Clear Sky Science · en
Toward cardiac electrophysiology digital twins with an efficient open source scalable solver on GPU clusters
Why building virtual hearts matters
When doctors treat heart rhythm problems, they are often working partly in the dark: they can measure electrical signals on the body surface but cannot easily see how those signals spread deep inside the heart. This study describes powerful new computer tools that can simulate a person’s heart beating in detail, fast enough to explore many “what if” scenarios. By combining realistic heart anatomy with cutting-edge graphics processors, the authors move us closer to safe, low-cost virtual experiments on digital copies of individual patients’ hearts.
From simple models to digital twins
Modern heart simulations can represent activity from the level of individual cells up to the whole organ, but faithfully reproducing how electricity moves through the heart is both mathematically complex and computationally expensive. A key goal for medicine is the “cardiac digital twin”: a personalized computer model that mimics a specific patient’s heart and can be used to test drugs, devices, or pacing strategies before they are tried in the clinic. To make this feasible, the software running these simulations must be both fast and accurate, must include important anatomical structures, and must be openly available so that researchers worldwide can check, improve, and reuse it.
Capturing the heart’s hidden wiring
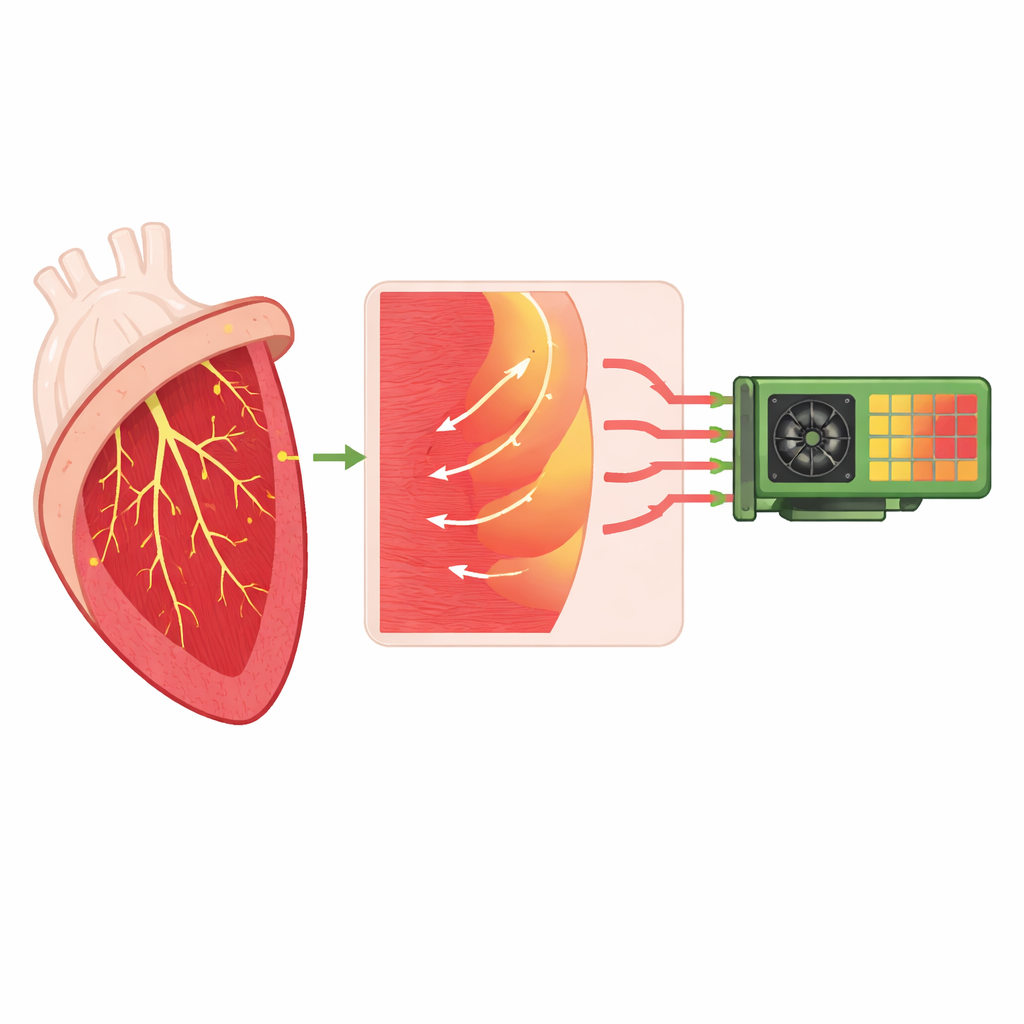
The work centers on an open-source simulator called monoalg3d, which models how electrical signals travel through the heart’s muscle and its specialized wiring network. That network, known as the Purkinje system, rapidly delivers the heartbeat-triggering signal from the upper conduction pathways to many tiny connection points spread across the inner surface of the ventricles. At each of these junctions, a thin Purkinje fiber must excite a much larger mass of heart muscle, introducing a natural delay and, under some conditions, a risk of blocked or abnormal conduction. The authors enhance monoalg3d so it represents this wiring and its junctions more realistically, including the asymmetry between signals traveling from Purkinje to muscle and from muscle back into the network.
