Clear Sky Science · en
Multidimensional transcriptome dataset for systematic evaluation of Jakyakgamcho-tang-induced cell signatures
Why this ancient remedy matters today
Many people turn to traditional herbal medicines for muscle cramps, pain, or general wellness, but how these age‑old remedies affect our cells is still largely a mystery. This study focuses on Jakyakgamcho‑tang, a simple two‑herb formula long used in East Asia, and asks a modern question: how do different ways of mixing and brewing the herbs change what they do to our cells? By mapping how genes in human and animal cells respond to many versions of this remedy, the researchers created a detailed reference that can help scientists test, compare, and refine herbal treatments with the same rigor used for conventional drugs.
A simple two‑herb formula under the microscope
Jakyakgamcho‑tang is made from just two ingredients: Paeoniae Radix (from peony root) and Glycyrrhizae Radix et Rhizoma (from licorice root). Traditionally, it is prescribed for muscle spasms and pain, and more recent work suggests it may also help with muscle wasting, inflammation, and memory problems. Yet the remedy is not always prepared the same way. Healers may use more peony or more licorice, boil the herbs in water or extract them with alcohol, and either cook them together or make separate extracts and then mix them. Each of these choices can change which plant chemicals end up in the final cup, and thus how the body responds. The authors set out to capture these differences in a systematic, data‑rich way.
Designing a broad and careful testing ground
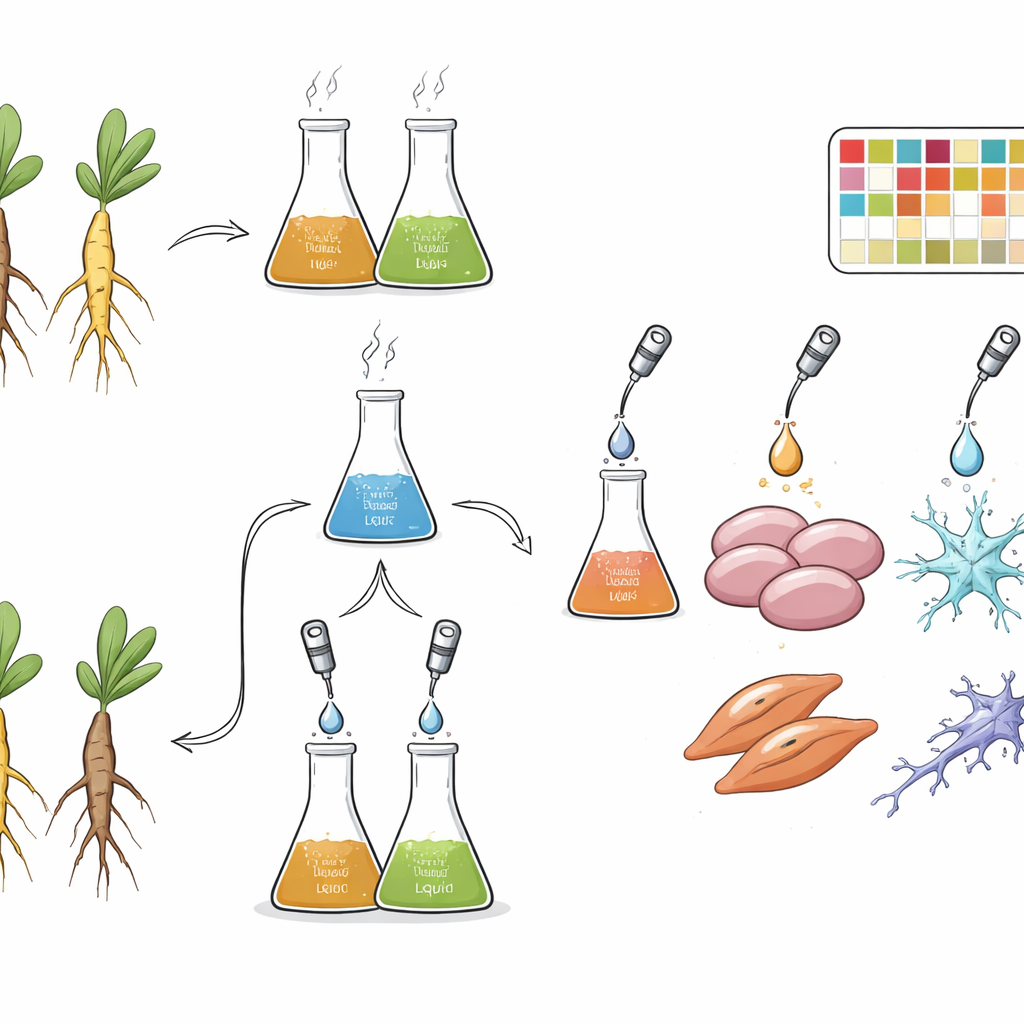
To do this, the team prepared many versions of the formula. They tested three peony‑to‑licorice ratios (more peony, equal parts, or more licorice), two solvents (plain hot water or 70% ethanol), and two preparation methods. In the combined method, the herbs were mixed first and then extracted together, allowing their chemicals to interact during boiling or sonication. In the individual method, each herb was extracted on its own and the dried extracts were mixed only afterward, favoring standardization and preservation of delicate compounds. The researchers also measured key marker chemicals using high‑performance liquid chromatography, confirming that peony‑related compounds increased in peony‑rich mixtures and licorice‑related compounds in licorice‑rich mixtures, and that ethanol generally pulled out higher amounts of many ingredients than water.
