Clear Sky Science · en
A Unified Dataset for Antibody and Nanobody Design Including Sequence, Structure, and Binding Affinity Data
Why tiny immune tools and big data matter
Antibodies and their smaller cousins, nanobodies, are the body’s precision-guided missiles against infections and cancer. Drug developers now try to design these molecules on computers, much as engineers design aircraft. But until recently, the raw material for such artificial-intelligence design—reliable data about antibody components, shapes, and how tightly they bind their targets—was scattered across many incompatible databases. This article introduces the Antibody and Nanobody Design Dataset (ANDD), a unified, public resource built to give researchers the clean, comprehensive data they need to craft the next generation of targeted therapies.
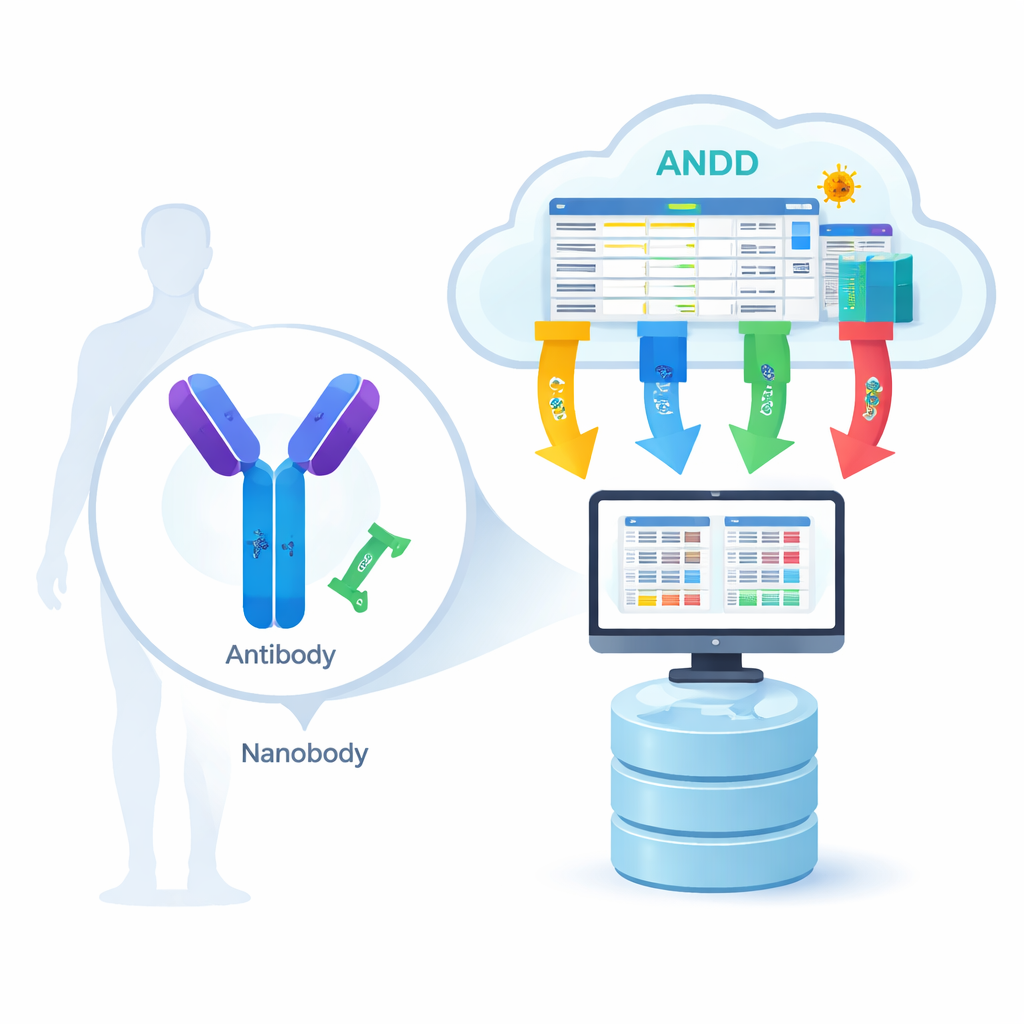
From biological lock-and-key to digital blueprint
Antibodies are large Y‑shaped proteins, while nanobodies are much smaller single‑piece versions found in animals such as llamas and alpacas. Both recognize specific “locks” on viruses, cancer cells, or other disease‑related proteins. For computer models to learn how this recognition works, they need four kinds of information for many different examples: the amino‑acid sequence (the parts list), the 3D structure (the shape), the antigen (the target), and the binding strength (how tightly the two stick together). Until now, most resources captured only one or two of these pieces at a time, forcing scientists to hop between databases and manually patch things together, which slowed progress and introduced errors.
Bringing scattered pieces into one organized library
The ANDD team gathered data from 15 major sources, including dedicated antibody and nanobody databases, general protein repositories, and even patent documents. They then passed these raw inputs through a carefully scripted pipeline: downloading, reformatting into a shared schema, cross‑checking identifiers, removing duplicates, and harmonizing naming rules. When different databases disagreed, curated sources and direct experiments were prioritized. The end result is a single table plus a set of structure files that connect sequence, structure, target, and binding information in a consistent way, with each record tagged so users can trace exactly where it came from and how it was processed.