Turning supercomputers and GPUs into heart labs
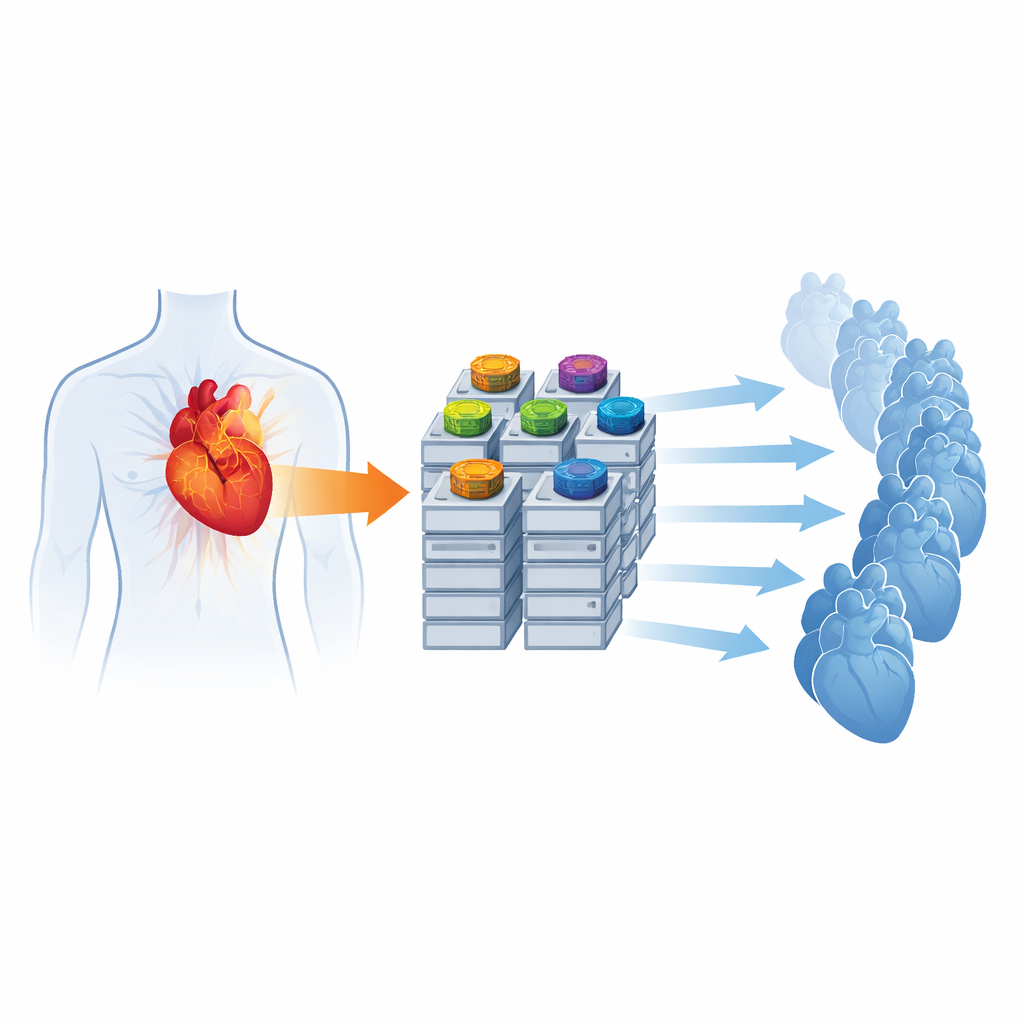
To run such detailed models quickly, the team re-engineers how the simulator uses hardware. They redesign the numerical core so that both the local cell dynamics and the spread of voltage across the heart are solved directly on graphics processing units (GPUs), devices originally created for rendering images but now widely used for scientific computing. By carefully organizing data for GPU libraries and avoiding frequent transfers between CPU and GPU, they show that a fully GPU-based setup can speed up a standard benchmark by nearly elevenfold compared with a traditional CPU-only version. They also introduce a new, compact output format to drastically cut storage needs and a message-passing feature that can launch hundreds of simulations in parallel across a supercomputer, each exploring a different set of physiological parameters.
Teaching virtual hearts to match real patients
Beyond raw speed, the authors test whether their improved solver can support realistic patient-specific simulations. Using a three-dimensional reconstruction of a human heart from medical imaging, they attach a synthetic Purkinje network and adjust key junction parameters: the resistance at each connection and how many neighboring muscle regions each Purkinje terminal excites. They automatically sweep through hundreds of combinations of these values, running simulations that start from a stimulus in the main conduction bundle, and measure both the small delays at each junction and the resulting electrocardiogram (ECG) signals on the chest. For a fine-resolution heart mesh, they identify parameter choices that produce physiologically plausible activation times and ECG waveforms that correlate well with a real patient’s recordings, all while keeping individual simulation times within a few hours even when 512 runs are executed concurrently.
Many possible hearts that look the same outside
An intriguing outcome is that different internal wiring configurations and junction strengths can lead to very similar ECGs at the body surface. In other words, several distinct digital hearts can appear equally consistent with the same clinical data. The authors show that some Purkinje junctions are especially influential, with their local delays varying substantially between equally good-fitting simulations, while overall activation timing across the ventricles remains stable. This non-uniqueness suggests that future digital twin frameworks will need additional data or constraints—such as more detailed imaging or invasive measurements—to pin down a single most likely internal configuration for a given patient.
What this means for future care
In summary, this study delivers a freely available, GPU-accelerated simulator that can handle detailed heart anatomy, realistic conduction wiring, and large batches of personalized simulations on modern computing clusters. For non-specialists, the key message is that it is becoming practical to run fast, high-fidelity virtual heart experiments that resemble individual patients, including subtle delays at the heart’s internal wiring junctions. While more work is needed to fully personalize these models and resolve internal uncertainties, the tools introduced here form an important building block for future digital-twin–based diagnosis, risk assessment, and treatment planning in cardiology.
Citation: Berg, L.A., Oliveira, R.S., Camps, J. et al. Toward cardiac electrophysiology digital twins with an efficient open source scalable solver on GPU clusters. Sci Rep 16, 9619 (2026). https://doi.org/10.1038/s41598-025-33709-w
Keywords: cardiac digital twin, heart electrophysiology simulation, Purkinje conduction system, GPU computing, personalized cardiology