Clear Sky Science · en
Telomeric G-quadruplex intermediates unveiled by complex Markov network analysis
Hidden knots at the ends of our DNA
At the tips of our chromosomes lie protective caps called telomeres, often compared to the plastic ends of shoelaces. Within these regions, the DNA can tie itself into unusual four-stranded knots known as G-quadruplexes. These tiny structures can influence how our genes behave and how cells age or turn cancerous. This study peeks behind the scenes of how one such knot gradually comes undone, using detailed computer simulations and advanced data analysis to reveal fleeting shapes that are almost impossible to catch in experiments.
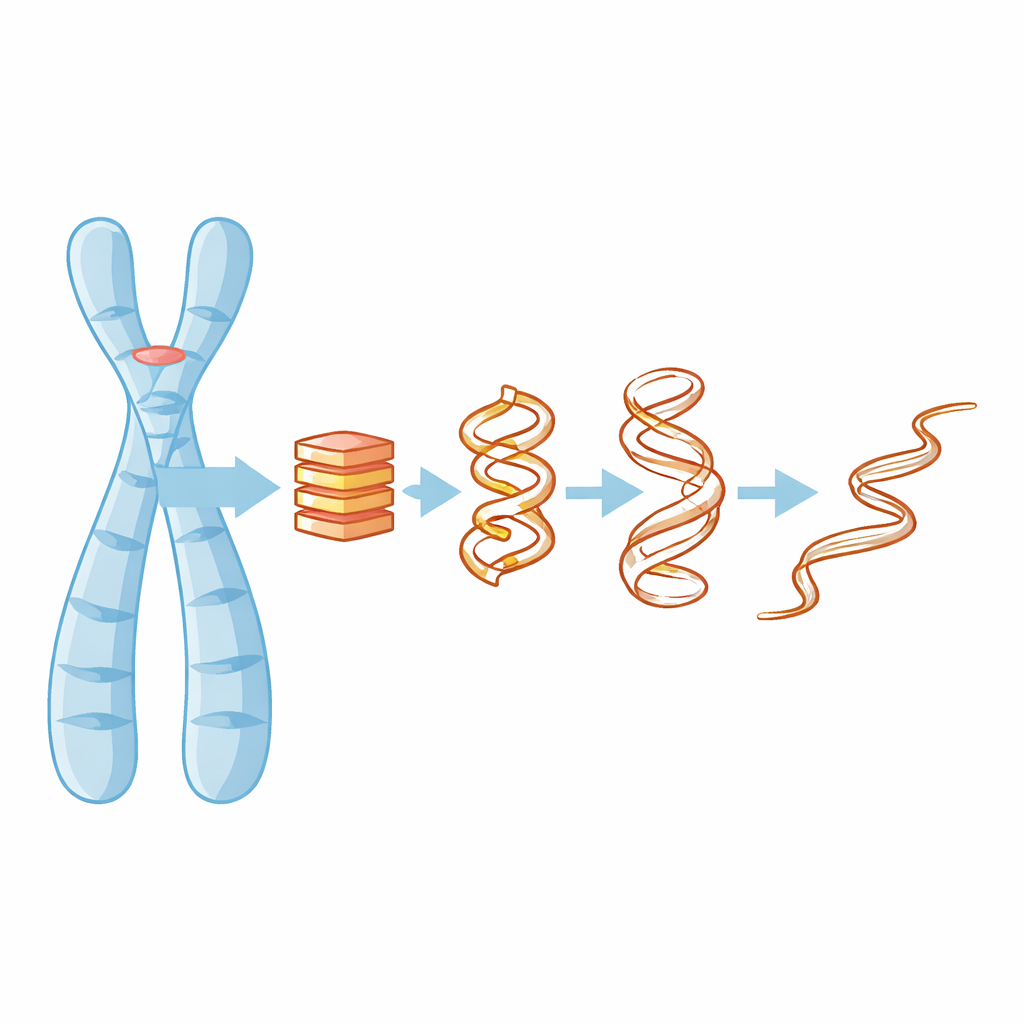
Why four-stranded DNA knots matter
Most of us learn that DNA is a double helix, but in guanine-rich stretches—such as those in telomeres—the molecule can fold into compact four-stranded stacks called G-quadruplexes. These stacks are held together by flat layers of guanine bases and tiny positively charged ions, like potassium, that sit in the middle and help glue everything together. When a G-quadruplex forms or falls apart, it can block or expose parts of the genome, affecting gene activity, DNA replication, and even how cancer cells maintain their telomeres. Understanding how these structures lose their shape is therefore key to figuring out how to target them with drugs or predict when they might fail.
Watching a molecular knot come undone
The researchers focused on a human telomeric G-quadruplex in which four segments of a single DNA strand line up in parallel and stack into a compact column. They used all-atom molecular dynamics simulations—virtual experiments that track every atom’s motion—to heat eight copies of this structure to temperatures near its melting point. To help the system explore different shapes, they used a strategy called replica exchange, where simulations at different temperatures occasionally swap configurations. In seven of the eight cases the G-quadruplex held together; in one, it fully unfolded. That rare unfolding event became a detailed case study, showing not just the before and after, but a whole sequence of intermediate shapes along the way.
The crucial role of tiny ions
A central finding is that the small ions inside the G-quadruplex act like linchpins. As long as at least one ion stays nestled between the guanine layers, the structure remains mostly intact, perhaps a bit loosened but still recognizable. When both ions eventually leave the central channel, however, the stack rapidly destabilizes. The guanine layers warp and individual strands peel away, and the overall DNA becomes more extended and flexible. Other simulated molecules that lost only one ion entered wobbly but still partly ordered states, suggesting that full unfolding demands the loss of both stabilizing ions—a clue to why some G-quadruplexes are so stubbornly robust in the cell.
