Clear Sky Science · en
BactoTraits: a trait database for exploring functional diversity of bacterial communities
Why tiny microbes matter for big environmental questions
Bacteria are everywhere: in soils, rivers, oceans, and even inside our bodies. These microscopic residents help clean up pollution, recycle nutrients, and support plant and animal life. Yet most studies still treat bacterial species as simple names on a list, without asking what they actually do. This article introduces BactoTraits, a large open dataset that turns scattered lab information about bacteria into practical "trait" profiles, helping scientists link who is present in a habitat to how that community functions and responds to environmental change.
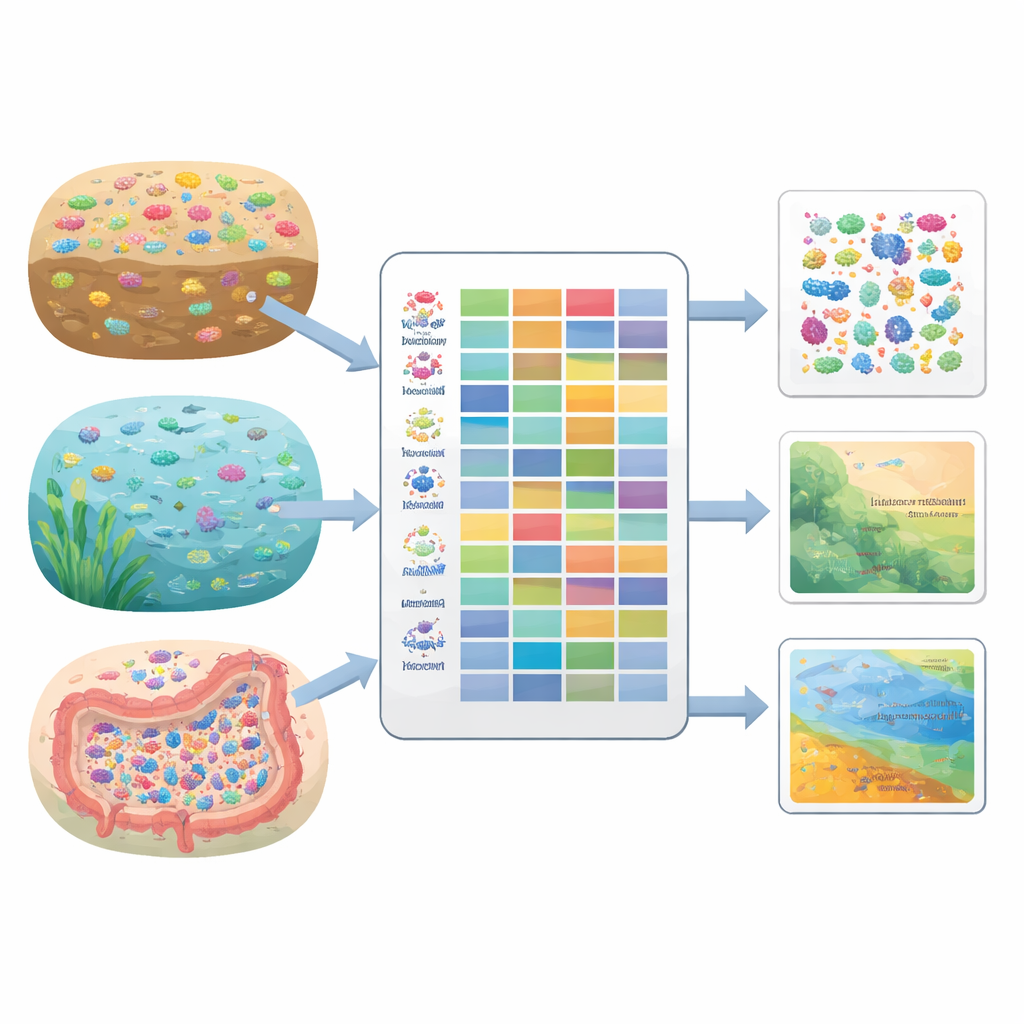
From plant traits to bacterial traits
Ecologists have long used traits—features such as leaf size or seed mass—to understand how plants cope with drought, pollution, or warming. Similar trait-based approaches exist for animals and soil invertebrates, and they have made it easier to predict how communities change under human pressures. For microbes, though, this way of thinking is still catching up, even though bacteria respond quickly to disturbance and can act as early-warning indicators of ecosystem problems. BactoTraits adapts this trait mindset to bacteria, defining traits as characteristics that influence how well strains survive, grow, and interact with their environment.
