Clear Sky Science · en
Large-Scale Histological Image Dataset with Metadata for Colorectal Cancer Microenvironment
Why mapping cancer’s hidden neighborhood matters
When doctors look at a colon tumor under the microscope, they do not just see cancer cells; they see a busy neighborhood of fat, immune cells, connective tissue, and more. This mix of cell types, called the tumor microenvironment, strongly influences how a patient responds to treatment and how long they live. Yet computers that could help doctors make sense of these complex scenes have been held back by a simple problem: they have not had enough well-labeled images to learn from. This study introduces one of the largest and most carefully annotated image collections of colorectal cancer tissue ever assembled, designed specifically to train and test modern artificial intelligence systems.
Building a massive picture library of colon tumors
The researchers created a resource they call HMU-CRC-Hist550K, built from tissue samples of 500 patients treated for colorectal cancer at a major cancer hospital in China. Each patient’s tumor was preserved, stained in the standard way used in pathology labs, and scanned into a high-resolution digital slide. From these slides, the team automatically cut out small square image tiles, each about the size of what a pathologist might see through a microscope at one time. In total, they produced about 550,000 such tiles, giving artificial intelligence models a huge and varied set of examples to learn what different tissues look like.
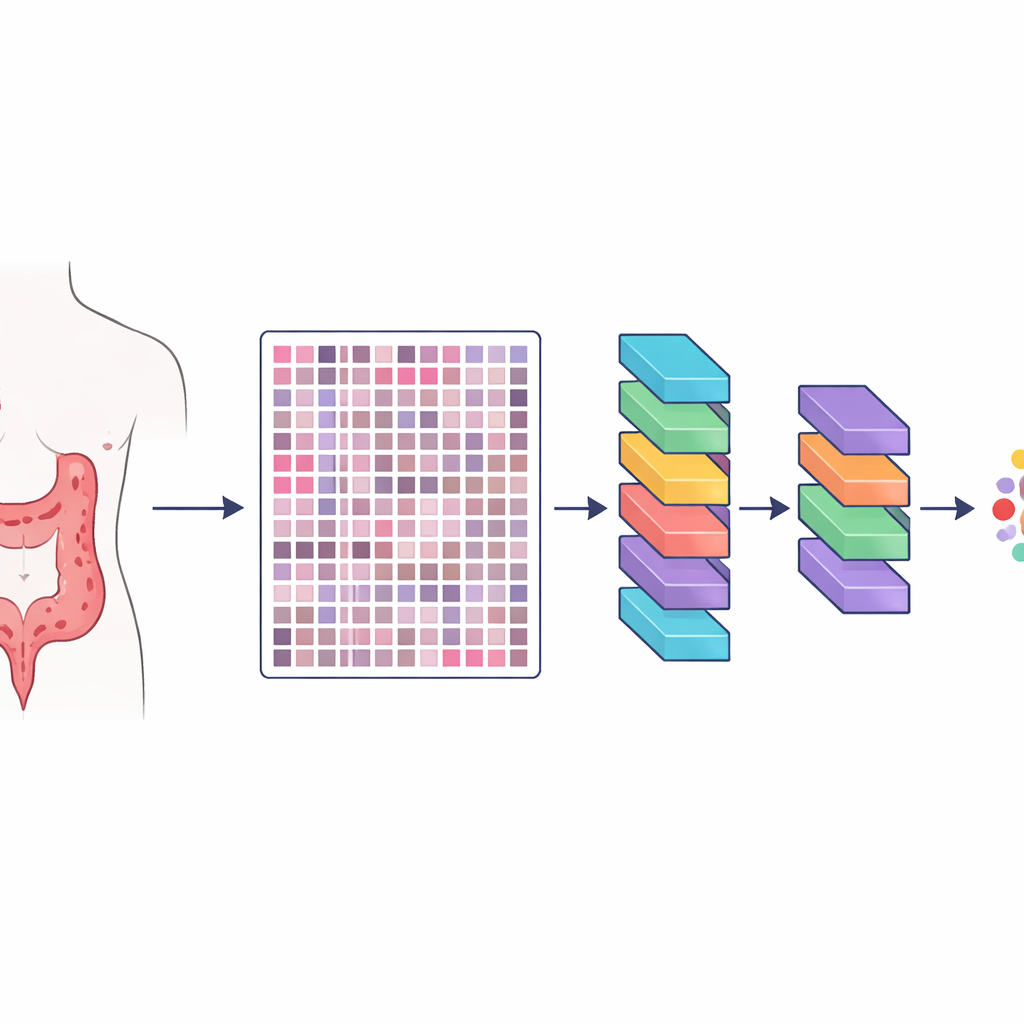
Careful human labeling of the cancer landscape
Creating a big image library is not enough; the images must also be labeled accurately. Three experienced pathologists worked together through a three-step process to mark out eight key components of the tumor’s surroundings: fat tissue, cellular debris, immune cells called lymphocytes, mucus, smooth muscle, normal colon lining, supporting connective tissue around the tumor, and the cancer cells themselves. Two pathologists first drew regions on the large slides independently, and then checked each other’s work. A senior specialist performed a final review, resolving disagreements and excluding unclear areas. This cross-checking greatly reduced personal bias and produced highly consistent labels at fine detail, so that each tile is tied to a specific tissue type within the tumor neighborhood.
