Clear Sky Science · en
Root-associated protein prediction using a protein large language model and hypergraph convolutional networks
Why roots and their hidden helpers matter
When we think about keeping crops healthy, we usually picture leaves and fruits. But much of a plant’s success happens out of sight, in the soil. There, special root-associated proteins help plants absorb water and nutrients and cope with stresses like drought or poor soil. Finding these crucial proteins with lab experiments alone is slow and expensive. This study introduces a powerful computer model, called Hypergraph-Root, that can quickly scan protein sequences and predict which ones are likely to be root-associated, offering a faster route to hardier crops and better harvests.
Hidden workhorses in the soil
Plant roots do more than anchor a plant in place. They constantly sense their surroundings, pull in minerals, and communicate with soil microbes. Root-associated proteins are central to all of this, shaping how roots grow, how they respond to heat, drought, or nutrient shortages, and how they interact with helpful microbes. Because these proteins strongly influence yield and resilience, farmers and breeders care about them even if they never see them directly. Yet many such proteins remain undiscovered, largely because traditional methods—like proteomics and gene expression studies—require costly instruments, complex analyses, and painstaking experiments.
Turning protein sequences into clues
Proteins are built from strings of amino acids, and patterns in those strings often reveal where a protein works in the plant and what it does. Previous computer models tried to exploit these patterns to spot root-associated proteins, but they topped out at accuracies below 80 percent. One problem is that they treated relationships between amino acids in a fairly simple way, usually as pairs. Another is that they relied on limited types of features extracted from sequences. The authors reasoned that richer representations of each protein, together with smarter ways of modeling amino acid relationships, could uncover subtler patterns linked to root functions.
Borrowing tricks from language and networks
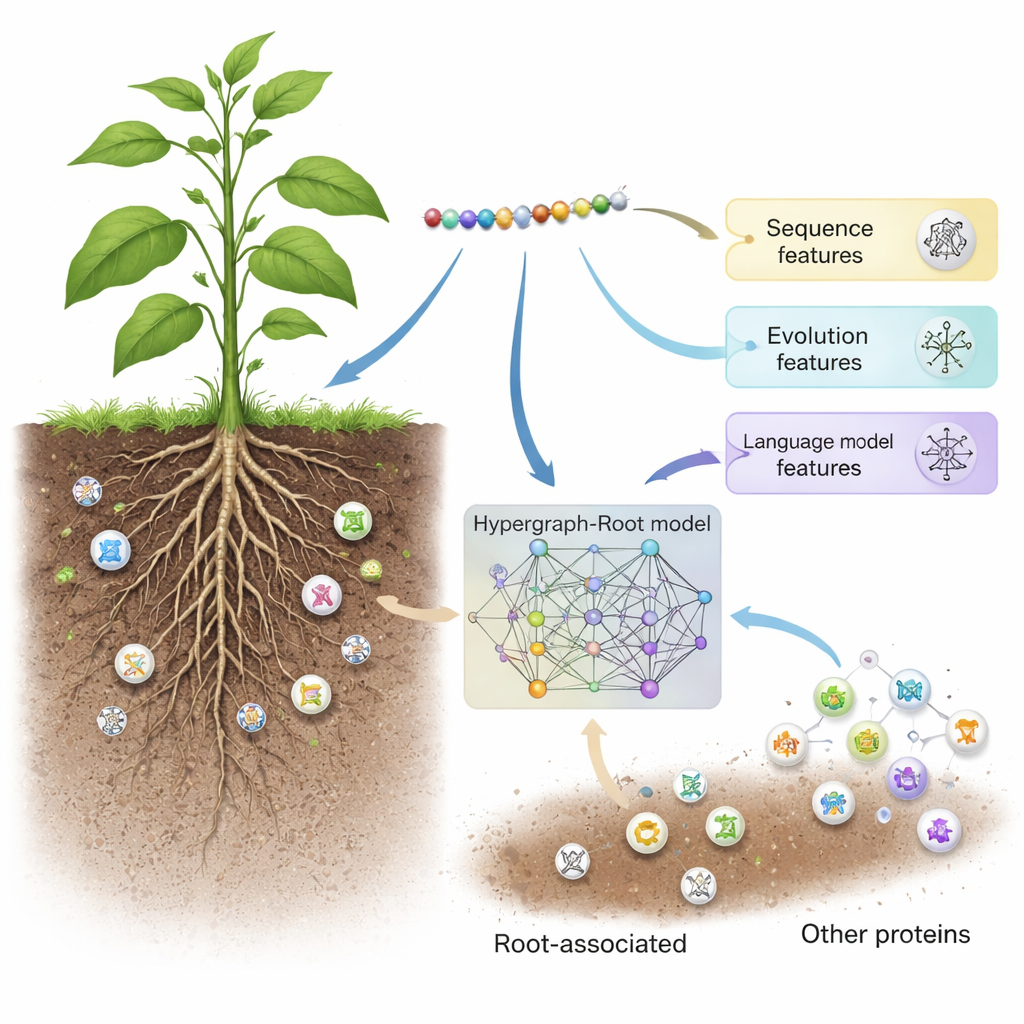
Hypergraph-Root starts by describing each protein in three complementary ways. It uses traditional sequence scoring schemes (BLOSUM62 and position-specific scoring matrices) that capture how amino acids tend to substitute for each other over evolution. It then adds a third, more modern description from a protein language model called ProtT5—software trained on millions of protein sequences, much like a text-prediction engine is trained on human language. ProtT5 produces a rich numerical “embedding” for each amino acid that encodes structural and functional hints. Together, these three views give a detailed fingerprint of every protein in the study.
