Clear Sky Science · en
A novel strain isolated from methane seeps in the Kattegat belongs to the planctomycetal species Novipirellula methanifontis sp. nov.
A hidden world in bubbling undersea springs
Far below the waves of the Kattegat strait between Denmark and Sweden, methane gas seeps up through rocky “bubbling reefs,” feeding a bustling but invisible community of microbes. This study tells the story of one such microbe—an unusual bacterium with a big genome, a taste for complex sugars, and a strange way of dividing—that turned out to be a completely new species. Understanding organisms like this helps scientists piece together how undersea ecosystems recycle carbon and may even point to future biotechnological tools, from new enzymes to potential antibiotics.
Life on the edge of methane springs
The newly described bacterium was originally collected decades ago from methane seeps in the Kattegat, where gas bubbles continuously rise from carbonate-cemented reefs on the seafloor. These shallow seeps host oxygenated sediments in which methane is rapidly oxidized, creating steep chemical gradients and a variety of ecological niches. The strain, stored for years in the collection of microbiologist Heinz Schlesner, was recently revived and given the laboratory name SH528T. Although it comes from a methane-rich habitat, the team showed that the bacterium does not actually “eat” methane. Instead, it breathes oxygen and feeds on organic matter, fitting into the broader community that surrounds the methane-fueled hotspot.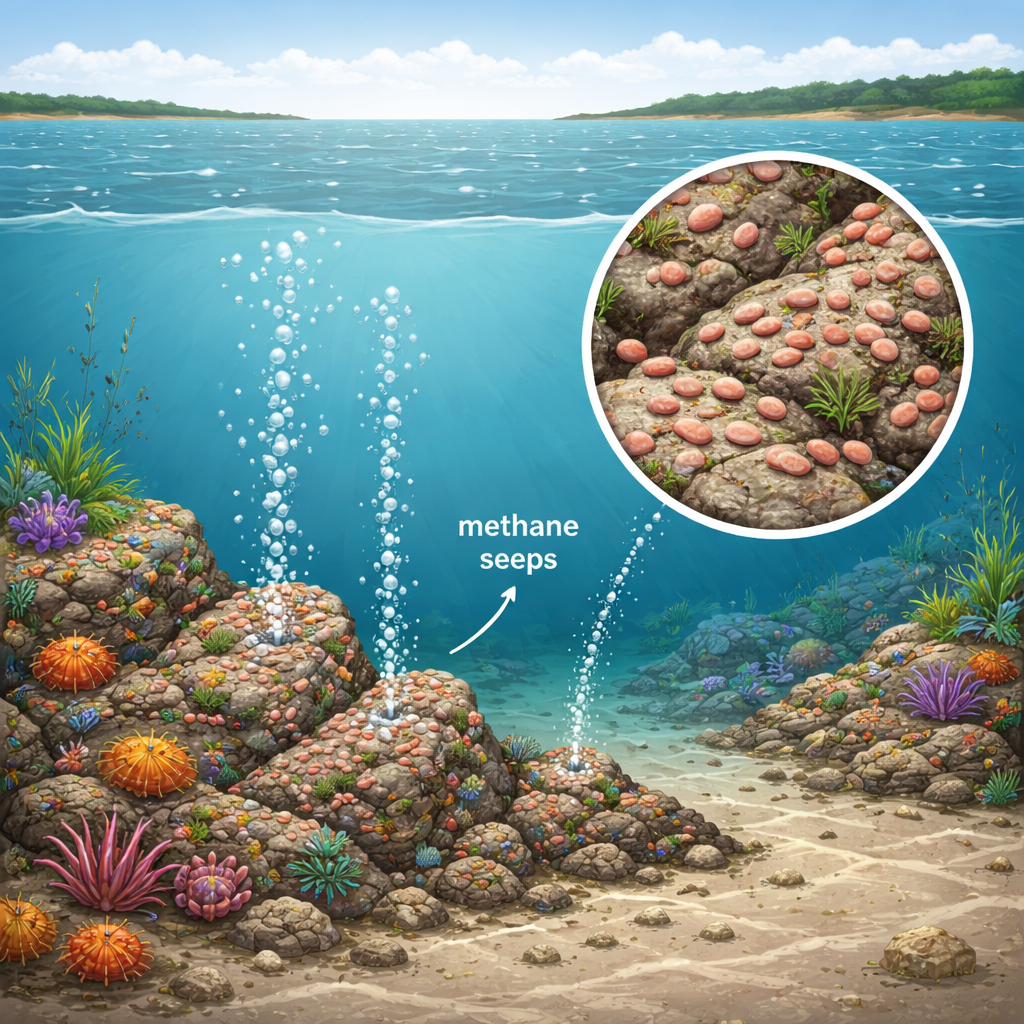 
An odd bacterium with a budding lifestyle
Under the microscope, SH528T looks and behaves very differently from textbook bacteria. It belongs to the phylum Planctomycetota, a group famous for unusual cell biology. The cells are salmon colored, ellipsoid to nearly round, and form small shapeless clumps in liquid culture. Rather than splitting in half down the middle, they reproduce by “polar budding”: a small round daughter cell grows out of one end of a larger mother cell, elongates, and then pinches off. The cells thrive at moderate temperatures around 24 °C and a near-neutral to slightly alkaline pH, matching the mild conditions of their coastal marine home.
