Clear Sky Science · en
Metagenomic sequencing identifies potential respiratory pathogens in PCR-negative subset of surveillance samples
Why hidden germs matter for everyone
When you come down with a sore throat or cough, doctors often rely on rapid lab tests to look for usual suspects like flu or COVID-19. But what happens when those tests say “nothing found,” even though you clearly feel sick? This study peeks behind that curtain by using a powerful DNA-based approach to search for germs that standard tests miss, revealing a more complex picture of respiratory infections and how we might track them in the future. 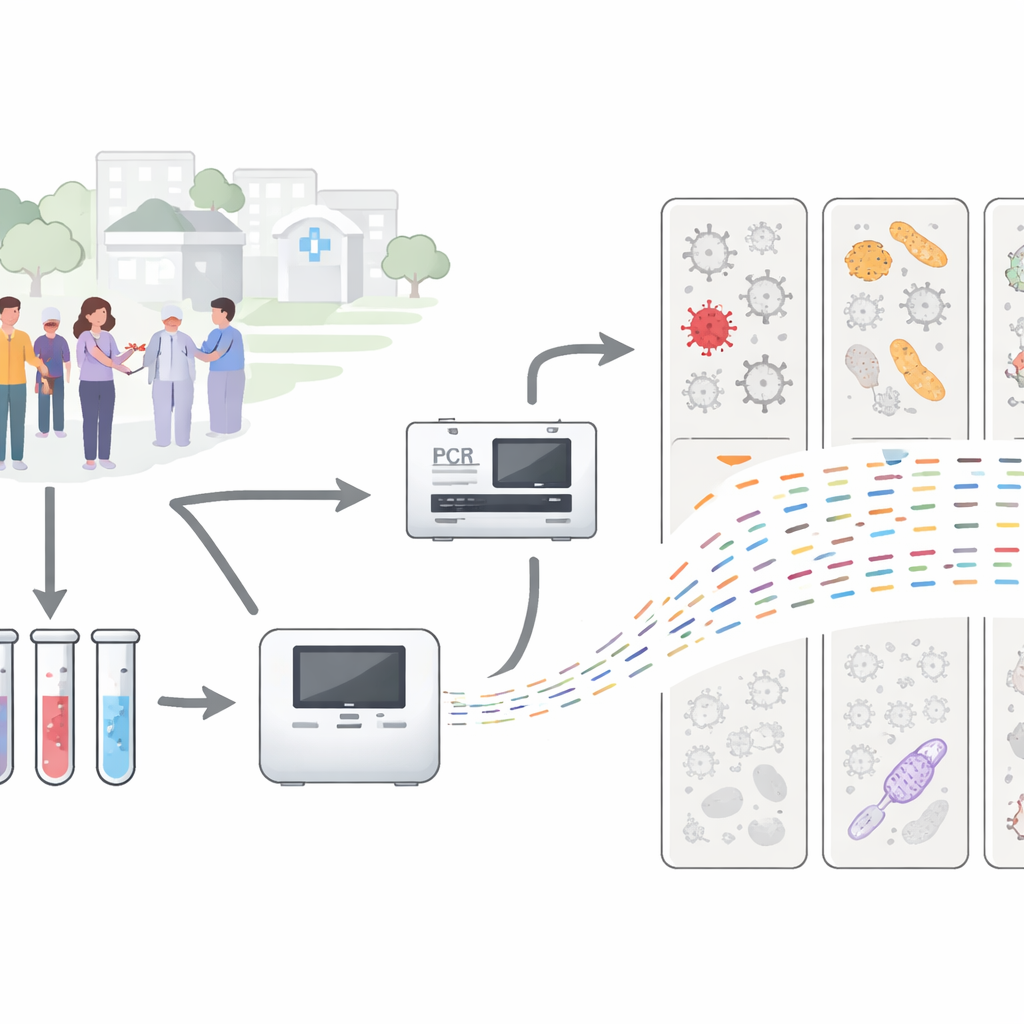
Looking beyond the usual test panel
During the COVID-19 pandemic, California ran a large program to monitor respiratory infections in people visiting clinics across several counties. Each person’s nose or throat sample was tested with common lab panels that look for a fixed list of viruses and bacteria, plus a separate test for SARS-CoV-2. More than half of these samples came back negative for every germ on the list, even though the patients had clear cold- or flu-like symptoms. The researchers behind this paper took a closer look at 305 of these “mystery” samples, along with 26 samples already known to be positive, to see whether more advanced sequencing could uncover what was really there.
