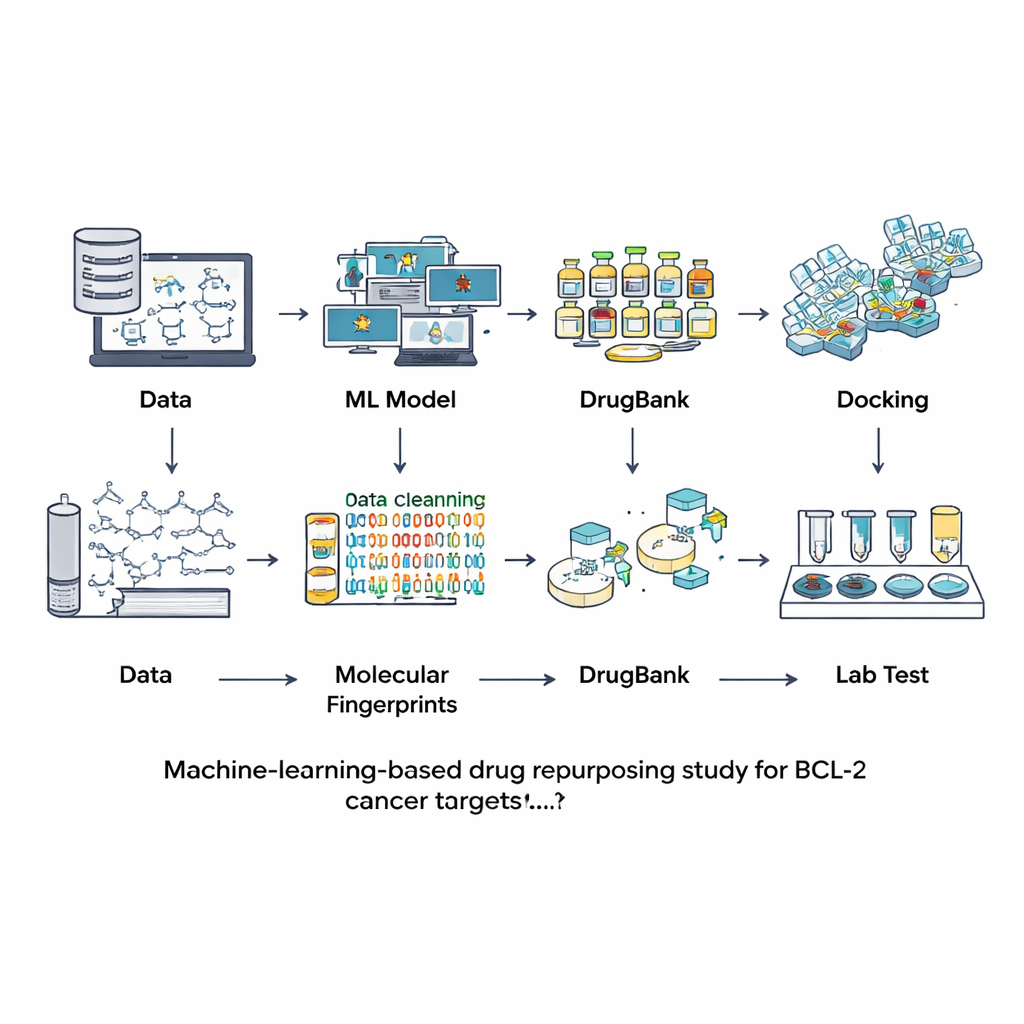
Picking the best model and scanning a drug library
The team evaluated the models using a separate test set that had not been seen during training, checking not only how often each model was right, but also how well it distinguished actives from inactives and how balanced its predictions were. A model called LightGBM—a modern, tree‑based boosting method—performed best across most measures, including overall accuracy and its ability to assign reliable probabilities. With this tuned model in hand, the researchers turned to DrugBank, a curated collection of more than 12,000 approved, experimental, and withdrawn drugs. After computing the same type of fingerprints, they asked LightGBM which of these molecules looked like potential BCL-2 inhibitors. Only nine compounds scored highly, roughly the top one‑tenth of one percent of the entire library, showing that the virtual screen was very selective. Four of the nine were already known BCL-2 inhibitors, reassuring the team that the approach was sound.
