Clear Sky Science · en
Conserved regions and molecular cloning of Acid and Alkaline phosphatases in Streptomyces sp. MMA-NRC
Why soil phosphorus matters to our food
Modern agriculture leans heavily on phosphorus, a key ingredient in plant fertilizers that helps crops build DNA, store energy, and grow strong roots. Yet much of the phosphorus we spread on fields quickly becomes locked into insoluble minerals that plants cannot use, while global reserves of mineable phosphate rock are finite and unevenly distributed. This study explores a biological alternative: harnessing soil-dwelling microbes, and even engineered bacteria, to unlock phosphorus from rock phosphate, potentially reducing our dependence on chemical fertilizers and easing environmental damage.
Tiny helpers that unlock “stuck” nutrients
In many soils, the total amount of phosphorus is high, but only a tiny fraction is in a form that plant roots can absorb. Excess fertilizer not only wastes money but also washes into rivers and lakes, triggering algal blooms and dead zones. The authors focus on a group of soil bacteria called Streptomyces, already known for making antibiotics and powerful enzymes. One strain, named Streptomyces sp. MMA-NRC, can dissolve rock phosphate, a cheap but poorly soluble fertilizer. The team set out to understand and harness two of its key enzymes, acid phosphatase and alkaline phosphatase—molecular tools that snip phosphorus-containing groups from otherwise unusable compounds.
Reading and modeling the enzymes’ blueprints
First, the researchers isolated the genes that code for these two phosphatase enzymes in the MMA-NRC strain. They amplified and sequenced the genes, which turned out to encode proteins of 488 and 560 building blocks (amino acids), and deposited the sequences in public databases. Using bioinformatics tools, they compared these proteins with related enzymes from other bacteria, showing that the MMA-NRC versions are closely related—about 99% similar—to those from another Streptomyces strain. They then built detailed three-dimensional computer models of the enzymes, checking their quality with established methods that assess whether each amino acid sits in a realistic position. The models passed these checks with high scores, suggesting that the virtual structures closely resemble the real ones.
Testing how well the enzymes grab rock phosphate