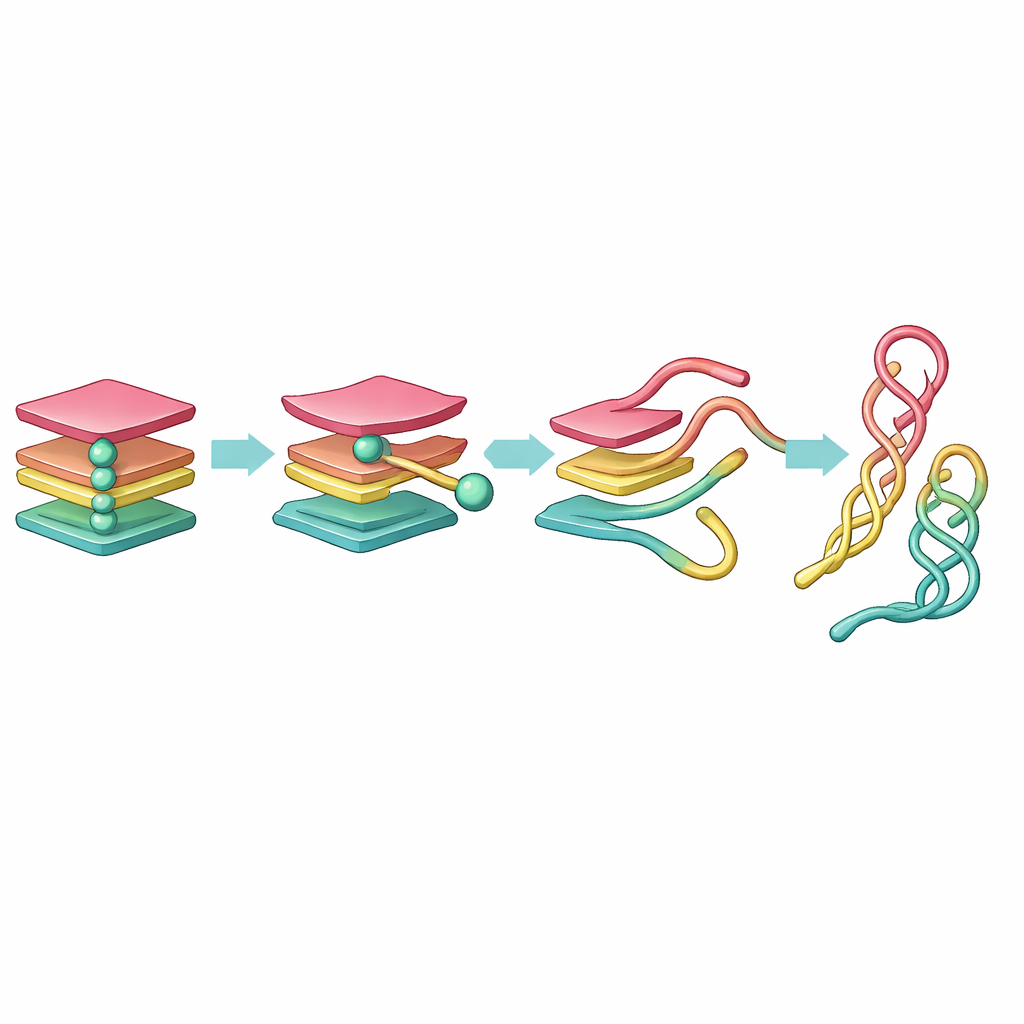
Uncovering hidden waypoints in the journey
The raw simulation produced an enormous amount of data, far too complex to interpret by eye. To tame this, the team first simplified the DNA by representing each base as a single bead, then used mathematical tools to reduce the motion to just a few key coordinates. They applied two methods: Principal Component Analysis (PCA), which highlights directions of largest motion, and time-independent Component Analysis (tICA), which filters for the slow, long-lived changes that mark true structural shifts. From these reduced coordinates they built so-called conformational Markov networks, in which each node is a typical DNA shape and links show how the molecule hops from one shape to another.
A preferred route from knot to loose strand
This network view revealed a clear unfolding pathway. Starting from a fully folded G-quadruplex, one strand first detaches, forming a three-stranded "triplex" state. That triplex then reorganizes into a two-stranded "hairpin" and a set of nearby loose strands. The hairpin and a related "cross-hairpin" arrangement prove surprisingly long-lived, acting as stable waypoints on the journey from a tight knot to a more relaxed DNA configuration. While PCA split this progression into many similar-looking states, tICA distilled it into a small set of distinct, kinetically meaningful steps, showing that triplex-like shapes are short-lived, while hairpin-like shapes linger much longer.
What this means for biology and medicine
To a non-specialist, the key message is that telomeric G-quadruplexes do not simply snap from folded to unfolded. Instead, they melt through a series of short-lived and longer-lived shapes, with tiny ions acting as gatekeepers of stability. The simulations suggest that three-stranded intermediates are fleeting, whereas two-stranded hairpins and cross-shaped forms are more stable resting points. Because these intermediate states may also appear in the reverse, folding direction, they are likely to be crucial in how G-quadruplexes form, change, and interact with potential drugs. Mapping this hidden landscape of shapes brings scientists a step closer to designing molecules that selectively stabilize or destabilize these DNA knots in the fight against cancer and other diseases.
Citation: Sáinz-Agost, A., Falo, F. & Fiasconaro, A. Telomeric G-quadruplex intermediates unveiled by complex Markov network analysis. Sci Rep 16, 8308 (2026). https://doi.org/10.1038/s41598-025-29993-1
Keywords: G-quadruplex, telomere DNA, molecular dynamics, protein-free DNA folding, cancer therapeutics