Clear Sky Science · en
Coding and non-coding RNA sequencing during Thalassiosira gravida resting cell formation
How tiny ocean drifters hit pause on life
Diatoms are microscopic algae that drift in the oceans, helping to fuel food webs and absorb carbon dioxide. Like seeds on land, many of these single-celled plants can enter a dormant state to survive dark, cold, or nutrient-poor periods. This study follows one such diatom, Thalassiosira gravida, as it shuts down into a resting stage and later wakes up, revealing a detailed molecular snapshot of how life presses pause without losing the ability to restart.
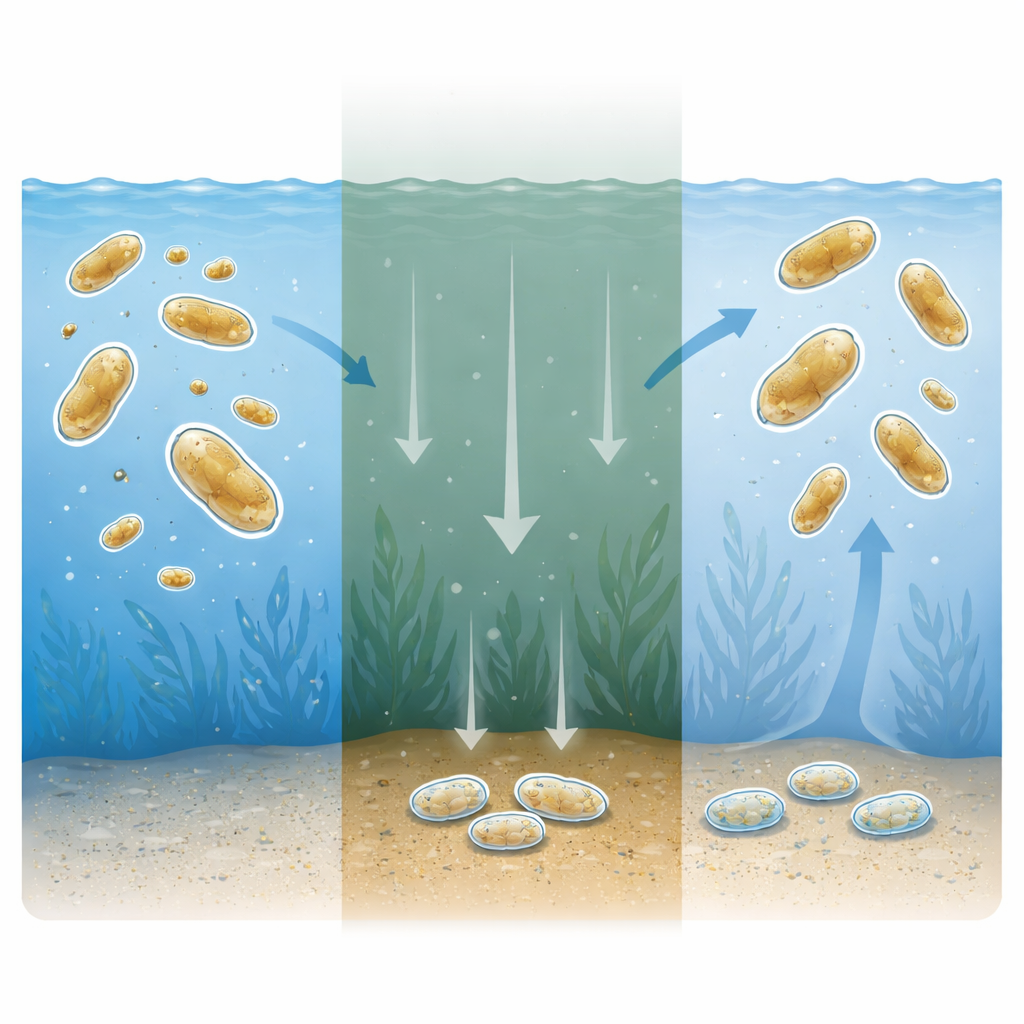
Why sleeping cells matter for the sea
Resting stages in plankton act like underwater seed banks. When conditions turn harsh—such as when nutrients run out—some diatoms transform into long-lasting resting cells that sink toward the seafloor and wait, sometimes for decades. When light and nutrients return, they reactivate, divide, and help spark new blooms. This hidden life cycle stabilizes marine ecosystems, shapes seasonal plankton cycles, and preserves genetic diversity. Yet, despite its ecological importance, we have known surprisingly little about the internal switches that move a diatom from active growth into this quiet, survival mode.
