Clear Sky Science · en
A chromosome level genome assembly of Homatula variegata from the Yangtze River basin
A tiny stream fish with a big genetic story
In the fast-flowing, stony streams that feed China’s upper Yangtze River lives a small, striped loach called Homatula variegata. It is prized both for the table and for home aquariums, yet until now almost nothing was known about its DNA blueprint. This study delivers the first nearly complete, chromosome-by-chromosome map of the species’ genome, opening the door to smarter conservation, more efficient breeding, and deeper insight into how life adapts to cold, rushing mountain waters.
Why map the DNA of a modest-looking fish?
Although this loach reaches only about 14 centimeters, it plays an outsized role in local river ecosystems and regional aquaculture. It thrives in mid‑elevation streams with pebbly bottoms and steady currents, feeding on insects, organic debris, and small fish. Because it is both tasty and colorful, there is growing interest in farming it as a native ornamental and food species. Yet breeding and protecting a species are far harder without a precise genetic reference. A complete genome acts like a detailed parts list and wiring diagram, revealing the genes that shape growth, color, disease resistance, and the ability to cope with swift, cool water. Until now, no loach in this branch of the fish family had such a high‑quality reference, leaving a major blind spot in freshwater fish genetics.
Building a chromosome-by-chromosome DNA map
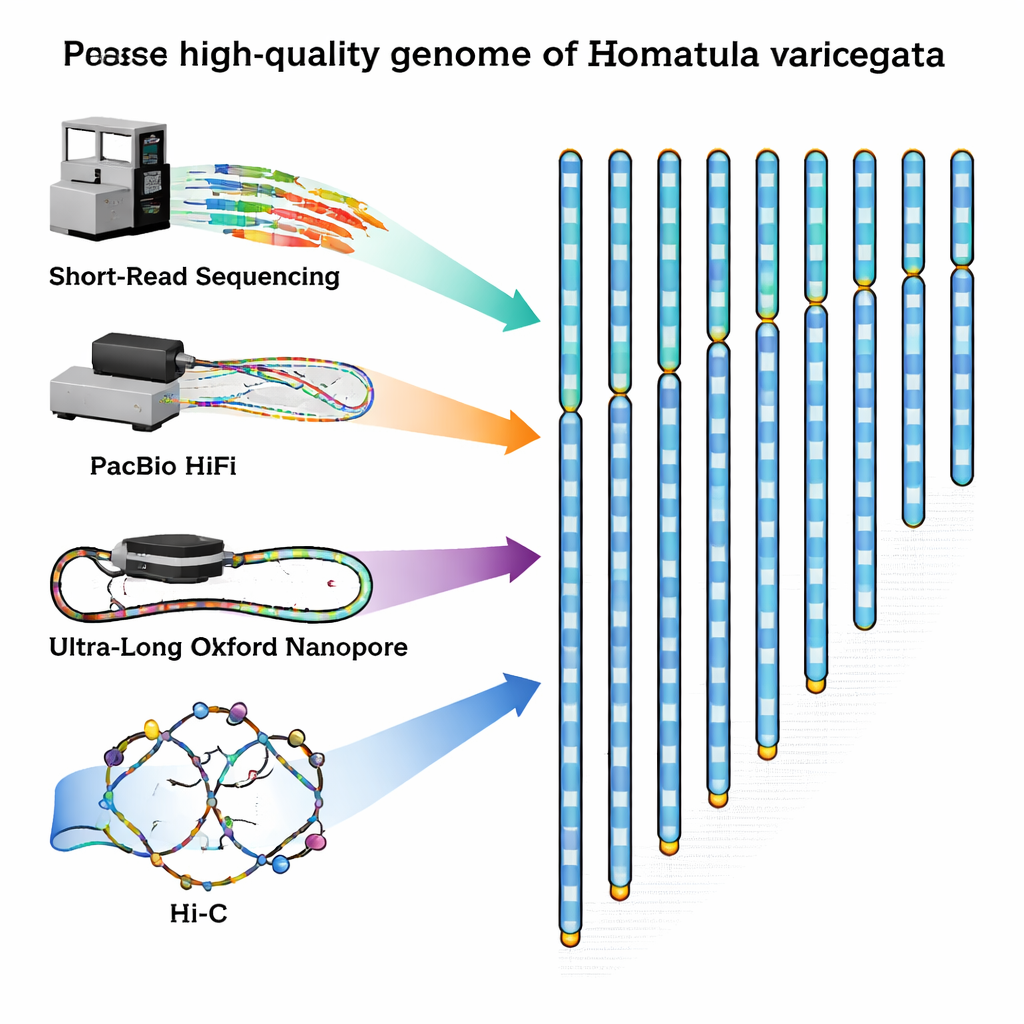
To fill this gap, the researchers captured a healthy adult male loach from the Qingyi River, a tributary of the Yangtze, and carefully extracted DNA and RNA from its blood. They then combined several state‑of‑the‑art sequencing methods, each with different strengths. Short‑read machines from Illumina produced huge numbers of highly accurate snippets of DNA. PacBio HiFi technology delivered somewhat longer fragments with excellent accuracy, while Oxford Nanopore devices generated ultra‑long strands that can stretch across hard‑to‑decode, repetitive regions. Finally, a method called Hi‑C captured how DNA strands fold and interact inside the cell nucleus, providing a 3‑D contact map that helps link fragments into full chromosomes in the correct order.
What the new genome reveals
By weaving these data together with modern assembly software and careful quality checks, the team produced a genome 641 million DNA letters long, neatly organized into 24 chromosomes. Remarkably, each chromosome is assembled as a single continuous piece, with 22 having no gaps at all and only tiny gaps in two others. They could pinpoint 24 likely centromeres—the central “belt” of each chromosome—and detect most of the protective telomere caps at chromosome ends. The scientists cataloged 24,479 protein‑coding genes, and were able to assign likely functions to about 93% of them by comparing against large international databases. They also mapped the landscape of repeated DNA, finding that more than a quarter of the genome consists of mobile genetic elements, especially DNA transposons, which can jump around the genome and sometimes drive evolution.
Testing the quality under the hood
High‑level numbers are only meaningful if the underlying map is trustworthy. The team therefore pushed the assembly through a battery of tests. Reads from all sequencing platforms aligned back to the new genome at very high rates, with even coverage across most of the chromosomes. Independent tools that count short DNA patterns suggested that nearly all expected sequence content is present, and standard gene completeness tests showed that the vast majority of universal fish genes are intact. The Hi‑C contact maps formed clean, square patterns along each chromosome, with little stray signal between them, indicating that segments are properly joined rather than scrambled.
From DNA map to real-world impact
To a non‑specialist, this work may sound like a technical triumph for its own sake, but its implications are practical and broad. Having a near end‑to‑end genome for Homatula variegata gives scientists a reference against which they can compare wild populations, track genetic diversity, and spot signs of inbreeding or local adaptation. Breeders can search for DNA markers linked to desirable traits such as fast growth, hardiness, or striking coloration, speeding up selective breeding while preserving the species’ natural character. Ecologists can explore how this loach has evolved to live in cool, fast‑moving mountain streams, lessons that may also inform the management of related species. In short, this chromosome‑level genome turns a modest river fish into a powerful model for understanding—and protecting—the rich life of Asian freshwater ecosystems.
Citation: Tang, Y., Wu, Q., Wang, Y. et al. A chromosome level genome assembly of Homatula variegata from the Yangtze River basin. Sci Data 13, 303 (2026). https://doi.org/10.1038/s41597-026-06667-9
Keywords: fish genome, chromosome assembly, freshwater biodiversity, aquaculture genetics, Yangtze River loach