Clear Sky Science · en
HMI-LUSC: A Histological Hyperspectral Imaging Dataset for Lung Squamous Cell Carcinoma
Seeing Cancer in New Colors
Lung cancer remains one of the world’s deadliest diseases, in part because spotting every last cancer cell on a microscope slide is difficult and time‑consuming. Pathologists usually rely on pink‑and‑purple stained tissue viewed under a microscope, a method that captures structure but misses subtle chemical clues. This paper introduces HMI‑LUSC, the first openly available collection of microscope images of lung squamous cell carcinoma captured not just in three colors, but in dozens of narrow color bands, giving computers and clinicians a far richer view of what makes tumor cells different from their healthy neighbors.
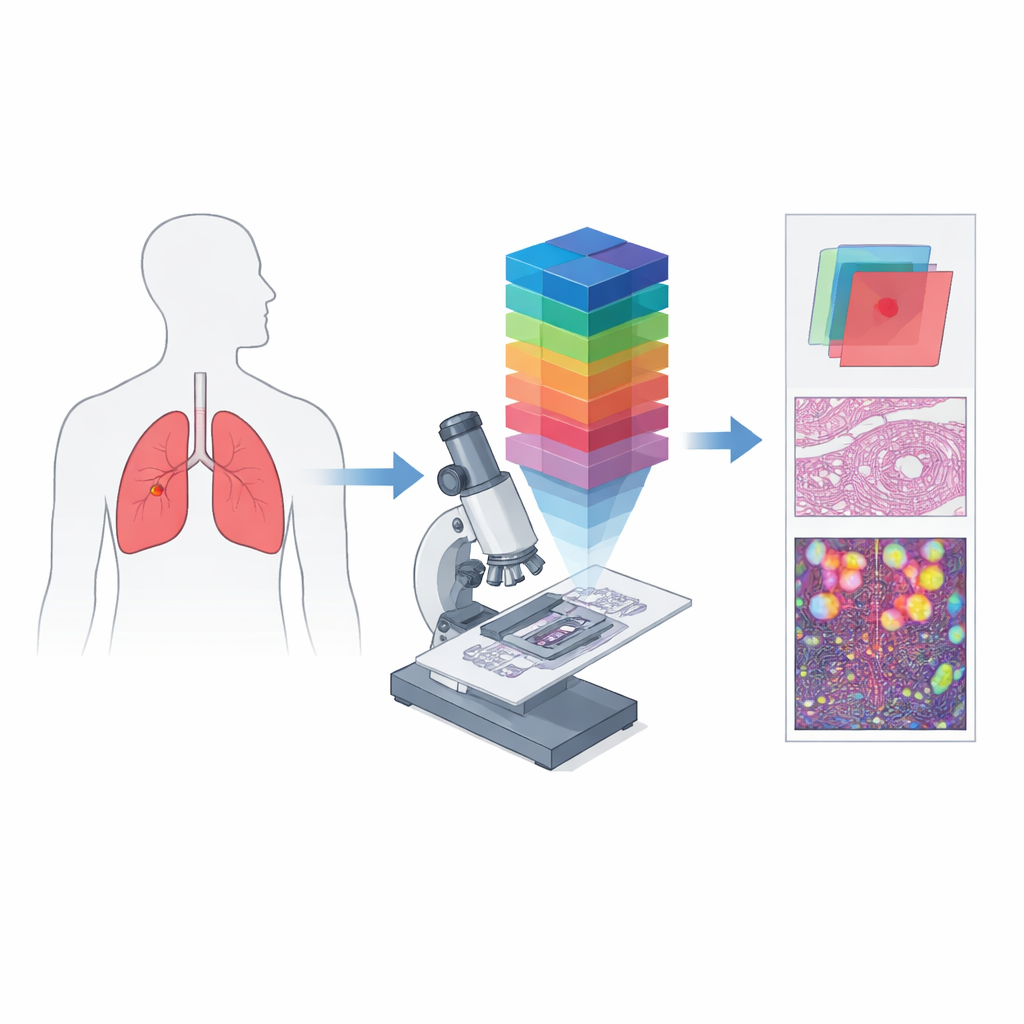
From Simple Color Pictures to Spectral Fingerprints
Conventional digital pathology works much like a phone camera: it records red, green, and blue channels to approximate what the eye sees. Hyperspectral imaging goes several steps further by splitting light into many closely spaced wavelengths, producing a three‑dimensional “data cube” in which every tiny spot of tissue has its own detailed color spectrum. When this idea is combined with a microscope, it becomes hyperspectral microscopic imaging, able to capture both fine structure and rich spectral information at the level of individual cells. Such data can reveal differences in how tissues absorb and reflect light that are invisible in standard images, creating unique spectral “signatures” for cancerous and non‑cancerous regions.
Building a New Library for Lung Cancer Study
The authors created HMI‑LUSC to fill a clear gap: before this work, there was no public hyperspectral dataset for lung cancer slides, making it difficult to test and compare computer‑based diagnostic methods. They collected tissue from ten patients undergoing lung tumor surgery, prepared standard hematoxylin‑and‑eosin slides, and scanned them at high resolution. Experienced pathologists marked tumor and normal areas, and representative regions were re‑imaged with a custom‑built hyperspectral microscope. Each resulting image covers a small patch of tissue but spans 61 wavelengths between 450 and 750 nanometers, at a resolution of 3088 by 2064 pixels. For every region, the dataset includes the raw spectral cube, a conventional RGB rendering, and masks that outline where tumor tissue is present.