Layered detail for different research needs
Not every entry in ANDD is equally rich, so the authors organized the collection into layers of increasing detail. At the broadest level, there are 48,683 antibody and nanobody entries with sequence information. A large subset adds 3D structures, and a smaller subset further includes the sequence of the target proteins. The most detailed layer—thousands of entries—adds measured or predicted binding strength. For antibodies, for example, 18,464 entries have sequences, the same number combine sequence and structure, over 8,000 also include antigen sequences, and 7,737 have full sequence, structure, antigen, and affinity data. A parallel hierarchy exists for nanobodies, giving both experimentalists and model builders flexibility: they can choose large, simple datasets or smaller, more information‑rich subsets.
Filling in the blanks on binding strength
Binding strength is crucial for drug design, but experimental values are scarce and unevenly reported. To address this gap without blurring the line between data and prediction, the authors used a specialized deep‑learning tool, ANTIPASTI, to estimate binding strength only for entries that had structures but lacked measurements. These 2,271 predicted values are clearly labeled and kept separate from the roughly 7,000 experimentally measured ones. The team then checked overall consistency using another model, AlphaBind, and by comparing mathematically related measures of binding. Strong correlations and low error suggested that the curated experimental values are reliable, and that the predicted values follow sensible trends without being treated as ground truth.
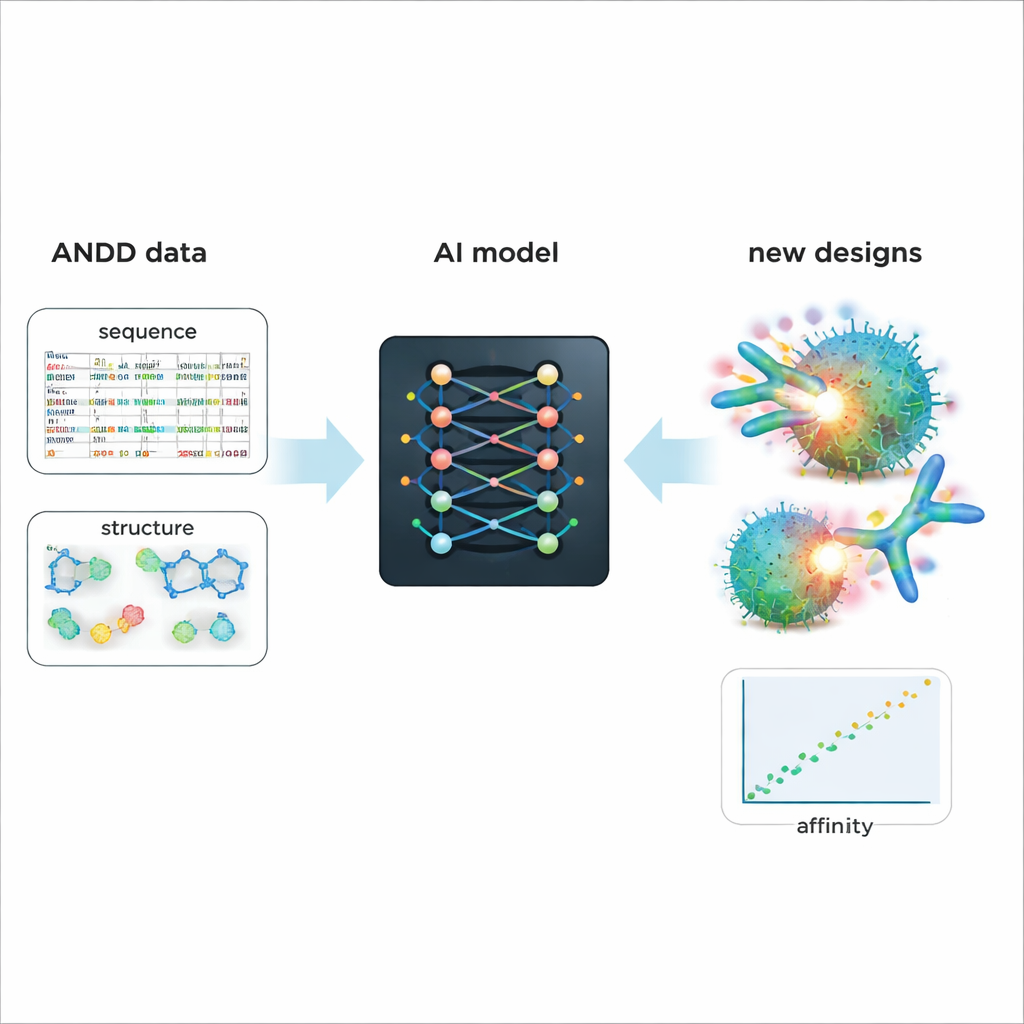
Powering smarter design of future medicines
To demonstrate ANDD’s practical value, the authors fine‑tuned an existing generative AI model that designs antibodies and nanobodies. Training on ANDD’s combined sequence, structure, target, and affinity information led to generated molecules with better predicted binding and more realistic shapes than a baseline model trained on older, simpler data. Beyond this case study, ANDD is openly available under a permissive license, comes with full documentation and a reproducible build pipeline, and is designed to be updated regularly. For non‑specialists, the key message is that ANDD turns a messy patchwork of antibody data into a coherent, trustworthy library—giving AI tools a far better starting point for designing precise, more effective biologic drugs.
Citation: Wu, Y., Liu, X., Hrovatin, K. et al. A Unified Dataset for Antibody and Nanobody Design Including Sequence, Structure, and Binding Affinity Data. Sci Data 13, 295 (2026). https://doi.org/10.1038/s41597-026-06878-0
Keywords: antibody design, nanobodies, binding affinity, biologic therapeutics, AI drug discovery