With the 3D models in hand, the scientists ran “docking” simulations to see how strongly the enzymes might bind to rock phosphate, their target substrate. In these simulations, the enzyme and mineral are brought together in many possible orientations, and a computer estimates which arrangements are most stable. Both acid and alkaline phosphatases from MMA-NRC showed very strong predicted binding energies to rock phosphate, meaning they should interact tightly with the mineral’s surface. Specific amino acids in the enzymes formed hydrogen bonds and hydrophobic contacts with the rock phosphate model, outlining probable active sites where chemical reactions to release phosphorus would occur.
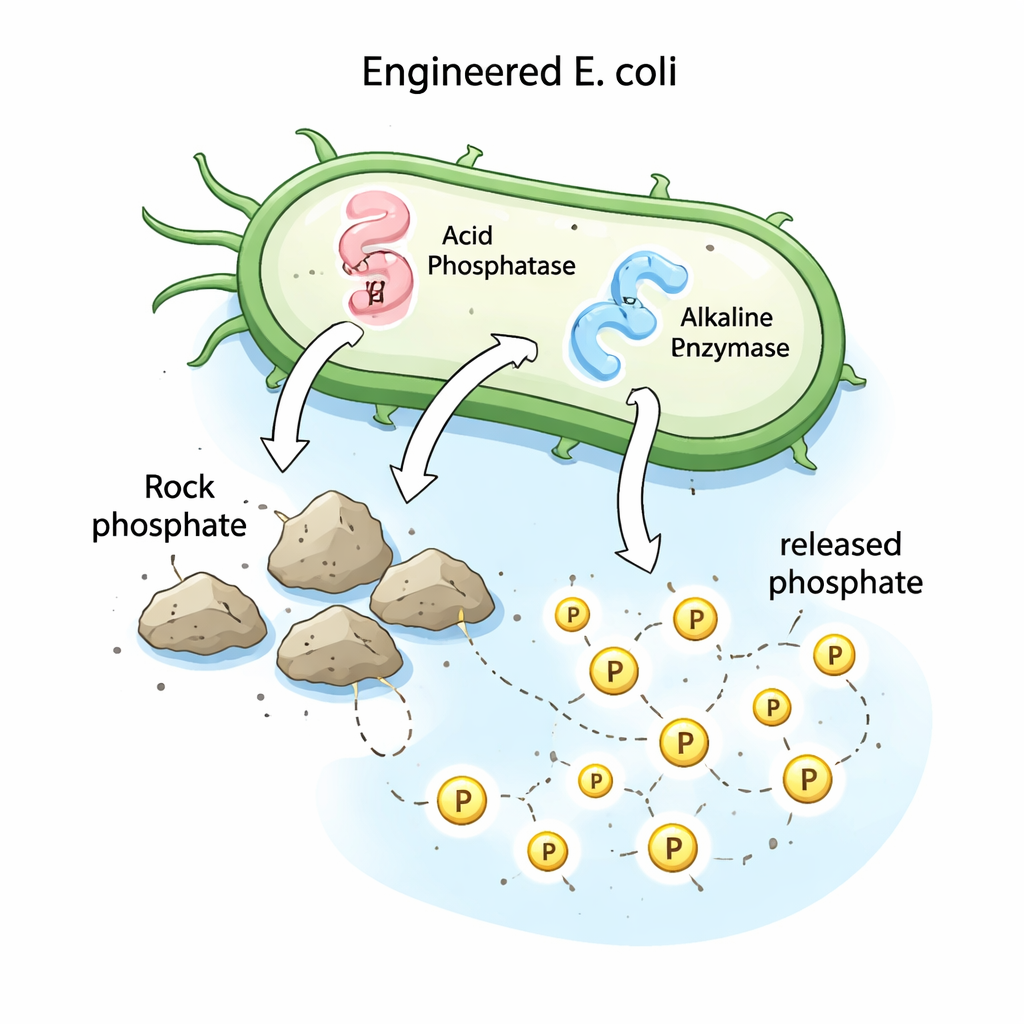
Turning a lab bacterium into a phosphorus releaser
To test these ideas experimentally and to create a practical tool, the team moved the phosphatase genes from Streptomyces into a well-studied laboratory bacterium, Escherichia coli DH5α. They inserted each gene into a standard cloning vector and transformed the plasmids into E. coli, selecting colonies that successfully took up the new DNA. These engineered strains were then grown in a medium containing rock phosphate as the only phosphorus source. While unmodified E. coli could not release measurable phosphorus, the recombinant strains producing either acid or alkaline phosphatase released about 53 and 57 milligrams of soluble phosphorus per liter after seven days—far higher than the original Streptomyces strain, which released about 35 milligrams per liter under the same conditions.
What this could mean for greener farming
For non-specialists, the takeaway is straightforward: by understanding and reusing nature’s own enzymes, scientists can turn inexpensive, poorly soluble rock phosphate into a more accessible nutrient source for crops. The high activity of the modeled and cloned phosphatases suggests that strains like Streptomyces sp. MMA-NRC, or engineered bacteria carrying its genes, could become components of biofertilizers that reduce reliance on conventional phosphate fertilizers. Such biological solutions could help farmers maintain yields while lowering pollution and easing pressure on limited global phosphate reserves, supporting more sustainable and resilient food production systems.
Citation: Abd El-Aziz, N.M. Conserved regions and molecular cloning of Acid and Alkaline phosphatases in Streptomyces sp. MMA-NRC. Sci Rep 16, 7493 (2026). https://doi.org/10.1038/s41598-025-33881-z
Keywords: phosphate-solubilizing bacteria, biofertilizer, Streptomyces, phosphatase enzyme, rock phosphate