Building a trait atlas for tens of thousands of strains
The authors compiled BactoTraits by mining three major open resources: BacDive, a metadatabase describing cultivated bacterial strains; rrnDB, which lists how many copies of the ribosomal RNA gene each strain carries; and genomesizeR, which predicts genome sizes from sequence records. From these sources they extracted information for 100,866 strains and converted it into 31 functional traits for 97,721 strains that had at least some usable data. These traits cover basic cell features (such as shape, size, and ability to form spores), environmental preferences (for temperature, salt, and pH), lifestyle and metabolism (such as oxygen use, energy and carbon sources, pigment and antibiotic resistance), and genomic properties (GC content, gene copy number, and estimated genome size).
Turning messy records into usable trait profiles
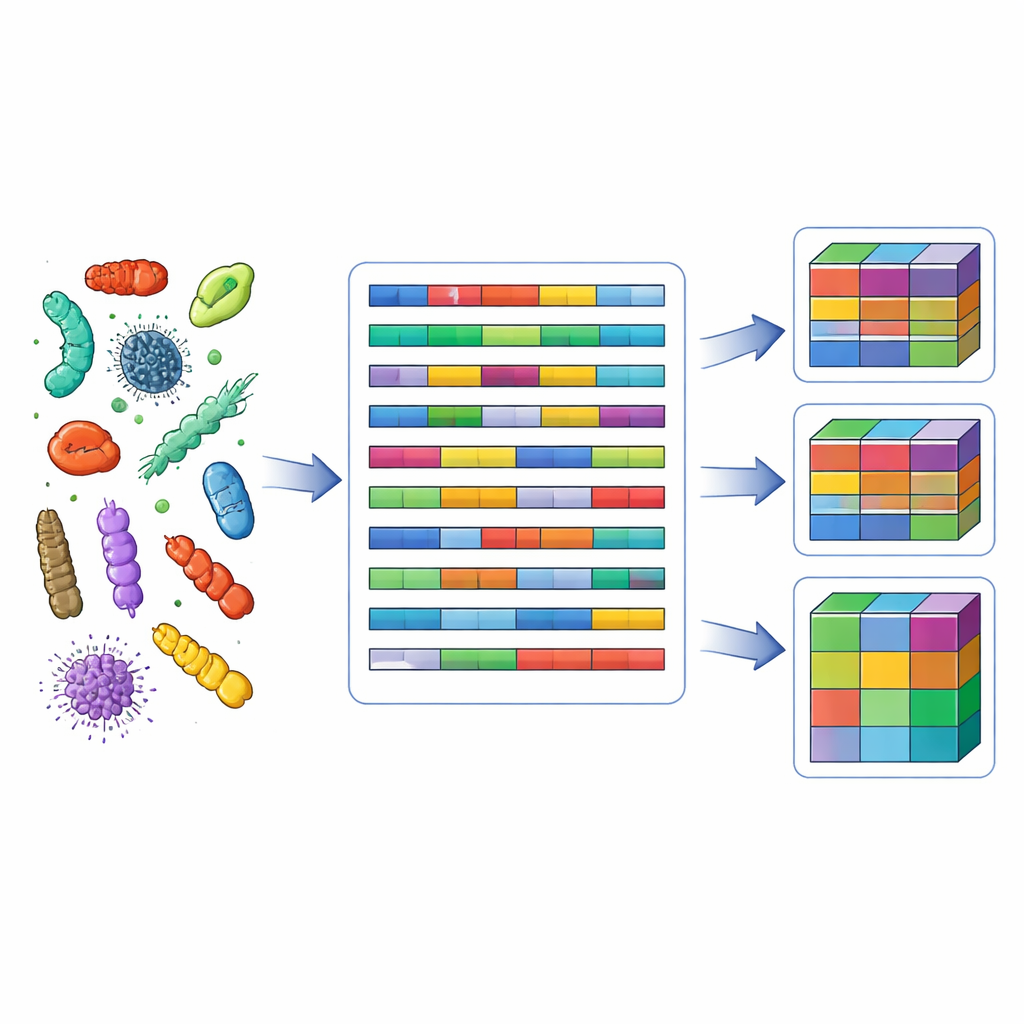
Data in the original databases are uneven and sometimes contradictory: one study may report a strain as motile, another as non-motile. The team addressed this by harmonizing terminology and then using a "fuzzy" coding approach. Instead of forcing each strain into a single category, they allowed it to have partial membership in several trait classes. For example, if most studies describe a strain as non-motile and a few describe it as motile, the strain’s profile reflects both possibilities with different weights that sum to one. Quantitative values such as temperature or pH were grouped into clear ranges defined from literature and data distributions, balancing biological meaning with the need to keep enough strains in each class. The result is a matrix where each strain is linked to a graded trait profile that captures both knowledge and uncertainty.
Linking DNA surveys to what bacteria can do
Modern environmental microbiology often relies on high-throughput sequencing of a marker gene (16S rRNA) to list bacterial types in samples from soil, water, or host organisms. On its own, that list says little about function. BactoTraits bridges this gap. The authors provide a series of R scripts that match each sequenced unit (OTU or amplicon variant) to strains in the database via taxonomic information from the SILVA reference set. When there are multiple matching strains, their