Reading all the genetic material in a sample
Instead of asking, “Is virus X present?” the team used metagenomic sequencing, which essentially asks, “What genetic material is in this sample, whatever it is?” They first extracted all DNA and RNA from each swab, copied it so there was enough to analyze, and then fed it into high-throughput sequencing machines. In a subset of samples, they added an extra step using a “probe-capture” panel designed to fish out viral genetic material, making it easier to spot viruses that might otherwise be drowned out by abundant human or bacterial material. Computer programs then compared millions of short genetic snippets to large reference databases to see which viruses, bacteria, and fungi were present. 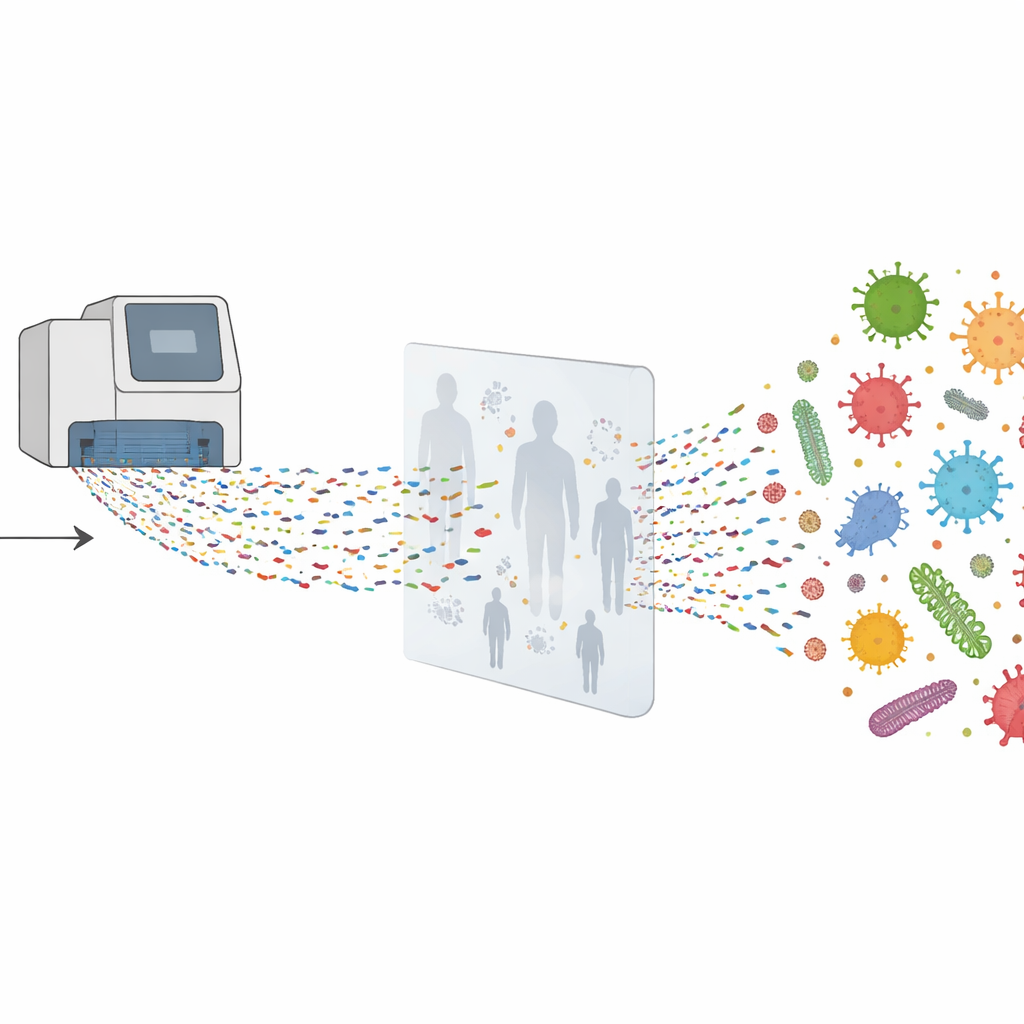
Uncovering overlooked viruses and microbes
Even among samples that had tested negative by routine methods, the sequencing approach found human respiratory viruses in about 5 percent of cases. These included influenza C virus, human bocavirus, rhinoviruses, and even a few SARS-CoV-2 infections that standard tests had missed. For many of these viruses, the team recovered almost complete genomes, allowing them to see how closely related the strains were to each other and to viruses found in other regions and years. They also found that some samples were dominated by a single kind of bacterium or fungus, such as certain Moraxella, Pseudomonas, or Penicillium species, hinting at possible bacterial or fungal involvement in respiratory illness or at least in shaping the local microbial community in the airway.
What missed infections can teach us
By reconstructing whole viral genomes, the researchers could tell, for example, that the bocavirus strains in neighboring counties were nearly identical, suggesting local spread, and that each rhinovirus infection tended to involve a distinct strain, including one closely related to a recently described new type. They also saw how the virus-enrichment step boosted the amount and completeness of viral genetic material, especially for harder-to-detect viruses like influenza C. At the same time, many negative samples still showed no clear pathogen, underscoring that some respiratory symptoms may come from non-infectious causes, poor-quality samples, or microbes at levels too low for detection.
What this means for future health monitoring
For everyday clinical care, rapid targeted tests will likely remain the workhorses: they are cheaper, faster, and easier to run than sequencing. But this study shows that when those tests come up empty—especially in severe or unexplained cases—broad metagenomic sequencing can reveal hidden infections, identify rare or unusual viruses, and provide full genomes for tracking variants over time. As the technology becomes more affordable and standardized, it could become a powerful complement to routine testing, helping public health officials spot new threats early and better understand how a wide range of viruses, bacteria, and fungi circulate in our communities.
Citation: Mascarenhas, A.C., Kantor, R.S., Thissen, J. et al. Metagenomic sequencing identifies potential respiratory pathogens in PCR-negative subset of surveillance samples. Sci Rep 16, 9308 (2026). https://doi.org/10.1038/s41598-025-33917-4
Keywords: respiratory infections, metagenomic sequencing, virus surveillance, diagnostic testing, pathogen discovery