Clear Sky Science · en
Chromosome-level genome assembly of the stonefly Rhopalopsole triangulispina Mo and Li, 2025 (Plecoptera: Leuctridae)
Why a Tiny Stream Insect Matters
In rushing mountain streams around the world, small, delicate insects called stoneflies quietly signal how healthy the water is. Their young stages, or nymphs, are so sensitive to pollution that their presence often means the stream is clean and well-oxygenated. Yet, despite this ecological importance, scientists have known surprisingly little about the detailed genetic makeup of many stoneflies. This study delivers a high-quality, chromosome-level map of the DNA of one such species, Rhopalopsole triangulispina, opening a new window into how these insects evolved and how they can help us better protect freshwater ecosystems.
Life in Clear, Fast-Flowing Water
Stoneflies in the superfamily Nemouroidea are among the most diverse and abundant groups of these insects, living in cold, clean streams, lakes, and ponds—often hidden among rocks or sediments. Their nymphs depend on good water quality: temperature, oxygen levels, chemical pollution, and even the type of streambed can make the difference between thriving and disappearing. Because of this sensitivity, scientists use them as natural “environmental sensors” when assessing river and stream health. Within Nemouroidea, the family Leuctridae and the genus Rhopalopsole are especially common in Asian mountain regions, including China, which harbors more than 60 known Rhopalopsole species. Despite this diversity, genetic reference maps, especially fully annotated genomes, have been missing, limiting our ability to study their evolution or use genomic tools in water-quality monitoring.
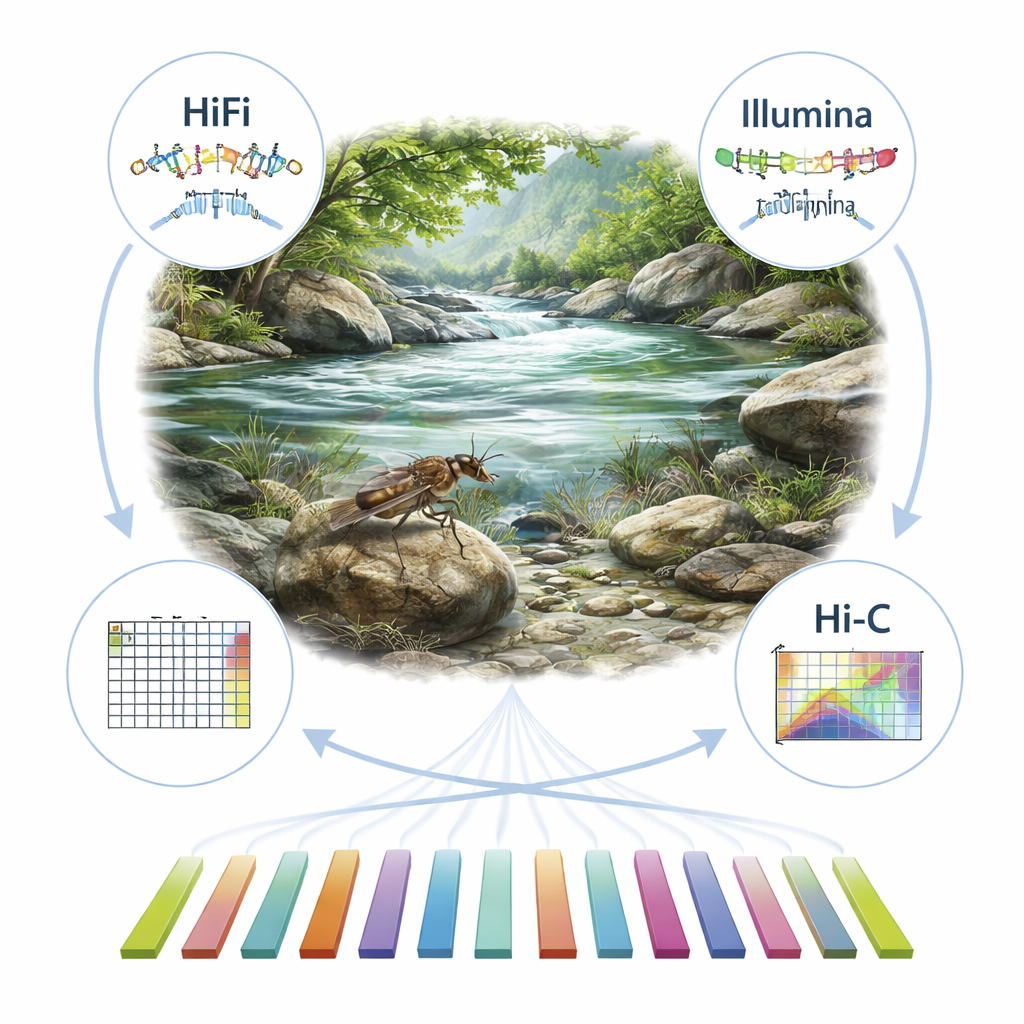
Building a Genetic Map from Scratch
To tackle this gap, the researchers collected adult Rhopalopsole triangulispina from a protected mountain reserve in southern China. After carefully preserving the insects, they extracted high-quality DNA and RNA (the molecule that reflects which genes are active). Using multiple cutting-edge sequencing technologies, they generated different views of the genome: very accurate long DNA reads (PacBio HiFi), massive numbers of short reads (Illumina), and special data that capture how pieces of DNA are folded and interact inside the cell nucleus (Hi-C). Together, these data types allowed the team to assemble the insect’s genetic material into long, continuous sequences, and then use the Hi-C information to arrange them into 13 chromosome-like structures, called pseudochromosomes.
