Clear Sky Science · en
Genome sequencing, de novo assembly and annotation of the commercially important bamboo, Bambusa tulda Roxb
A Fast-Growing Grass With Big Potential
Bamboo may look like a simple garden plant, but it is actually a powerful natural resource for building, paper, and even future biofuels. One widely used species, Bambusa tulda or Bengal bamboo, grows quickly, stores large amounts of woody material, and flowers only rarely. Until now, scientists lacked a complete “instruction manual” for this species. This article describes how researchers decoded and organized the entire DNA sequence of B. tulda, creating a foundational resource that will help improve bamboo for industry, conservation, and climate-friendly technologies.
Why Decode a Bamboo’s DNA?
Bambusa tulda is common across the Indian subcontinent and parts of Southeast Asia, where its strong culms (stems) are used in rural construction, furniture, and handicrafts. It is also drawing interest as a source of paper pulp and renewable energy. Yet B. tulda behaves in puzzling ways: it can grow very fast, store a lot of tough woody material, and then wait about 50 years before flowering, sometimes all plants in an area doing so at once. Without a full genome sequence, scientists could only guess which genes control these traits. By reading and assembling its DNA, the authors aimed to build a reference map that future researchers can use to study growth, flowering, disease resistance, and more.
Measuring and Reading a Giant Genome
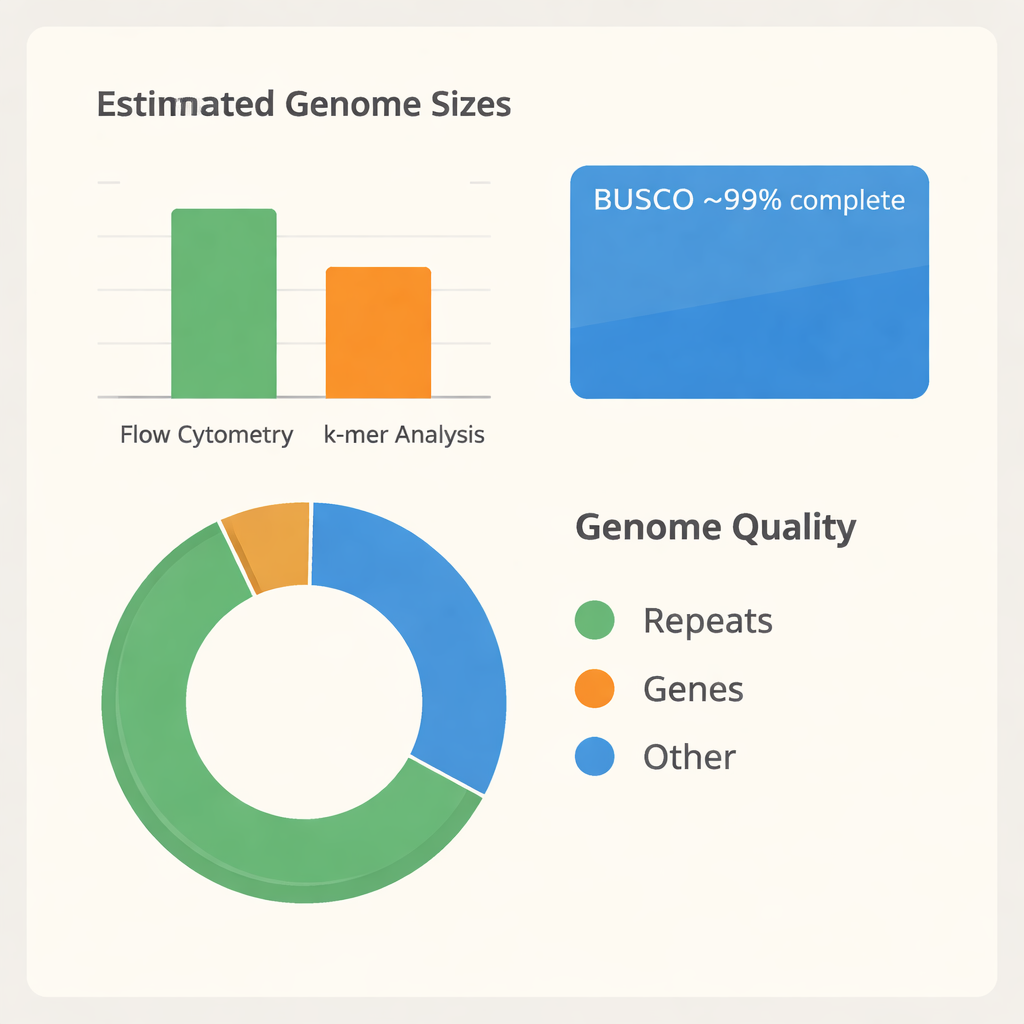
The team first needed to understand how large the B. tulda genome is. Using a technique called flow cytometry, they compared the DNA content of B. tulda leaf cells with that of tomato and maize, two plants whose genome sizes are already known. This suggested a diploid genome size of about 3 billion DNA “letters.” They then used a second, independent approach based on how short DNA fragments overlap (k-mer analysis), which estimated a slightly smaller size of about 2.34 billion letters and revealed that much of the genome is repetitive and likely duplicated. With these measurements in hand, they extracted very long, high-quality DNA from young leaves and sequenced it using advanced PacBio HiFi technology, generating over 116 billion bases of raw data—enough to read the genome dozens of times over.
Piecing Together the Bamboo Blueprint
