Clear Sky Science · en
Metabolomic and Transcriptomic Profiling of Two Closely Related Species within the Genus Oldenlandia
Why these humble herbs matter
Two small weedy plants used in traditional Asian medicine, Oldenlandia diffusa and Oldenlandia corymbosa, are widely taken for problems ranging from inflammation to liver disease and cancer. Yet, despite their long history in herbal remedies and their frequent mixing in the marketplace, we still know surprisingly little about how they actually differ inside: which chemical ingredients they make, in which plant parts, and which genes control those ingredients. This study opens up that hidden world by providing a detailed map of both the molecules and the gene activity in different tissues of the two species.
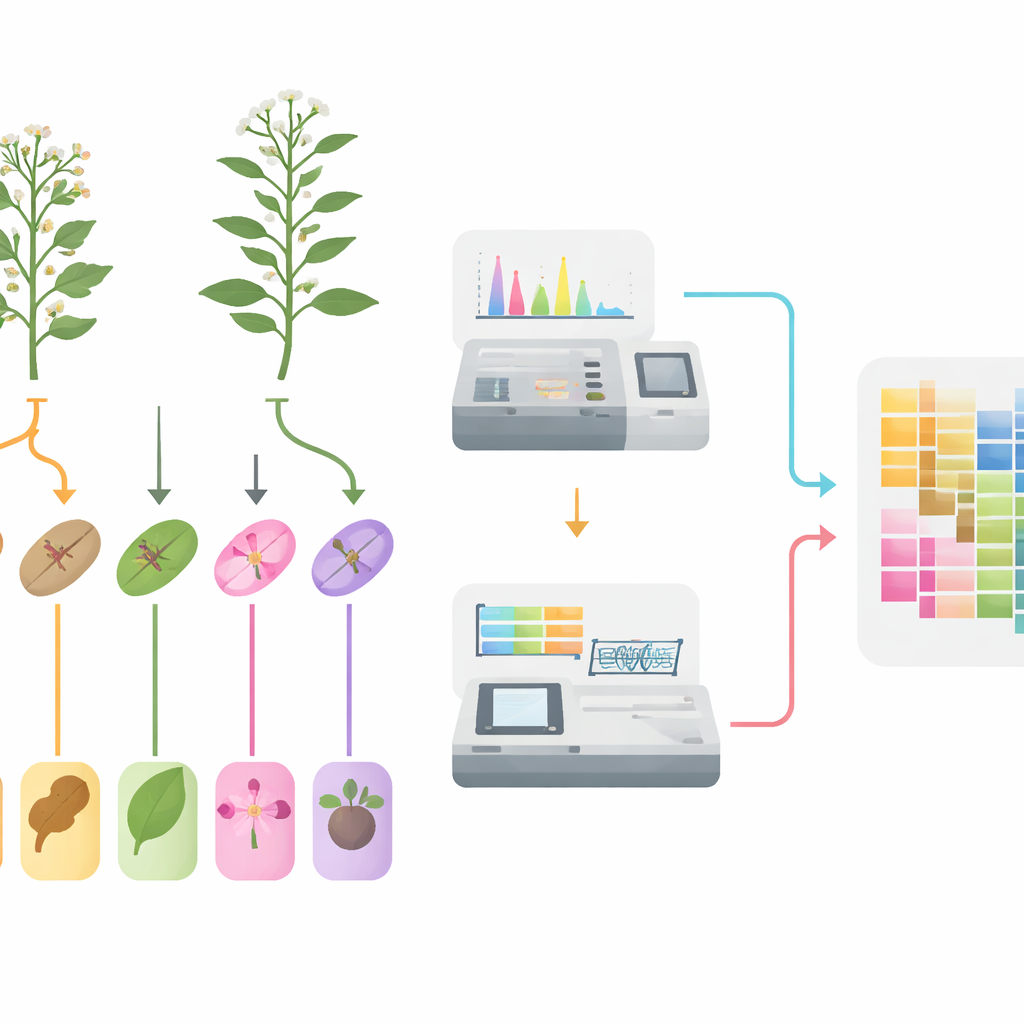
From field to frozen samples
The researchers began by growing both species side by side in the same field conditions to avoid environmental bias. During the flowering period, they carefully separated five tissues—roots, stems, leaves, flowers, and fruits—from multiple plants of each species. To capture a reliable picture of natural variation, they pooled material from many individuals and created six biological replicates for every tissue type. Each tissue sample was then split in two: one part for measuring small molecules, and one part for reading out which genes were switched on.
Taking stock of thousands of plant chemicals
To catalog the plants’ chemical makeup, the team used a sensitive mass-spectrometry–based approach that can track a very large number of known plant metabolites in one run. Across 60 samples they detected 1,342 distinct small molecules, dominated by groups already suspected to be important for medicine: nearly one quarter were flavonoids, followed by phenolic acids, alkaloids, and terpenoids. Statistical analyses showed that tissue samples clustered cleanly by type, and quality control checks confirmed that measurements were stable and reproducible. Comparing the same tissue between the two species revealed numerous molecules that were far more abundant in one species than the other, especially in pathways linked to color pigments, defense compounds, and other specialized plant chemicals.