Mapping complex connections inside proteins
To go beyond simple pairwise comparisons, the researchers predicted how close amino acids are in a protein’s 3D structure and used this information to build a hypergraph—a network in which a single connection can link more than two amino acids at once. A specialized neural network, the hypergraph convolutional network, processes this structure-aware network and refines the protein fingerprints into higher-level features. A multi-head attention module then learns which parts of the protein carry the most useful signals for deciding whether it is root-associated. Finally, a standard classifier turns these distilled features into a probability score: root-associated or not. Across many training runs and both balanced and imbalanced test sets, Hypergraph-Root reached accuracies above 83 percent and an area under the ROC curve (AUC) around 0.9, clearly outperforming earlier models.
What the model reveals and why it matters
Beyond raw accuracy, the model provided insights into which information matters most. Features from the ProtT5 language model contributed more than traditional sequence and evolutionary features, suggesting that large, pre-trained models can capture subtle biological signals that older methods miss. The hypergraph component also proved important: removing it or replacing it with a simpler graph model reduced performance. When the researchers applied Hypergraph-Root to proteins not previously labeled as root-associated, it highlighted a handful whose known functions—such as membrane transport and protein tagging in roots—strongly suggest they do play roles in root biology. These candidates now give experimental biologists clear shortlists to test in the lab.
From smart predictions to stronger crops
In everyday terms, Hypergraph-Root is like an expert librarian for plant biology: given only the “letters” of a protein, it estimates whether that protein likely works in the roots. By combining language-model insights, evolutionary history, and complex structural relationships, it greatly improves on past prediction tools. While it does not replace experiments, it can narrow down thousands of possibilities to a manageable few, saving time and money. In the long run, such models could speed the discovery of root-associated proteins that help crops survive heat, drought, or poor soils—an important step toward more resilient agriculture in a changing climate.
Citation: Chen, L., Xun, X. & Zhou, B. Root-associated protein prediction using a protein large language model and hypergraph convolutional networks. Sci Rep 16, 4876 (2026). https://doi.org/10.1038/s41598-026-35110-7
Keywords: root-associated proteins, plant bioinformatics, deep learning, protein language models, crop resilience