Listening to cells in liver, muscle, and nerve
Next, the scientists exposed three well‑established cell types to these different extracts: liver‑like HepG2 cells, muscle‑like C2C12 cells, and nerve‑like PC12 cells. These lines were chosen because they are closely connected to conditions where the formula is used, such as muscle spasms, pain, and broader metabolic or nervous‑system problems. Careful dose‑finding ensured that the concentrations used disrupted cell activity without killing the cells, so that gene responses would reflect pharmacological effects rather than simple toxicity. Each condition was tested in triplicate, and more than 500 RNA samples were sequenced, ultimately yielding 513 high‑quality gene expression profiles that record how thousands of genes in each cell type respond to each preparation of the herbal formula.
Ensuring trustworthy and reusable data
Because this work is meant as a shared resource, the team devoted significant effort to quality checks. The herbs were authenticated by experts and DNA barcoding, and their chemical make‑up was documented. RNA from treated cells was examined for purity and integrity, and sequencing reads showed high quality scores and strong mapping to the appropriate reference genomes. Replicate experiments were highly consistent, with very strong correlations between repeat samples across species and batches. The team also compared pathway‑level patterns from their data to an independent drug‑response resource called the Connectivity Map. For three well‑known drugs used as controls, the gene activity patterns in this new dataset matched those from the external database much more closely when the same drug was used, supporting the reliability and broader relevance of the measurements.
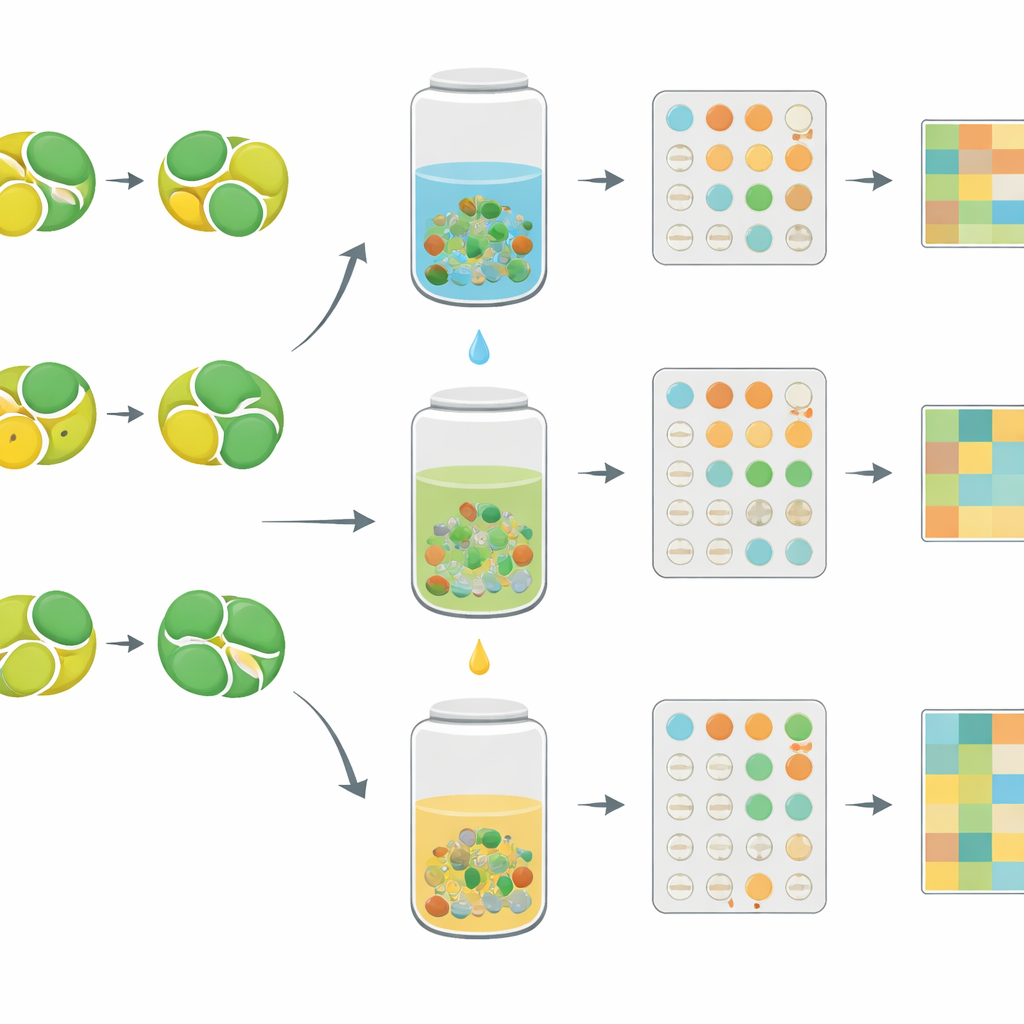
From traditional recipes to data‑driven refinement
All raw and processed gene expression data, along with detailed preparation and chemical information, have been deposited in public databases where other researchers can freely explore them. In everyday terms, this study turns many slight variations of a traditional two‑herb tea into a searchable “fingerprint” library of how cells react at the genetic level. This makes it possible to ask, for example, which mixing ratio and solvent best support muscle‑protective responses while limiting inflammatory signals, or which preparation most closely mimics the action of a known drug. By building such bridges between long‑used remedies and modern molecular tools, the work lays a foundation for more precise, evidence‑based optimization of herbal medicines.
Citation: Baek, SJ., Lee, H., Park, SM. et al. Multidimensional transcriptome dataset for systematic evaluation of Jakyakgamcho-tang-induced cell signatures. Sci Data 13, 367 (2026). https://doi.org/10.1038/s41597-026-06759-6
Keywords: herbal medicine, gene expression, RNA sequencing, drug response data, Jakyakgamcho-tang