Clear Sky Science · en
Near complete 12S DNA reference library for the freshwater fish of French Guiana, northern Amazonian region
Why Hidden Fish Matter
Rivers in the Amazon’s northern fringe are home to an astonishing variety of fish, many of which are rarely seen by people and difficult for scientists to study. Yet these species are vital for food webs, fisheries, and the health of some of the world’s last great rainforests. This study tackles a surprising obstacle to understanding that underwater life: not a lack of high‑tech tools, but a missing “phone book” that links anonymous snippets of DNA to actual fish species. By building an almost complete DNA reference library for the freshwater fishes of French Guiana, the authors unlock powerful new ways to monitor biodiversity, track human impacts, and guide conservation across the wider Amazon region.
Reading Rivers Through Traces
Instead of catching every fish, modern surveys can now “read” rivers by filtering water and analyzing the tiny fragments of genetic material animals leave behind, known as environmental DNA or eDNA. When these fragments are amplified and sequenced, they can be matched against a reference library to reveal which species are present. However, in megadiverse tropical systems, most species still lack suitable DNA entries, and commonly used genetic markers are often too long or not specific enough for waterborne traces. For fishes, a short section of a gene called 12S has emerged as a sweet spot: it is compact enough to be recovered from degraded DNA, yet usually distinct enough to tell closely related species apart—if a good reference library exists.
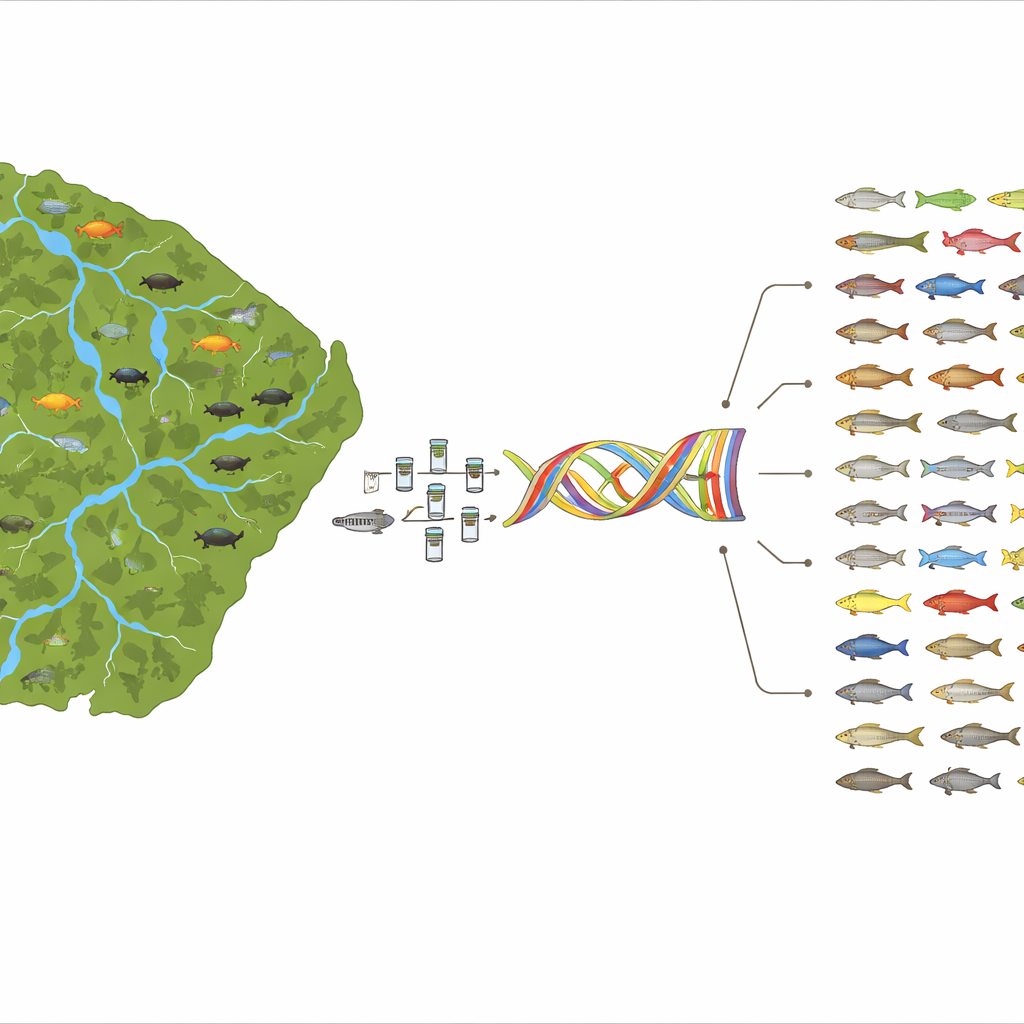
Building a Genetic Catalog of a Wild Frontier
French Guiana, a largely forested territory on the Guiana Shield between Brazil and Suriname, is crisscrossed by eight major river basins and extensive swamps. These waters shelter at least 415 freshwater fish species, about a quarter of them found nowhere else. Over 15 years and more than 25 expeditions, the team sampled fishes throughout these rivers using nets, traps, and other standard techniques. For each captured specimen, they took a small fin clip, preserved it in ethanol, and later extracted its DNA in the laboratory. They also tapped museum collections in Geneva to fill gaps for species that had not been recently collected in the field. In total, they obtained 12S gene sequences for 1,557 specimens, representing 369 species across 51 families and all 16 freshwater fish orders known from French Guiana—coverage of nearly 89% of the region’s freshwater fish fauna.
