Clear Sky Science · en
A high-quality Chromosome-level genome assembly of Gynostemma guangxiense (Cucurbitaceae)
A forest vine with hidden healing power
Deep in the limestone forests of southern China grows a little-known climbing vine, Gynostemma guangxiense. Local people have long used this plant as a traditional remedy for liver troubles and chronic coughs, and it is rich in natural compounds similar to those found in ginseng. As demand for its better-known cousin Gynostemma pentaphyllum has surged, wild populations have been strained. To protect these valuable plants and better understand their medicinal promise, scientists have now decoded the full genetic blueprint of G. guangxiense at the level of individual chromosomes.

Why this little vine matters
Gynostemma plants occupy a special place in herbal medicine because they are one of the very few groups, besides ginseng, that produce dammarane-type saponins—molecules linked to immune support, heart health and anti-cancer activity. G. guangxiense looks much like G. pentaphyllum but tastes sweeter and has been reported to contain high levels of saponins, flavonoids and essential trace elements such as iron, manganese, zinc and copper. These features make it an attractive candidate both as an alternative medicinal resource and as a window into how such health-boosting compounds arise in plants. Yet, until now, scientists had only fragments of its genetic information, such as its chloroplast DNA, limiting deeper study.
Reading the plant’s instruction manual
To unlock this hidden information, the research team collected healthy leaves from a wild female plant in Guangxi Province and extracted its DNA and RNA. They then used a combination of modern sequencing approaches that capture both short and long stretches of DNA, along with a technique called Hi-C that reveals how these stretches are folded and linked inside the cell. By carefully piecing together these different data streams, and cross-checking them with RNA readouts from several tissues—buds, leaves, stems, flowers and fruits—they assembled a highly complete version of the plant’s genome and checked its accuracy using widely accepted quality tests.
Building chromosomes from countless fragments
The final result is a genome of about 565 million building blocks of DNA, organized into 11 pseudochromosomes that cover more than 99 percent of the assembled sequence. Quality measures show that almost all standard plant benchmark genes are present and intact, indicating that very little information is missing. The researchers also mapped where repeated DNA stretches lie along the chromosomes and found that over two-thirds of the genome consists of such repeats, especially a type known as long terminal repeat elements. They cataloged 27,527 genes that code for proteins, and remarkably, nearly 98 percent of these genes could be connected to known functions or families by comparing them to international databases.
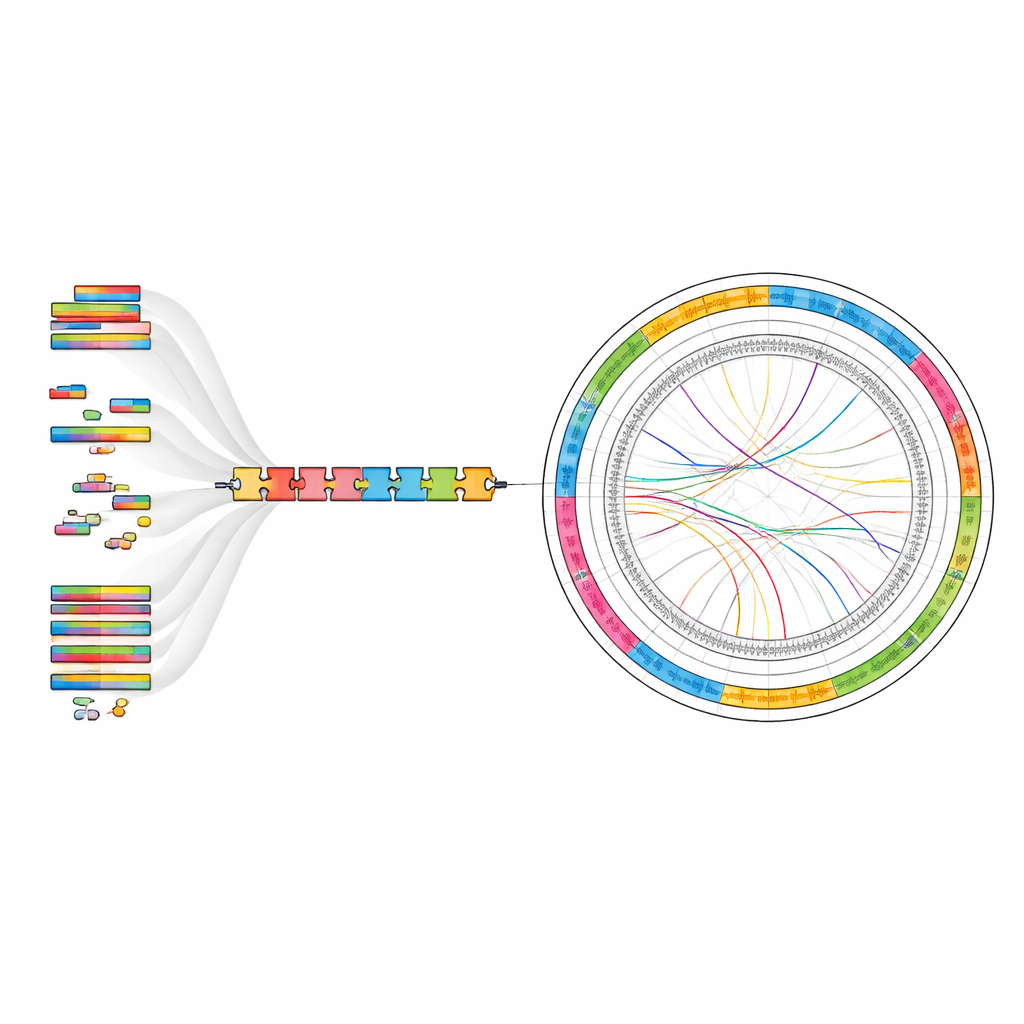
What the genome reveals about this plant
Beyond listing genes, the genome map shows how different features—genes, repeats and non-coding RNAs—are arranged along each chromosome, and how regions correspond to one another across the genome. This provides clues to past chromosome rearrangements and the forces shaping the plant’s evolution. The team also identified thousands of pieces of non-coding RNA, including transfer RNAs and ribosomal RNAs, which help run the cell’s protein-making machinery. Together, these findings offer a detailed reference for exploring how G. guangxiense produces its rich blend of saponins and other bioactive molecules, and how it adapts to its rocky forest habitat.
A new foundation for medicine and conservation
By delivering a carefully checked, chromosome-level genome for G. guangxiense, this work gives scientists a powerful reference manual for a promising medicinal vine. With this blueprint in hand, researchers can more easily pinpoint genes involved in valuable compounds, guide breeding programs to enhance desired traits, and compare G. guangxiense with related species to clarify their relationships. Just as importantly, the genome supports better conservation planning at a time when overharvesting threatens some members of the genus. In short, this detailed genetic map transforms a little-known forest climber into a well-charted resource for future medicine, agriculture and biodiversity protection.
Citation: Zhang, X., Zhang, H., Chen, C. et al. A high-quality Chromosome-level genome assembly of Gynostemma guangxiense (Cucurbitaceae). Sci Data 13, 503 (2026). https://doi.org/10.1038/s41597-026-06889-x
Keywords: medicinal plants, plant genome, Gynostemma, genetic resources, conservation biology