Clear Sky Science · en
An annotated dataset of Gram stains from positive blood cultures
Why fast infection answers matter
When bacteria or fungi slip into the bloodstream, every hour without the right treatment can mean the difference between life and death. Doctors rely on a quick laboratory test called a Gram stain to see what kind of germ is present and to choose early antibiotics. But reading these stained microscope slides is a skilled, manual job that takes time and can vary from one technologist to another. This study describes a new, carefully annotated image collection of real hospital blood culture slides, built to help computers learn to read Gram stains automatically and support faster, more reliable care.
Turning real hospital slides into data
The researchers gathered 57 different kinds of bacteria and fungi that had been grown from patients’ positive blood culture bottles as part of everyday hospital work. From January to May 2024, once a blood culture signaled positive, staff prepared Gram-stained smears on glass slides and confirmed the exact species using a high-precision identification method called MALDI-TOF mass spectrometry. Without changing normal routines or collecting extra samples, the team then captured high-resolution digital images of typical fields under a 100× oil-immersion microscope, resulting in 505 large color images that mirror what technologists see in real practice.
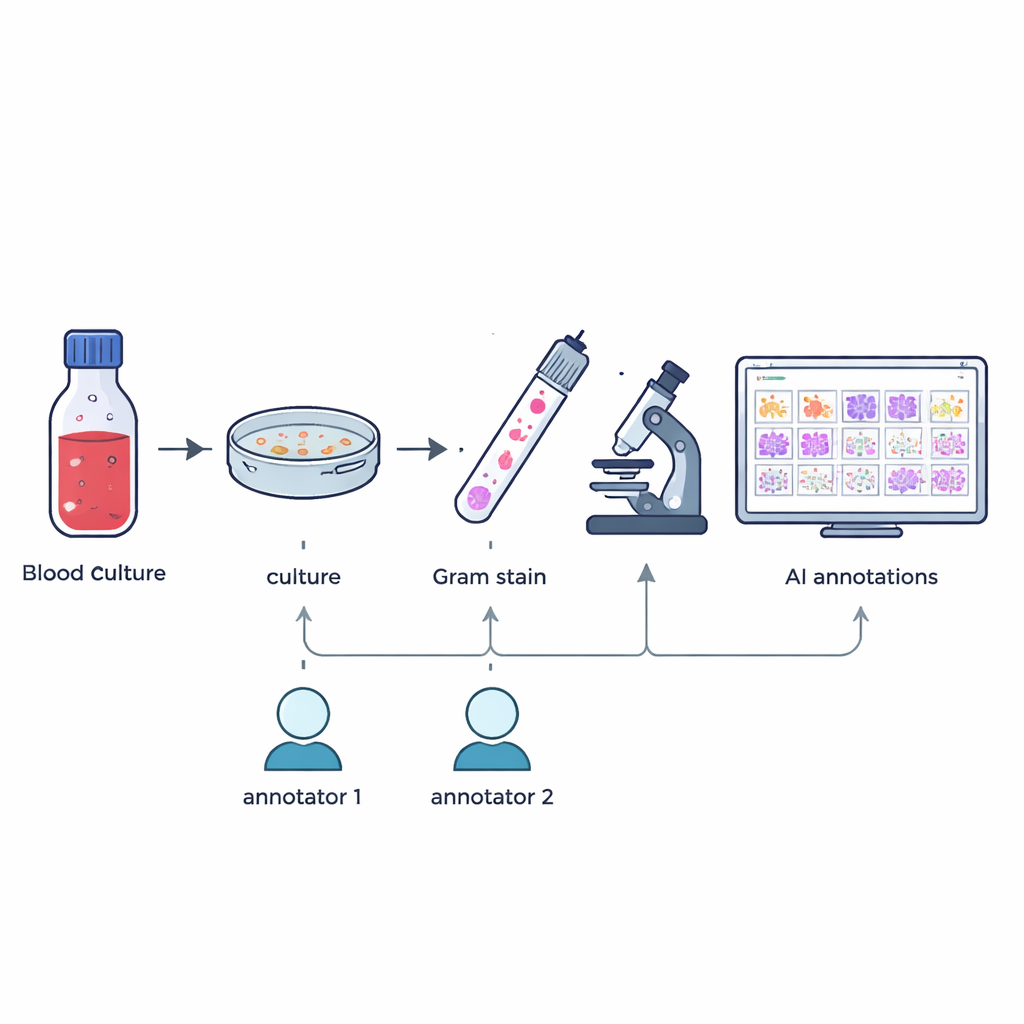
Careful labeling of tiny shapes
Building a useful teaching set for artificial intelligence means knowing exactly where each microbe is in each picture. Two experienced microbiology technologists independently drew boxes around individual microbial cells or clusters in every image, guided only by what they saw under the microscope. A custom software tool compared the two sets of markings: boxes that overlapped enough were merged, and any mismatches or disagreements were flagged. A senior expert with more than 20 years of experience then reviewed these cases by hand. This multi-step process produced 7,528 checked annotations that highlight cocci (round cells), bacilli (rod-shaped cells), and fungi, while leaving out partial or doubtful objects.
What the dataset contains
The finished resource combines several layers of information. All 505 images are provided as high-resolution JPEG files, and the final, expert-verified boxes are stored in standard COCO JSON format that is widely used in computer vision research. Extra files link each image to its microbe species, whether it is Gram-positive or Gram-negative, its broad shape group, the type of blood culture vial it came from, and how long the culture took to turn positive. Because each image contains only one species, all boxes within a given image share the same biological traits. Users can choose between a single large annotation file or separate files per image, and a simple Python script is included to visualize any image with its boxes overlaid.
