Clear Sky Science · en
Transcriptomic characterization of the aberrant alternative splicing in skeletal muscles of sarcopenia patients
Why aging muscles lose their strength
Sarcopenia—the gradual loss of muscle mass and strength with age—is one of the main reasons older adults struggle with everyday tasks, from climbing stairs to carrying groceries. This study dives inside aging muscle at the level of RNA, the messages that tell cells which proteins to make. The authors show that in sarcopenia, many of these RNA messages are cut and stitched together in abnormal ways, potentially disturbing energy production in muscle and pointing to new treatment targets.
Muscle loss and the body’s gene “editing” system
Our genes are written in DNA, but cells work from RNA copies of those genes. Before an RNA message is used, cells normally trim and splice it—much like editing a film—so that one gene can produce several versions of a protein. This process, called alternative splicing, is especially active in muscle, which needs finely tuned proteins to contract, repair, and adapt to exercise. When splicing goes wrong, the wrong protein versions can be produced or important ones can be lost, contributing to diseases such as muscular dystrophy and some brain disorders. The authors suspected a similar hidden layer of RNA mis-editing might be driving sarcopenia.
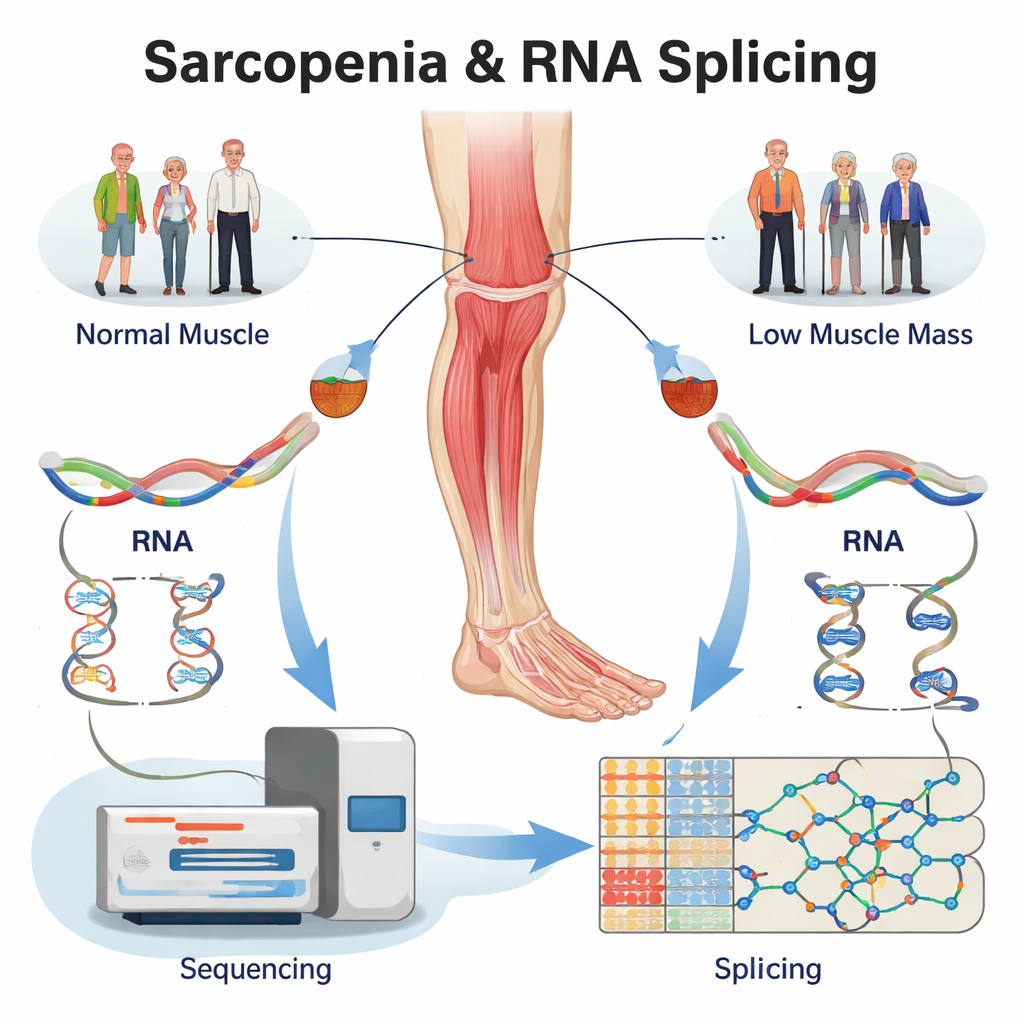
Re-reading existing data to find hidden RNA errors
Instead of recruiting new volunteers, the team re-analyzed an existing large RNA sequencing dataset from human thigh muscle biopsies. The samples came from four groups: older adults with full-blown sarcopenia, people with only low muscle mass, people with only low muscle strength, and age-matched healthy controls. Using specialized software, they first confirmed that hundreds of genes were turned up or down in the diseased muscles. More importantly, they then looked at how each gene’s RNA was spliced, cataloguing more than 5,000 splicing changes across the three problem groups compared with healthy muscle. These changes were concentrated in a few basic types—such as skipping an exon, or choosing an alternative cut site—patterns that are known to strongly alter protein structure.