A giant genome packed with sugar-degrading tools
When the researchers sequenced the bacterium’s DNA, they found a single circular chromosome of about 10.5 million base pairs—large even by the standards of its already gene-rich relatives. The genome carries more than 7,000 protein-coding genes, including an impressive collection of nearly 300 genes for so-called carbohydrate-active enzymes. These enzymes specialize in breaking down and modifying complex polysaccharides, the tough sugar-based molecules that make up things like algal cell walls. Such a toolkit suggests that SH528T is well equipped to survive where simple sugars are scarce, by digesting more recalcitrant organic matter and helping recycle carbon in marine sediments.
Fitting the newcomer into the bacterial family tree
Placing SH528T in the tree of life required multiple lines of genetic evidence. The team compared key molecular markers—such as the 16S ribosomal RNA gene, overall average nucleotide identity, and the percentage of shared proteins—with those of known relatives. All analyses firmly placed the strain within the family Pirellulaceae and the genus Novipirellula, but the similarity values consistently fell below accepted species-level thresholds. Pangenome analysis, which compares the full gene repertoires of related strains, revealed that SH528T carries more than 1,700 unique gene clusters, underscoring its distinct genetic identity and hinting at specialized ecological adaptations, including its rich enzyme set and capacity to produce various secondary metabolites. 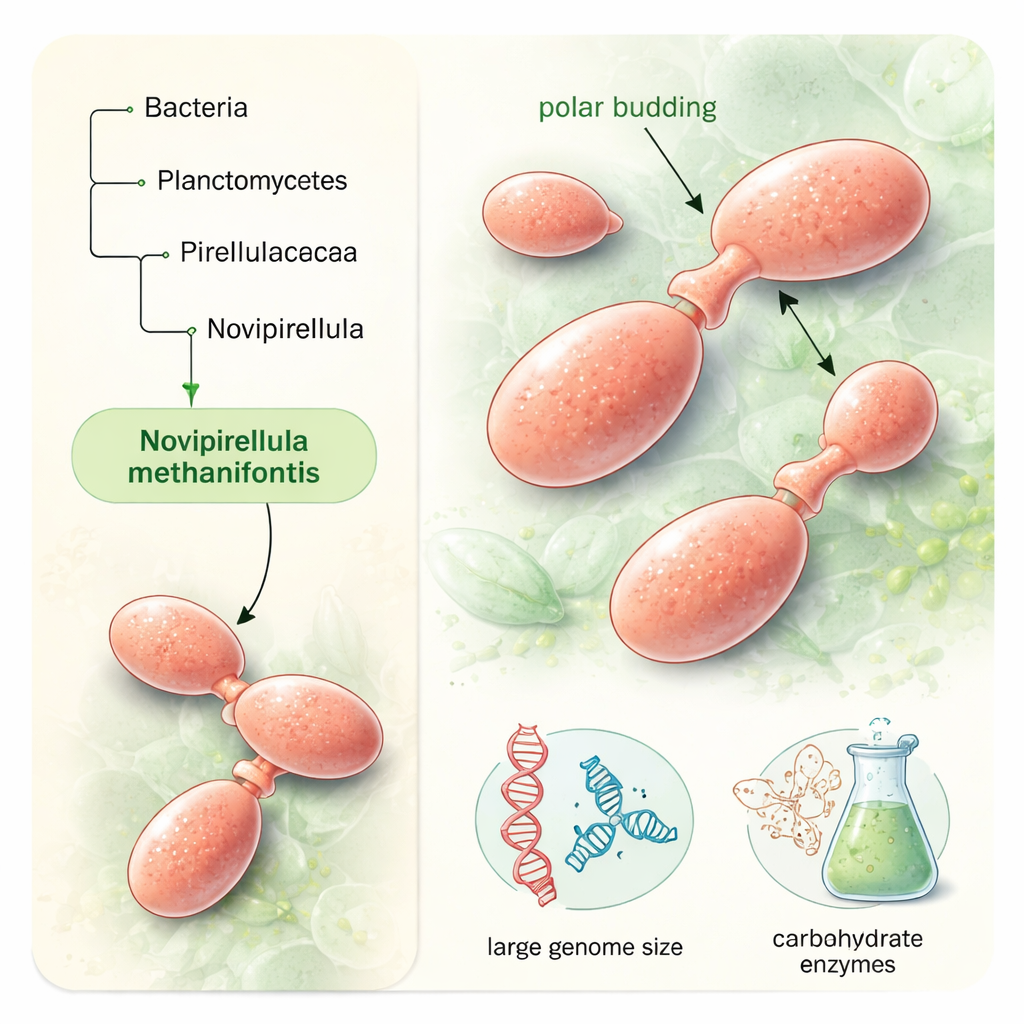
A new name and why it matters
Taken together—its origin at methane seeps, distinctive cell shape and budding division, unusually large and diverse genome, and clear genetic separation from close relatives—these findings justified recognizing SH528T as a new species. The authors propose the name Novipirellula methanifontis, meaning “of a methane spring.” For non-specialists, the key message is that even well-studied coastal seas still hide unknown microbial players that quietly drive elemental cycles. By cataloging and characterizing such bacteria, scientists deepen our understanding of marine ecosystems and uncover new enzymes and bioactive compounds that could eventually be harnessed in biotechnology, medicine, or environmental applications.
Citation: Staack, M., Haufschild, T., Hammer, J. et al. A novel strain isolated from methane seeps in the Kattegat belongs to the planctomycetal species Novipirellula methanifontis sp. nov.. Sci Rep 16, 6537 (2026). https://doi.org/10.1038/s41598-025-33410-y
Keywords: marine bacteria, methane seeps, planctomycetes, microbial diversity, carbon cycling