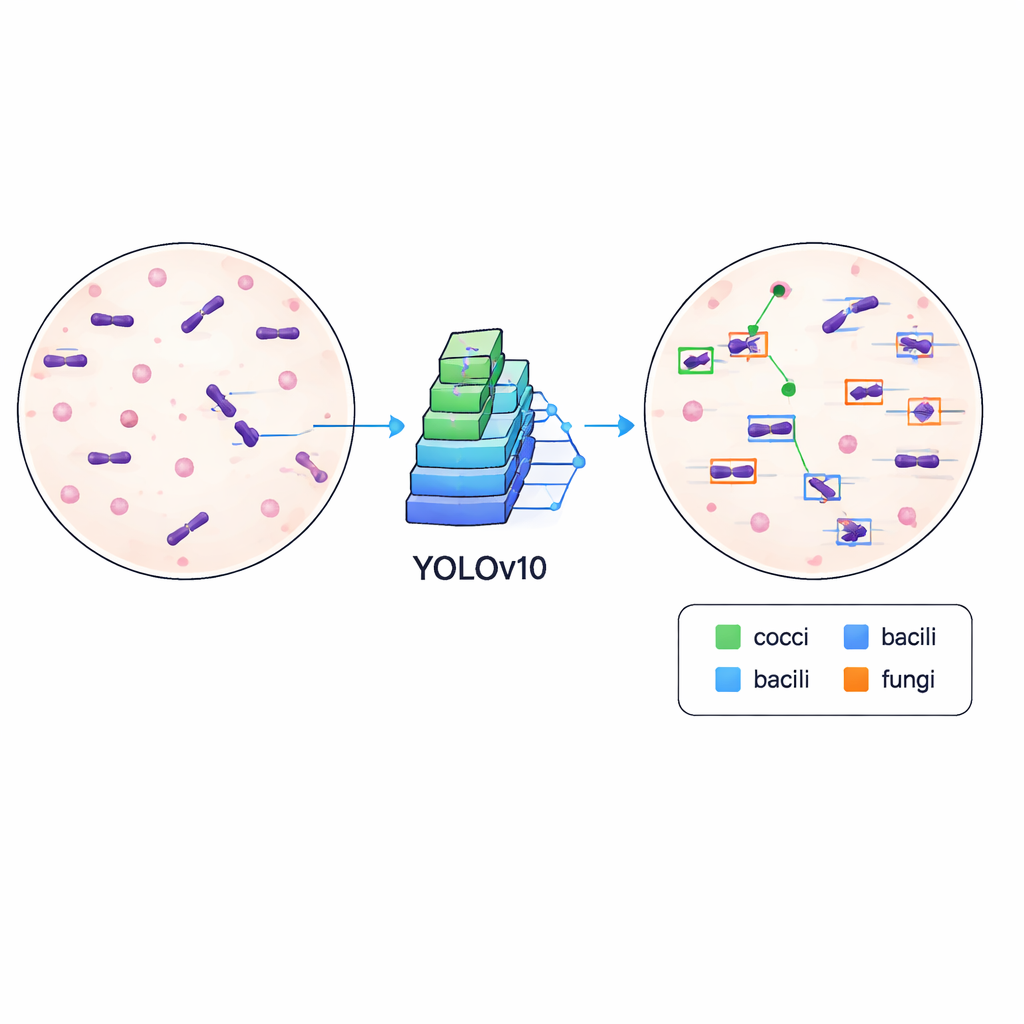
Teaching computers to spot germs
To show that the dataset is not just tidy but also practical, the authors trained a modern object-detection algorithm known as YOLOv10 to find and classify microbes in the images. They split the data into training and validation sets and ran the model for 500 training rounds on a high-end graphics card, tracking how well it learned to draw accurate boxes and distinguish between different cell types. The trained system reached a mean average precision of about 84.6% at a standard matching threshold, indicating that it can reliably locate and label microbes across varied slide appearances, including differences in stain intensity, background debris, and focus.
How this resource can be used
Because the data follow common formats, they can plug into many existing computer vision pipelines. Researchers might first train a system simply to tell true microbes from debris, helping laboratories filter out false-positive culture signals. They can also group microbes into broad shapes, matching what clinicians need for an early, so-called “Tier 1” report that guides initial antibiotic choices. A more ambitious goal is to distinguish individual species by subtle visual cues. The authors note limits: some cells are grouped, some slides come from a single source per species, and focus can vary—just as in real life. Still, every included box has been carefully checked, making the dataset a trustworthy starting point.
What this means for patients
In plain terms, this work turns routine blood culture slides into a shared training ground for smart software. By making both the images and expert markings publicly available, the study lowers the barrier for teams around the world to build and test AI tools that can read Gram stains quickly and consistently. While such systems will not replace human microbiologists, they could help flag dangerous infections sooner, reduce interpretation errors, and support better use of antibiotics. For patients, that could translate into faster, more precise treatment when it matters most.
Citation: Yi, Q., Gou, X., Zhu, R. et al. An annotated dataset of Gram stains from positive blood cultures. Sci Data 13, 294 (2026). https://doi.org/10.1038/s41597-026-06651-3
Keywords: bloodstream infections, Gram stain, medical imaging dataset, artificial intelligence, microbiology diagnostics