Checking Names and Faces in the DNA
Turning this trove of sequences into a reliable reference required careful quality control. Fish experts first identified each specimen using field guides and, when necessary, specialist taxonomists. The researchers then checked whether each species matched its known geographic range in French Guiana’s rivers to avoid confusing look‑alike species. Finally, they built a DNA‑based family tree to see whether specimens assigned to the same species clustered together as expected. Any sequences that fell outside their expected group—suggesting mislabeling, contamination, or unresolved species boundaries—were discarded. The team then trimmed the full 12S sequences down to the specific short regions targeted by two widely used eDNA primer sets, often called Teleo1 and 12S‑V5. They tested how well these shorter “barcodes” could distinguish species, finding that Teleo1 could assign about 95% of sequences to a species, while 12S‑V5 resolved roughly 83% to that level, with the remainder reliably placed at genus or family.
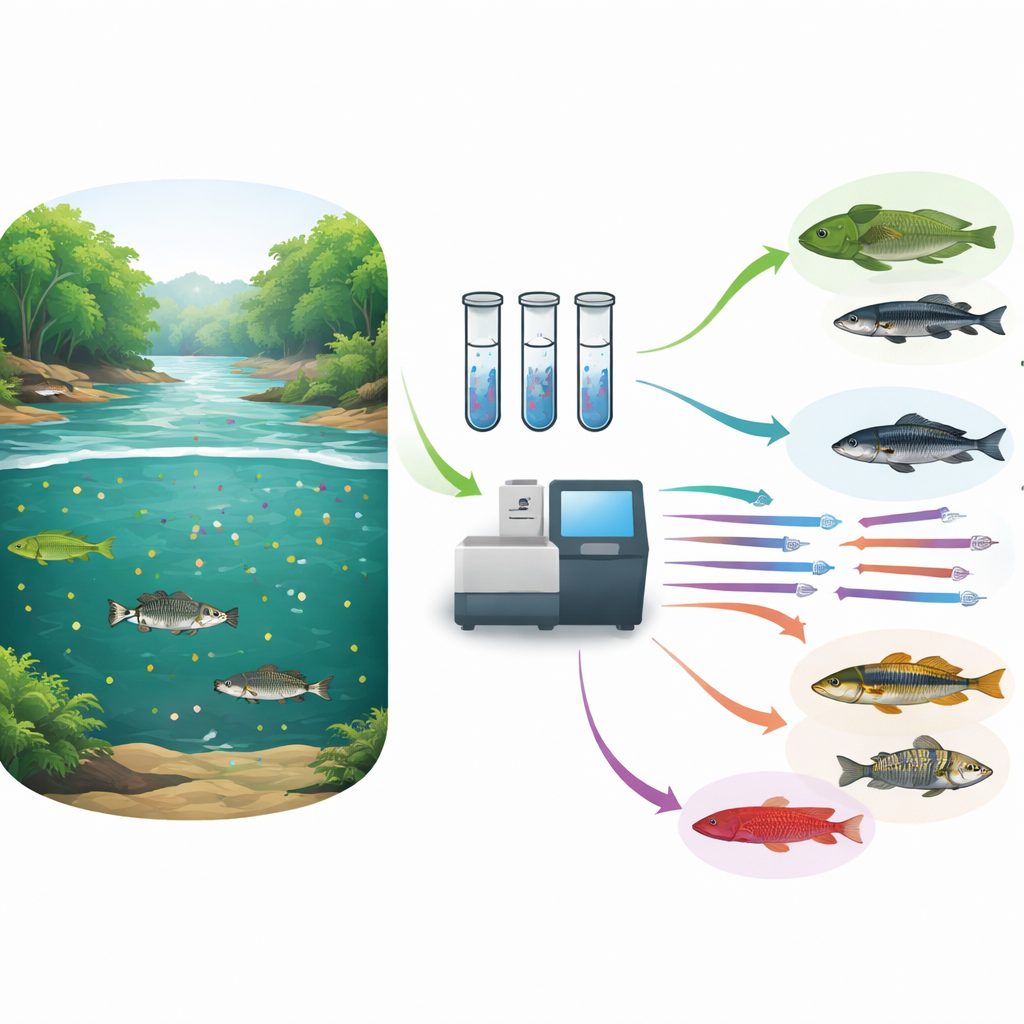

From Local Rivers to a Continental View
Although the library was built for French Guiana, many of its species and lineages are shared with rivers across tropical South America. By comparing their entries with a continent‑wide database of freshwater fish occurrences, the authors show that their 12S sequences cover about 6% of all described Neotropical freshwater fish species but a far larger fraction of higher groups: roughly 23% of genera, 56% of families, and 43% of orders. This means that even when an exact species match is not available outside French Guiana, eDNA surveys can still reliably identify which broader lineages are present, allowing scientists to measure patterns of functional diversity—how different kinds of fish contribute to ecosystem roles—over a vast region.
Why This Library Changes the Game
For a general reader, the key message is that the authors have built the missing reference book that lets scientists translate anonymous DNA traces in Amazonian waters into real fish species. Their near‑complete 12S library for French Guiana makes it possible to rapidly inventory fish communities from water samples, including rare, elusive, or declining species, and to monitor how activities such as deforestation and gold mining are reshaping river life. Beyond one small territory, the library strengthens eDNA studies throughout the Neotropics by anchoring many fish groups in a well‑vetted genetic framework. In a region where traditional surveys are costly and logistically daunting, this resource turns rivers themselves into readable archives of biodiversity, improving our chances of detecting change in time to protect these hidden freshwater worlds.
Citation: Brosse, S., Cuenot, Y., Condachou, C. et al. Near complete 12S DNA reference library for the freshwater fish of French Guiana, northern Amazonian region. Sci Data 13, 408 (2026). https://doi.org/10.1038/s41597-026-06811-5
Keywords: environmental DNA, freshwater fish, French Guiana, biodiversity monitoring, genetic reference library