What the Genome Reveals
The finished genome is about 347 million “letters” of DNA long, with nearly 97% of it placed onto the 13 pseudochromosomes. Quality checks showed that the assembly is both highly complete and extremely accurate, with error rates estimated at less than one in ten million bases and over 95% of all sequencing reads mapping back to it. Almost half of the genome consists of repetitive DNA—stretches that occur many times and often reflect mobile genetic elements, or “jumping genes,” that have shaped the genome over evolutionary time. On top of this foundation, the team predicted 12,857 protein-coding genes and more than 2,400 non-coding RNA genes, which include molecules involved in building ribosomes, processing RNA, and guiding protein production. By comparing these genes to large international databases, they were able to assign likely functions to the vast majority of them and connect many to known biological pathways.
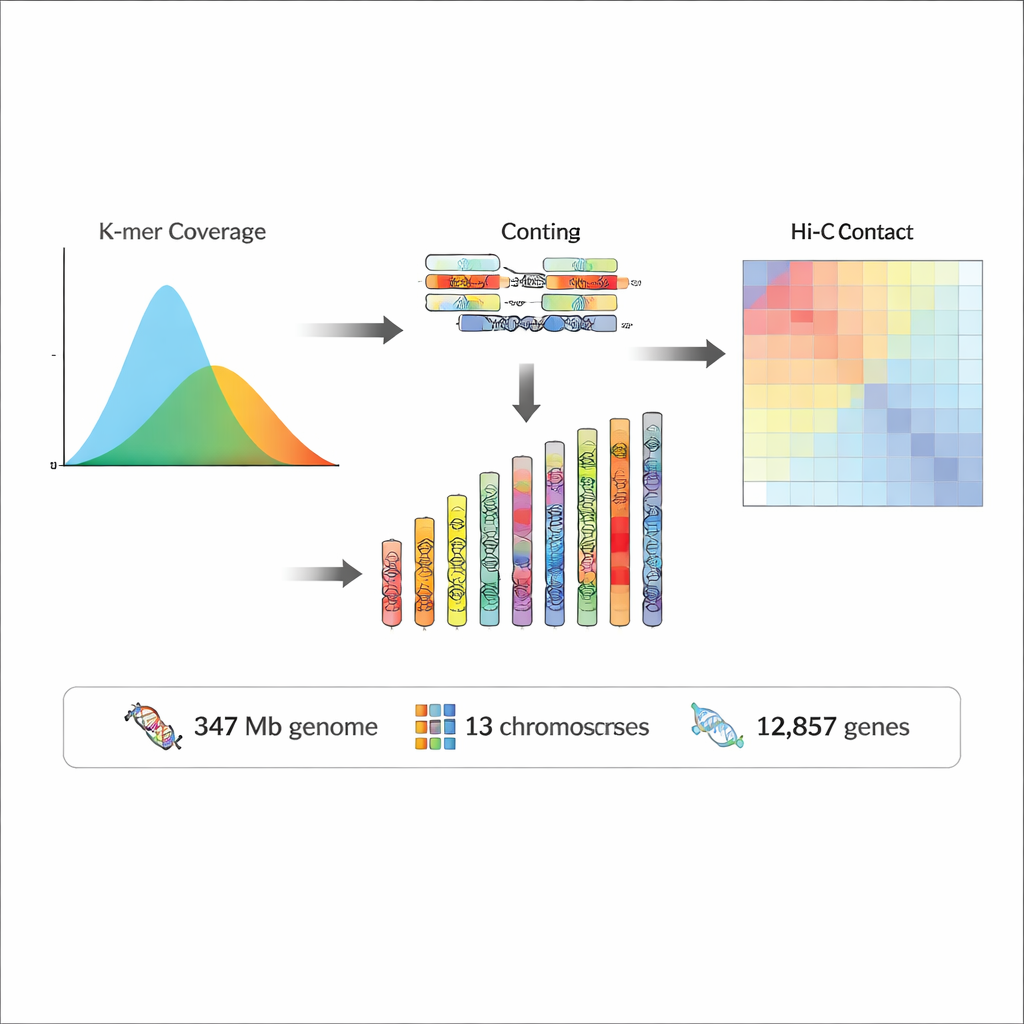
A New Resource for Evolution and Ecology
Beyond cataloging genes, this genome acts as a reference book for future studies. Scientists can now investigate which genes help stoneflies adapt to cold, fast-moving water, how they respond at the molecular level to pollution or climate change, and how different stonefly lineages are related to one another. Until now, most evolutionary work on Nemouroidea relied on physical traits, mitochondrial DNA, or partial gene sets. This complete, annotated genome offers a much richer and more precise basis for reconstructing the stonefly family tree and for comparing genomes across insects, including those with very different lifestyles. Because all raw data and annotations are publicly available, researchers worldwide can reanalyze the information, integrate it with other datasets, and build new tools for biomonitoring.
From DNA Blueprint to Cleaner Streams
For non-specialists, the key takeaway is that we now have a detailed, chromosome-level blueprint of a stonefly species that serves as a frontline guardian of freshwater health. This genome will help scientists understand how these insects live and adapt, trace their deep evolutionary history, and design more sensitive, DNA-based methods to detect them in rivers and streams. In practice, that means better early-warning systems for pollution and environmental change. By uncovering the genetic underpinnings of a small insect in mountain streams, this work ultimately contributes to safeguarding the clean water on which people and ecosystems depend.
Citation: Lin, A., Cao, J., Murányi, D. et al. Chromosome-level genome assembly of the stonefly Rhopalopsole triangulispina Mo and Li, 2025 (Plecoptera: Leuctridae). Sci Data 13, 292 (2026). https://doi.org/10.1038/s41597-026-06631-7
Keywords: stonefly genome, freshwater bioindicators, chromosome-level assembly, insect evolution, environmental DNA