Connecting microscope views to patient stories
What makes this dataset especially powerful is that the images are paired with rich clinical information for each patient. For every case, the team collected basic details such as age and sex, as well as tumor stage, tumor location along the colon and rectum, how abnormal the cancer cells looked, whether nerves or lymph nodes were invaded, and how long the patient survived after treatment. They also recorded results of common lab tests that reflect the tumor’s genetic and protein makeup. All personal identifiers were removed so that patients cannot be recognized. By combining tissue patterns with these clinical features, researchers can explore how specific microenvironment layouts relate to real-world outcomes, such as which patients fare better or worse.
Putting AI to the test on the new dataset
To show that the dataset is truly useful, the scientists trained three different deep learning models—modern pattern-recognition systems that excel at image tasks—to identify the eight tissue types in the tiles. They used strict rules to split patients between training and testing groups so the models would be judged on patients they had never seen before. The models, including both classic image networks and a newer “vision transformer” design, all achieved very high accuracy, with performance scores close to perfect on several test sets. The team also compared results to other advanced image segmentation methods and found similarly strong performance. Visual tools were used to highlight which parts of the tissue the models relied on, confirming that they focused on medically meaningful regions rather than random patterns.
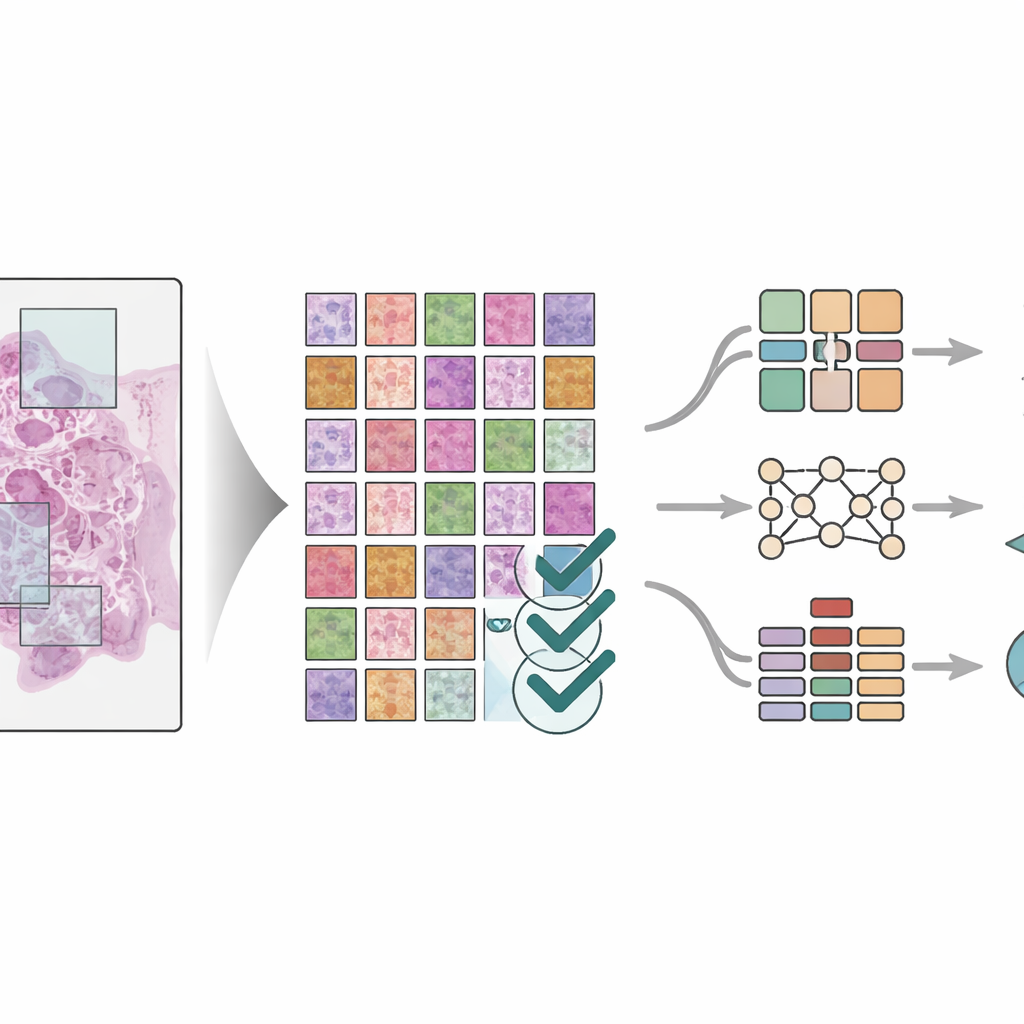
What this means for future cancer care
For non-specialists, the key message is that this work does not introduce a new treatment, but rather a powerful foundation for smarter diagnosis and prognosis. By sharing a large, well-organized, and openly available image library tied to detailed patient records, the authors enable researchers worldwide to build and compare artificial intelligence tools on the same solid ground. Such tools could eventually help pathologists more quickly and consistently map the tumor neighborhood, predict which patients are at higher risk, and suggest more personalized treatment strategies. Although the current data capture only single time points rather than changes over months or years, this resource is an important step toward using digital pathology and AI to better understand, and ultimately better treat, colorectal cancer.
Citation: Wang, H., Li, H., Xue, J. et al. Large-Scale Histological Image Dataset with Metadata for Colorectal Cancer Microenvironment. Sci Data 13, 431 (2026). https://doi.org/10.1038/s41597-026-06675-9
Keywords: colorectal cancer, tumor microenvironment, digital pathology, deep learning, medical imaging dataset