Energy factories and cell “fuel gauge” pathways are disrupted
The splicing changes turned up in genes that sit at the heart of muscle biology. Many affected genes build parts of the mitochondria, the tiny energy factories that power muscle contraction. In sarcopenic muscle, 16 genes involved in oxidative phosphorylation—the process that turns nutrients into usable energy—showed altered splicing, especially components of the mitochondrial complexes that handle electron transport and proton pumping. Other changes appeared in genes that make or use NAD⁺, a key molecule for cellular energy and repair. Rather than being simply more or less abundant, important enzymes such as CD38, PARP2, and several sirtuins produced different RNA variants, suggesting that the balance between NAD⁺ production and consumption is distorted at the splicing level. The affected genes also clustered in well-known muscle signaling routes, including AMPK and FoxO pathways, which act like cellular fuel gauges and stress responders.
Shared patterns across different stages—and across species
Strikingly, many of the same genes showed splicing changes not only in people with full sarcopenia, but also in those who had only low muscle mass or only low strength. This suggests that abnormal splicing begins early and cuts across the different clinical faces of the disease. To test whether these changes could be reproduced in an animal model, the researchers induced muscle wasting in mice with the steroid dexamethasone, a standard way to mimic some features of sarcopenia. The mice developed weaker grip strength and poorer performance on running and balance tests. When the team sequenced mouse muscle RNA, they again found hundreds of genes with altered splicing. About one-fifth of these overlapped with the human sarcopenia genes, and many were involved in controlling transcription, chromatin structure, and splicing itself—higher-level switches that can reshape a cell’s entire protein output.