A laboratory model of going to sleep
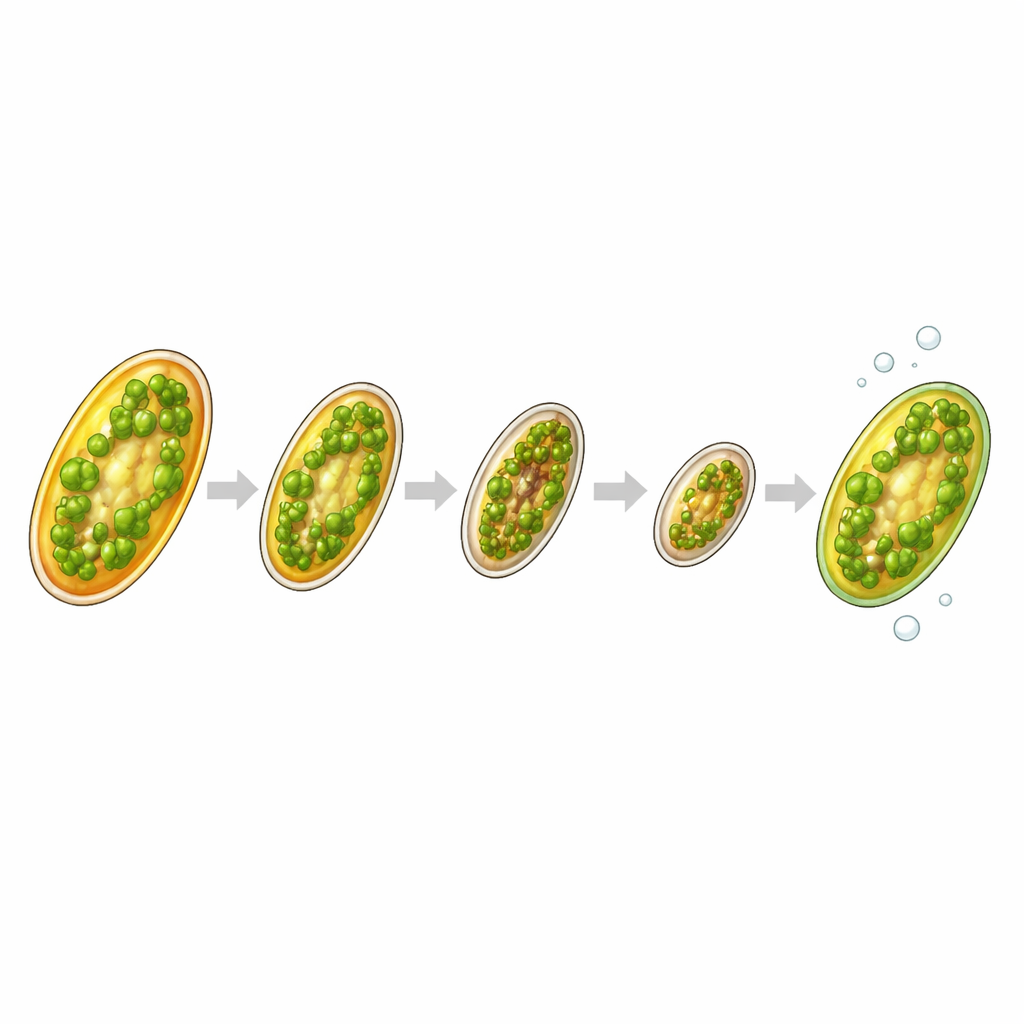
The researchers focused on T. gravida, a widespread diatom known for producing bioactive compounds that can affect small crustaceans and other marine life. In the lab, they grew genetically identical cultures under two conditions: one with normal nutrients and one lacking nitrogen, a key ingredient for growth. Over seven days, nitrogen-starved cells gradually stopped dividing and developed a glassy look, with their green chloroplasts squeezed against the cell wall—clear signs of resting cell formation. A subset of these resting cells was then held in cold, dark conditions for a month to test whether they could truly persist in a dormant state and later reawaken.
Reading the cell’s messages over time
To find out what happens inside the cells during this transition, the team tracked the activity of many types of RNA—the molecules that help carry and control genetic information. They sampled diatoms at four stages: at the start of the experiment, during the early shift toward resting cells, when the resting state was fully established, and after a month of cold, dark storage. For each time point, they sequenced not only the standard protein-coding messages (mRNA) but also long non-coding RNAs and small RNAs, including microRNA-like molecules that can fine-tune gene activity. By comparing patterns between nutrient-rich and nitrogen-deprived cultures, and across time, they assembled a rich, time-resolved view of which genes and regulatory RNAs turn up or down as cells shut down and maintain dormancy.

Reliable data from quiet cells
The authors carefully checked that their cultures behaved as expected. Cell counts showed that nutrient-rich cultures kept growing, while nitrogen-starved ones slowed and stabilized, consistent with entering a quiescent state. When long-stored resting cells were returned to favorable conditions, they resumed growth after a brief adjustment period and regained their normal shape and internal structure, confirming that dormancy was reversible. On the technical side, most sequencing reads were high quality and mapped cleanly to the diatom genome, and samples grouped logically by treatment and time point in statistical analyses. This indicates that the dataset faithfully captures real biological changes rather than experimental noise.
A new map for marine dormancy
Instead of delivering a single mechanism, this work provides a foundational dataset: a detailed catalog of coding and non-coding RNA changes as T. gravida shifts from active growth into a resting state and back. For non-specialists, the key takeaway is that we now have a molecular “movie” of how a common ocean microbe powers down and survives lean times, guided not only by genes that build proteins but also by regulatory RNAs that act more like switches and dimmers. These data are freely available and are expected to guide future studies on how marine microbes endure environmental stress, how their dormant stages shape ocean productivity, and how microscopic life in the sea copes with a changing climate.
Citation: Sepe, R.M., Orefice, I., Di Marsico, M. et al. Coding and non-coding RNA sequencing during Thalassiosira gravida resting cell formation. Sci Data 13, 358 (2026). https://doi.org/10.1038/s41597-026-06744-z
Keywords: diatom dormancy, resting cells, marine phytoplankton, RNA sequencing, nitrogen depletion