trait profiles are averaged. If matches fail at the genus level, the scripts move stepwise up to family, order, class, or phylum, always noting which level was used. Finally, for each environmental sample, the scripts calculate a community-weighted mean profile: how strongly the entire bacterial community expresses each trait, taking into account both trait values and relative abundances.
How researchers can use this new resource
The BactoTraits dataset and scripts are designed to be transparent, flexible, and easy to update as BacDive, SILVA, rrnDB, and genome records grow. Researchers can combine BactoTraits with existing gene-based prediction tools to get a richer picture of communities: not just which metabolic pathways might be present, but also how bacteria differ in size, shape, stress tolerance, growth strategy, and potential pathogenicity. Earlier work using a previous version of the dataset has already shown that certain trait combinations can signal metal or hydrocarbon contamination in soils, or the coexistence of aerobic and anaerobic bacteria in oil-affected environments. The expanded version now covers far more strains, traits, and taxonomic levels, making such applications more robust.
What this means for understanding living communities
To a lay reader, the core message is that BactoTraits turns a huge mass of scattered microbiology facts into a coherent map of how bacteria live and behave. By connecting routine DNA surveys to concrete features like temperature preference, salt tolerance, or ability to resist antibiotics, it becomes possible to track not only which bacteria are present, but how their collective abilities shift under pollution, climate change, or management actions. This can improve biomonitoring, guide conservation and restoration, and help scientists test ideas about how microbial communities are assembled. In short, BactoTraits provides a powerful new lens for seeing the hidden workings of bacterial life across ecosystems.
Citation: Laderriere, V., Usseglio-Polatera, P., Maunoury-Danger, F. et al. BactoTraits: a trait database for exploring functional diversity of bacterial communities. Sci Data 13, 337 (2026). https://doi.org/10.1038/s41597-026-06652-2
Keywords: bacterial traits, microbial ecology, functional diversity, environmental DNA, biomonitoring