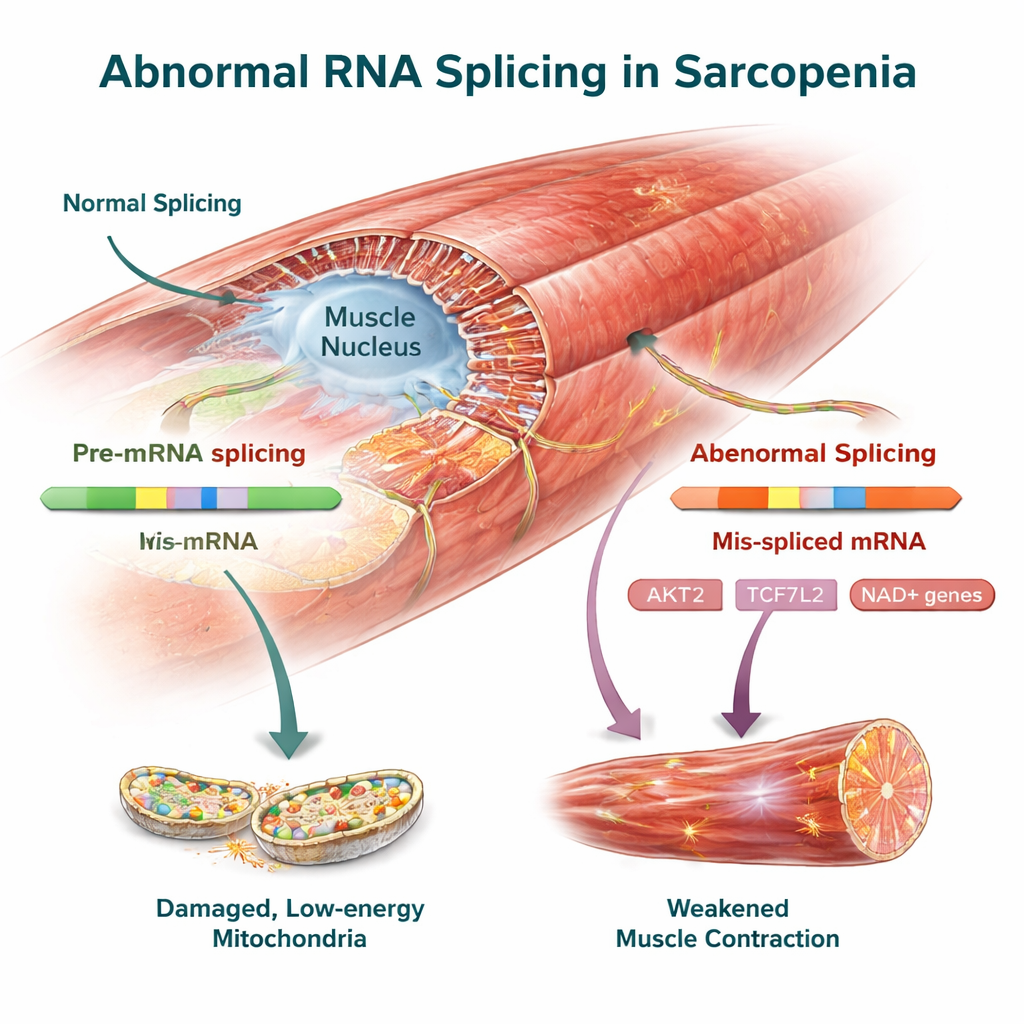
Key muscle control genes as potential treatment targets
Among the cross-species changes were several genes already linked to muscle health. AKT2, a central regulator of growth and protein building in muscle, showed distinct splicing shifts in both humans and mice that may blunt its ability to maintain muscle mass. TCF7L2, part of the Wnt signaling pathway, along with FMNL2 and USP40, which help organize the muscle’s internal scaffolding and protein quality control, also carried altered RNA variants. The team confirmed these splicing changes experimentally in mouse muscle, strengthening the case that they are real features of the disease rather than artifacts of data analysis.
What this means for preventing frailty
For non-specialists, the message is that sarcopenia is not just about muscles “wearing out” or genes being switched on or off. It also involves a widespread mis-editing of RNA messages that guide how muscle cells make and maintain their machinery, especially the systems that generate energy and sense stress. Because splicing can, in principle, be corrected with precisely designed RNA-based drugs, the conserved splicing errors discovered here—particularly in genes like AKT2 and TCF7L2—offer a new set of leads for therapies to help aging muscles stay stronger for longer.
Citation: Li, Y., Guo, X., Li, S. et al. Transcriptomic characterization of the aberrant alternative splicing in skeletal muscles of sarcopenia patients. Sci Rep 16, 5334 (2026). https://doi.org/10.1038/s41598-026-35002-w
Keywords: sarcopenia, muscle aging, RNA splicing, mitochondria, AKT2