Turning millions of DNA reads into an ordered genome is like assembling a massive jigsaw puzzle without the picture on the box. The researchers used specialized software to build both a combined primary assembly and two separate haplotypes, reflecting the two parental copies of the genome. After removing duplicate and organelle-derived pieces, they arrived at a streamlined “haploid” assembly of 43 large segments, covering about 1.37 billion bases. These segments fall into three subgenomes, labeled A, B, and C, consistent with B. tulda’s complex, polyploid origin. A widely used quality test called BUSCO showed that about 99% of expected plant genes are present and intact, indicating that the assembly is both complete and reliable for downstream studies.
Genes, Repeats, and Evolutionary Clues
Once the genome was assembled, the next step was to identify its working parts. By combining three lines of evidence—predictions from the DNA sequence itself, similarity to genes from other bamboos, and RNA data from actively expressed genes—the team annotated 56,890 protein-coding genes, which occupy roughly one fifth of the genome. They also cataloged large numbers of non-coding RNAs, including over a thousand transfer RNA and ribosomal RNA genes that support protein production. Strikingly, about two-thirds of the genome consists of repetitive elements, especially mobile DNA segments that copy and move around. These repeats help explain why earlier size estimates differed and point to a dynamic evolutionary history. Comparing protein families across twelve other bamboo species, along with maize and banana as relatives, placed B. tulda firmly among paleotropical woody bamboos with a hexaploid background, confirming that its genome is built from multiple ancestral copies.
A New Foundation for Future Bamboo Research
For non-specialists, the key outcome is that B. tulda now has a high-quality reference genome—an indexed, searchable blueprint of its DNA. This resource will let scientists home in on genes that control rapid growth, woodiness, and delayed flowering, and compare them with those in other grasses. It will also support efforts to breed or engineer bamboo varieties better suited for construction, paper, or energy while preserving natural populations. In short, by charting the genetic landscape of this commercially important bamboo, the study lays the groundwork for smarter use of one of the world’s most versatile plants.
Citation: Kundu, S., Rupp, O., Dey, S. et al. Genome sequencing, de novo assembly and annotation of the commercially important bamboo, Bambusa tulda Roxb. Sci Data 13, 175 (2026). https://doi.org/10.1038/s41597-026-06679-5
Keywords: bamboo genome, Bambusa tulda, plant genetics, woody grasses, renewable biomaterials