Turning Rough Outlines into Cell‑Level Maps
While slide‑level markings are useful, training modern algorithms often requires information at the level of individual cells. Manually tracing every cell is impractical, so the team designed a semi‑automatic workflow. First, they grouped pixels into clusters based on their spectral similarity using a standard computer‑vision method. Then pathologists inspected these clusters overlaid on the tissue image and assigned them to four categories: tumor cells, non‑tumor cells, non‑cell tissue such as stroma or blood, and empty background. A second pathologist reviewed and adjusted these results, with disagreements resolved by consensus. The outcome is a set of detailed pixel‑wise masks that capture subtle mixtures of cell types and confusing border zones, providing much richer teaching material for machine‑learning systems.
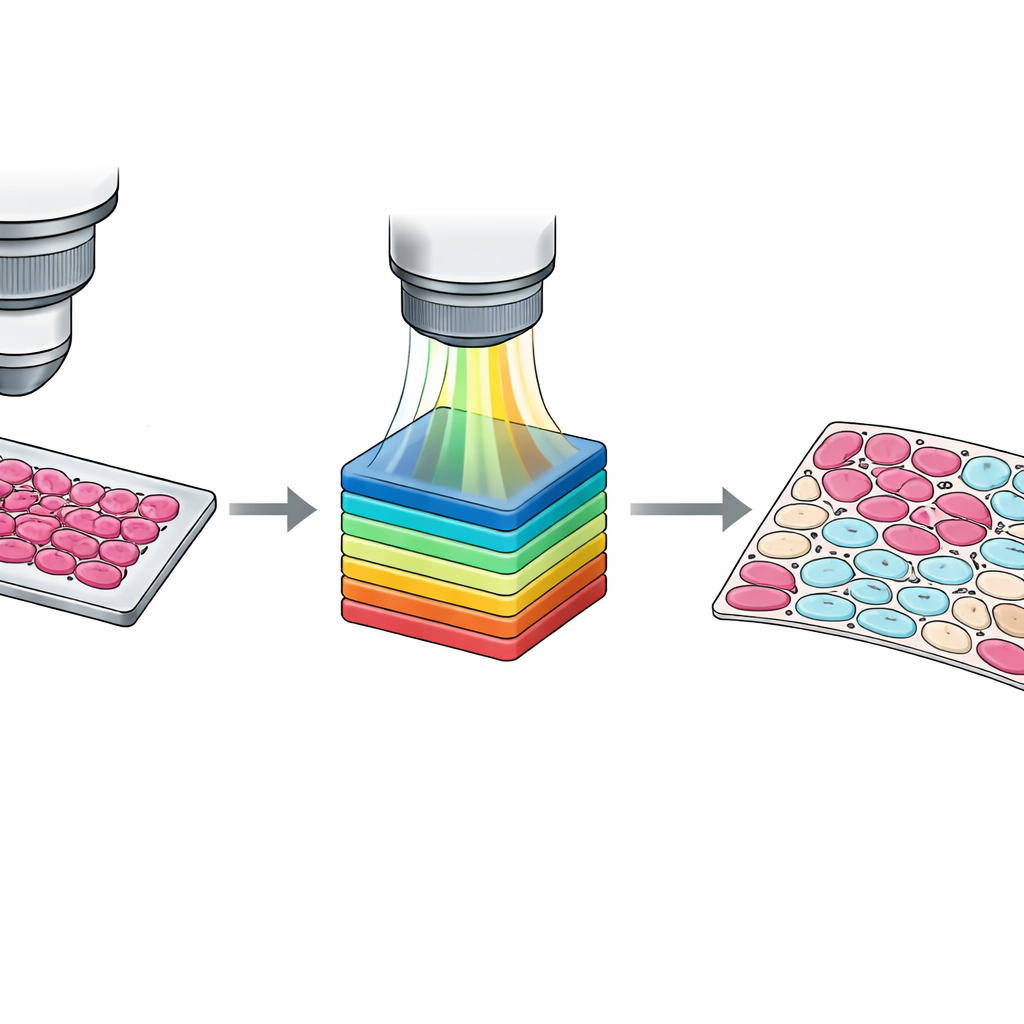
Ensuring Sharp and Reliable Data
To make the dataset trustworthy, the authors thoroughly tested their imaging system. They verified that the microscope can resolve fine patterns down to about one micron—small enough to distinguish individual cells—and that image noise is low across most wavelengths. They also compared the measured spectrum of a standard light source with reference curves and with a commercial hyperspectral camera, finding excellent agreement. Finally, they demonstrated how the data can be used by running baseline computer models, from classic machine‑learning methods to simple deep‑learning networks, to segment tumor regions. Even without heavy optimization, these models achieved solid accuracy, showing that the dataset is well suited as a benchmark for future methods.
What This Means for Future Lung Cancer Care
HMI‑LUSC does not replace large collections of standard slides, nor is it yet a clinical tool on its own. Instead, it offers researchers a carefully curated window into how lung tumor cells differ from nearby tissue across many wavelengths of light. By making these data, labels, and code openly available, the authors provide a common testbed for developing and comparing algorithms that use spectral information, from simple classifiers to advanced neural networks. In the long run, such work could help computers assist pathologists in spotting tumors more accurately and quickly, and may reveal spectral patterns linked to tumor type or treatment response that ordinary images cannot show.
Citation: Yan, Z., Huang, H., Guo, Y. et al. HMI-LUSC: A Histological Hyperspectral Imaging Dataset for Lung Squamous Cell Carcinoma. Sci Data 13, 415 (2026). https://doi.org/10.1038/s41597-026-06766-7
Keywords: hyperspectral imaging, lung cancer, digital pathology, tumor segmentation, medical imaging dataset