Reading the plants’ instruction manuals
In parallel, the scientists extracted RNA from the same tissues to see which genes were active, thereby capturing the plants’ internal instructions at the time of sampling. High-throughput sequencing generated over 500 gigabases of high-quality data and led to the identification of 37,644 genes in Oldenlandia diffusa and 20,825 genes in Oldenlandia corymbosa. Again, samples from the same tissue clustered together, underscoring data reliability. By comparing tissues, the team found thousands of genes that behaved differently between roots and above-ground parts, or between stems, leaves, flowers, and fruits. Many of these genes are involved in energy capture, basic metabolism, and enzymes that modify plant compounds—exactly the kinds of activities that would shape where medicinal compounds are made and stored.
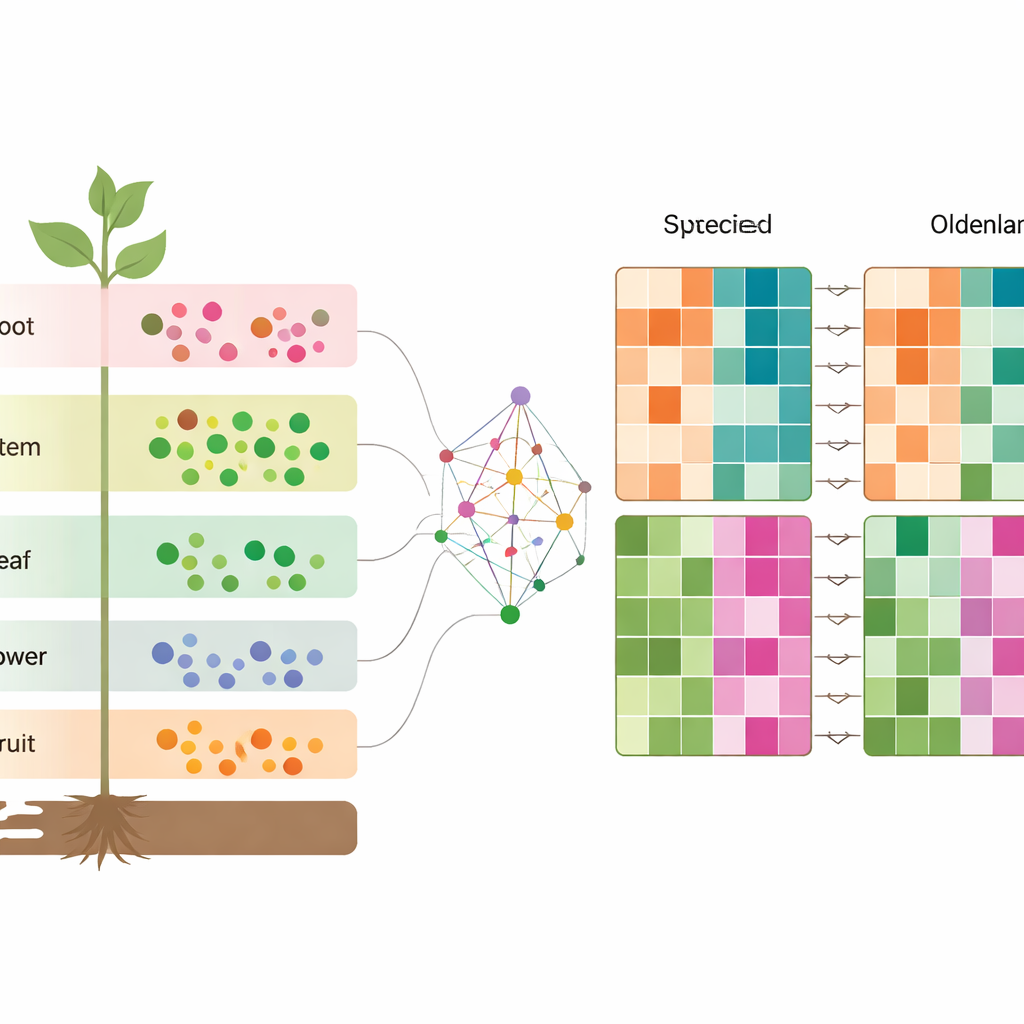
Zooming in on differences between the two species
To understand what truly separates the species, the authors matched genes in O. diffusa and O. corymbosa that are direct counterparts, then compared their activity in each tissue. They discovered thousands of genes turned up or down in one species relative to the other in roots, stems, leaves, flowers, and fruits. These genes fell into pathways that make and process key compounds, including terpenoids, phenylpropanoids, and various nitrogen-containing molecules. When combined with the metabolite data, this reveals a layered picture: each tissue has its own chemical and genetic fingerprint, and those fingerprints shift in characteristic ways between the two closely related herbs.
What this means for medicine and future research
For non-specialists, the main takeaway is that these two look-alike medicinal plants are not interchangeable. Beneath their similar appearance, their roots, stems, leaves, flowers, and fruits differ in both the amounts of important small molecules and the activity of genes that make those molecules. By releasing this large, carefully validated dataset, the study provides a reference atlas that other scientists can mine to pinpoint which ingredients may drive specific health effects, and which genes could be targeted to breed or engineer plants with more consistent and potent properties. Ultimately, this kind of deep mapping can help transform traditional herb mixtures into better-understood, more precisely used medicines.
Citation: Chen, P., Huang, Z., Wen, Y. et al. Metabolomic and Transcriptomic Profiling of Two Closely Related Species within the Genus Oldenlandia. Sci Data 13, 378 (2026). https://doi.org/10.1038/s41597-026-06745-y
Keywords: Oldenlandia, medicinal plants, metabolomics, transcriptomics, plant secondary metabolites