From computer hits to molecular interactions
Among the remaining high‑scoring molecules, the researchers focused on three not previously linked to BCL-2: Dersalazine, Opelconazole, and Zongertinib. To see whether these candidates could plausibly fit into the BCL-2 binding pocket, they used computer docking, a technique that predicts how a small molecule might nestle into the protein’s surface. The simulations suggested that Opelconazole and Zongertinib, in particular, form networks of favorable contacts with the same key amino acids that grip a well‑studied reference drug, ABT-737. Their predicted binding strengths were close to those of established inhibitors, hinting that the machine‑learning model had indeed uncovered molecules capable of disarming BCL-2. 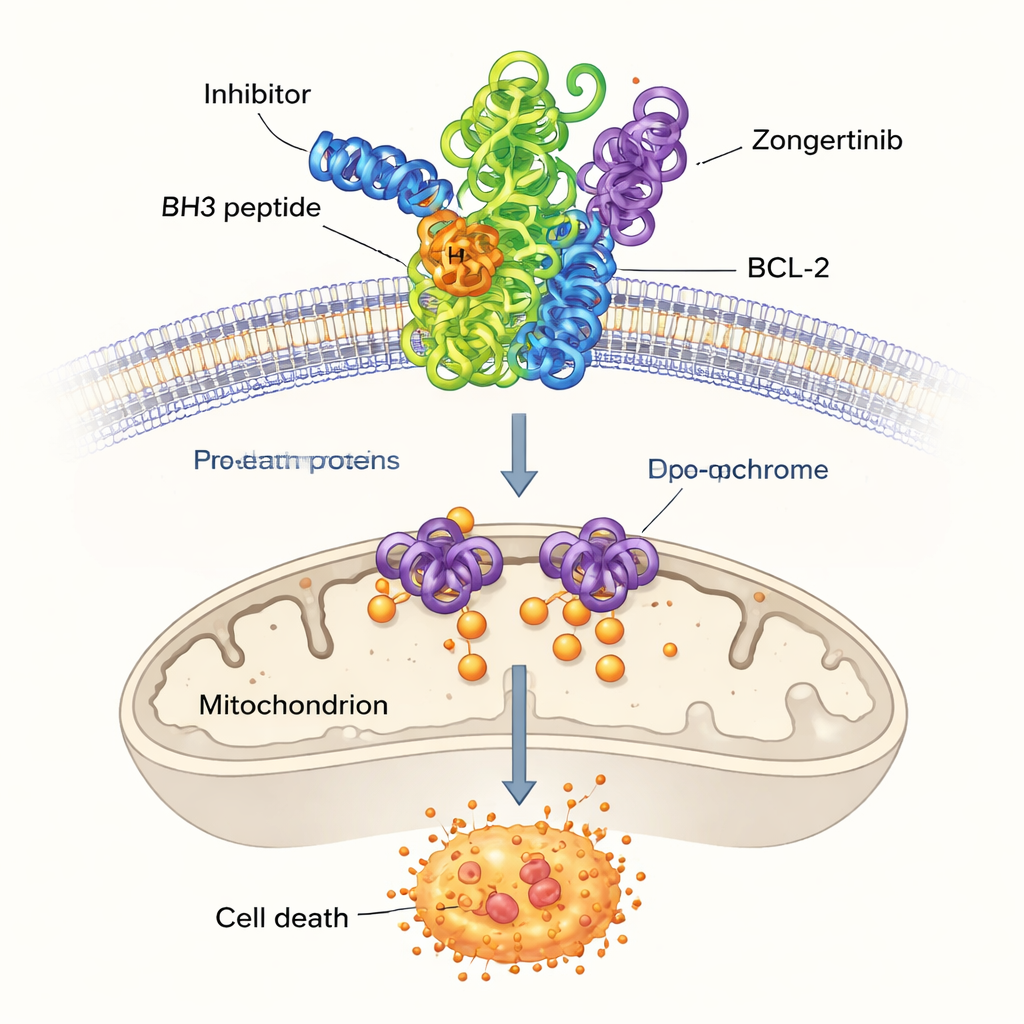
Putting predictions to the test in the lab
Computational hints are useful only if they hold up under real‑world conditions. The team therefore tested the three candidates in a biochemical assay that measures how well a compound can prevent BCL-2 from binding to one of its natural partners. At a range of concentrations, Dersalazine showed little effect. Opelconazole and Zongertinib, however, both reduced BCL-2 activity at high doses, with Opelconazole nearly shutting down the signal. While these concentrations are higher than what would be ideal for a clinic‑ready drug, they demonstrate that the candidates truly interact with BCL-2 and validate the overall discovery pipeline.
What this means for future cancer therapies
To a non‑specialist, the key message is that the researchers successfully trained a computer system to recognize what a BCL-2‑blocking molecule “looks like,” then used it to mine a large library of existing drugs and drug‑like compounds. The approach rediscovered known BCL-2 drugs and highlighted new candidates, two of which showed real inhibitory activity in lab tests. Although much work remains—improving potency, understanding safety, and testing in cells and animals—this study shows how machine learning and smart data curation can speed up the hunt for better cancer medicines by recycling and re‑evaluating compounds we already know.
Citation: Park, J., Cho, S., Lee, H. et al. DrugBank mining with machine learning reveals novel candidates for BCL-2 inhibition. Sci Rep 16, 5482 (2026). https://doi.org/10.1038/s41598-026-35117-0
Keywords: BCL-2 inhibitors, machine learning, drug repurposing, apoptosis, cancer therapy