Clear Sky Science · en
A chromosomal haplotype-resolved genome assembly of Cuphea hookeriana
From Garden Shrub to Genetic Blueprint
Cuphea hookeriana is a small evergreen shrub best known for its bright flowers in gardens and hedges. But behind its showy blossoms lies a treasure trove of useful oils and natural chemicals that could support greener fuels, cosmetics, and industrial products. This study turns C. hookeriana from a pretty plant into a powerful scientific resource by building a highly detailed map of its DNA, offering a foundation for future breeding, ecology, and evolution research.
A Colorful Plant with Hidden Value
Cuphea hookeriana is native to tropical and subtropical regions, including Mexico, and is popular in landscaping for borders and ground cover. Its seeds are unusually rich in medium-chain fats, similar to those found in coconut and palm oil. These fats are valuable as ingredients for bio-based soaps, detergents, lubricants, and biodiesel, and they also support the growing market for plant-based cosmetics. The plant’s varied flower shapes and colors, including distinctive spur-like petals thought to reflect long-term partnerships with bees, birds, and moths, make it a favorite model for studying how plants and pollinators shape one another over time.
Why a High-Quality Genome Matters
Despite its economic and scientific promise, research on Cuphea has been held back by the lack of a complete reference genome—a trusted master copy of the species’ DNA. The challenge is made greater because C. hookeriana is triploid, meaning it carries three sets of chromosomes instead of the usual two. When those sets are very similar, it is hard for computers to untangle which piece of DNA belongs to which copy. A clear, chromosome-level genome makes it far easier to track genes that influence oil content, flower form, stress tolerance, and other traits, and it also allows scientists to compare Cuphea with related species to trace its evolutionary path.
Building the DNA Map, Layer by Layer
The researchers started with leaves from a single, vegetatively propagated plant and extracted its DNA using a refined laboratory protocol. They first used short-read DNA sequencing to estimate the overall genome size and to confirm that the plant carried three chromosome sets. They then turned to long, highly accurate "HiFi" DNA reads and a technique called Hi-C, which captures how pieces of DNA sit next to each other in the cell nucleus. Together, these approaches allowed them to assemble the genome from millions of fragments into long stretches that matched full chromosomes, and to separate the DNA into two distinct haplotypes—two slightly different versions of the genome present in the triploid plant.
What the New Genome Reveals
The team assembled 16 chromosomes totaling just under 500 million DNA letters, divided into two haplotypes labeled A and B. Each haplotype contained about 30,000 genes, and independent quality checks showed that over 97% of expected plant genes were present and correctly assembled. They also cataloged repeated DNA elements, which made up nearly 38% of the genome and were dominated by a common class of plant repeats known as long terminal repeat retrotransposons. Some of the third chromosome set, which should have produced a second B-like haplotype, could not be fully separated because it was nearly identical to the first, leading to a "collapsed" view of those extra copies—an expected difficulty in complex plant genomes.
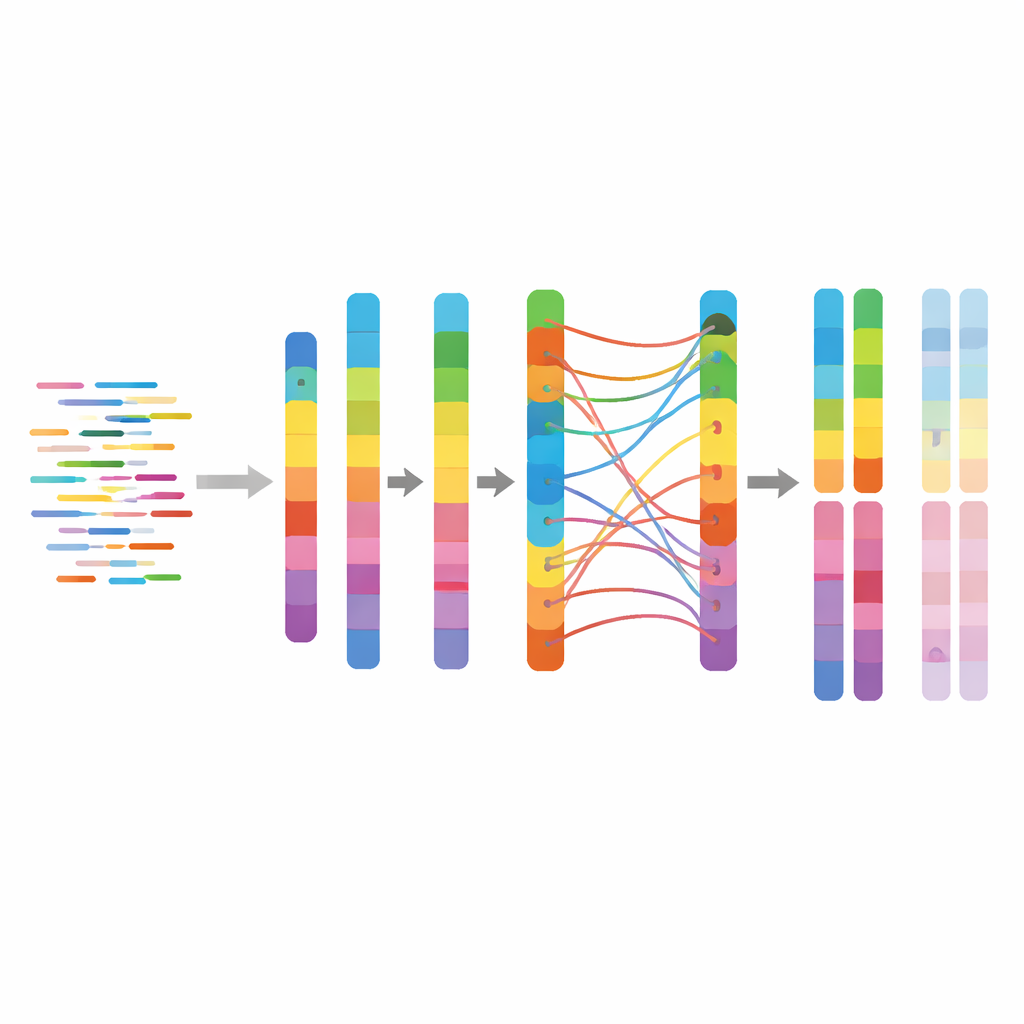

A Lasting Resource for Breeding and Discovery
All of the raw data, the finished genome assembly, and the gene and repeat annotations have been made publicly available in major sequence and data archives. For non-specialists, the key message is that C. hookeriana now has a solid genetic reference, much like a detailed road atlas, that other researchers can use and build upon. This resource will speed efforts to develop new varieties for ornamental planting and sustainable oil production, and it will enable deeper studies of how its striking flowers and ecological roles evolved. In short, the study transforms an attractive garden plant into a well-charted genetic model for future science and innovation.
Citation: Gu, C., Wang, J., Zhang, G. et al. A chromosomal haplotype-resolved genome assembly of Cuphea hookeriana. Sci Data 13, 445 (2026). https://doi.org/10.1038/s41597-026-06830-2
Keywords: Cuphea hookeriana, plant genome assembly, ornamental oilseed, triploid chromosomes, haplotype-resolved DNA