Clear Sky Science · en
Global deep-sea hydrothermal deposit metagenomes and metagenome-assembled genomes over time and space
Life in the darkest parts of the ocean
Far below the reach of sunlight, deep-sea hot springs called hydrothermal vents create oases of life on the otherwise barren seafloor. These sites host strange, heat-loving microbes that help drive global chemical cycles and may even resemble some of Earth’s earliest life forms. The study described here does not focus on a single new organism, but instead delivers a massive, long-term genetic catalog of the microbes that thrive in these extreme environments—an open resource that will fuel discoveries about evolution, biotechnology, and the future impacts of deep-sea mining.
Hidden hot springs around the world
Hydrothermal vents form where seawater seeps into the ocean crust, heats up, and rises back to the seafloor loaded with metals and gases. As this hot fluid meets cold seawater, it builds chimney-like mineral deposits that are quickly colonized by microbes that can use chemicals instead of sunlight for energy. Until recently, most of these heat-loving bacteria and archaea were known only from fragments of a single gene, leaving scientists largely in the dark about what they can do. This new effort brings together samples collected across 21 vent fields in the Atlantic, Pacific, and Indian oceans over 16 years, turning scattered expeditions into a coherent global time series. 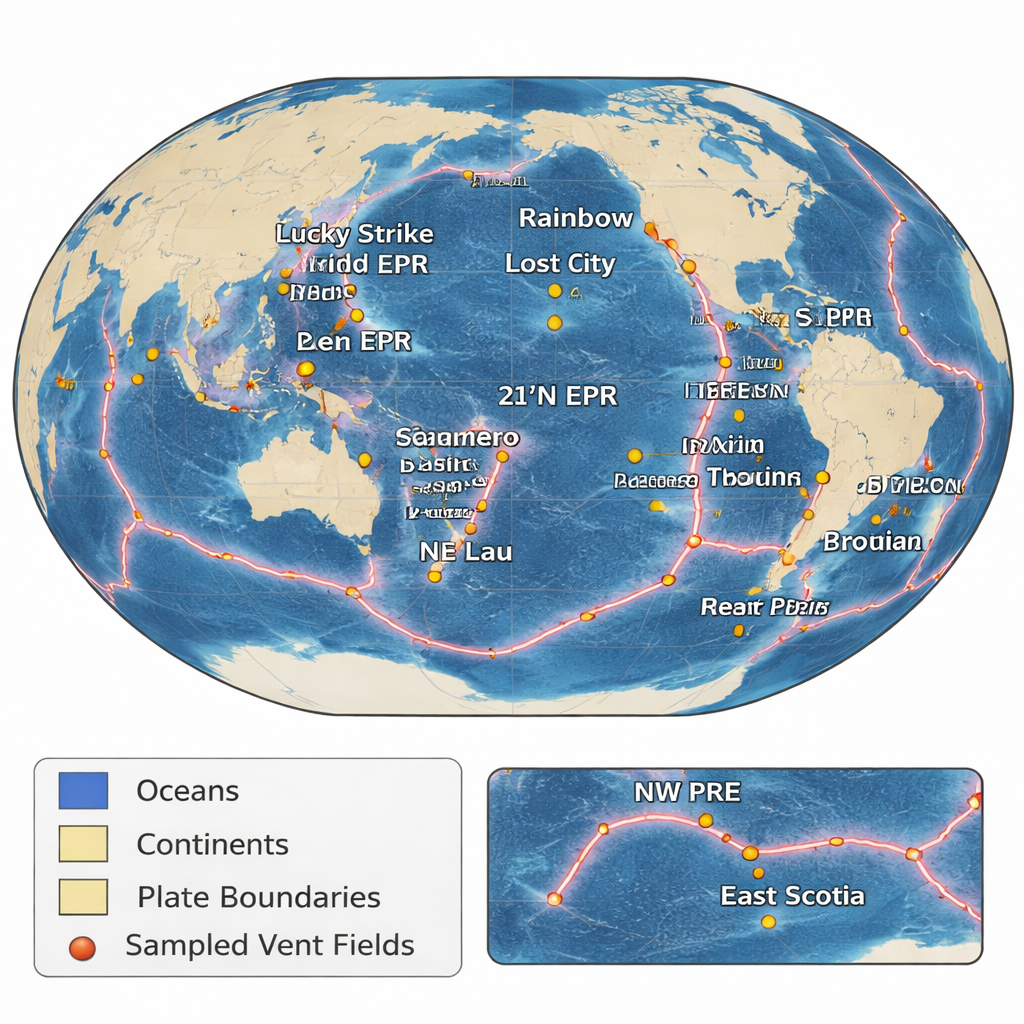
Turning seafloor minerals into genomes
To see which microbes live in these deposits and what functions they carry out, the team used a “metagenomic” approach: instead of trying to grow each species in the lab, they extracted all the DNA directly from the vent minerals. High-throughput sequencing produced an enormous 3.56 trillion base pairs of raw data from 70 samples. After careful quality control to remove low-quality and contaminant sequences, sophisticated software stitched the remaining DNA pieces into longer fragments and then grouped them into thousands of draft genomes, called metagenome-assembled genomes, or MAGs. This stepwise process—from chimney sampling to DNA sequencing to genome reconstruction—creates a kind of molecular census of each vent community.
A vast family album for deep-sea microbes
The result is the DSV70 dataset: 7,422 medium- to high-quality genomes from heat-loving bacteria and archaea. These genomes span at least 16 archaeal groups and 85 bacterial groups, including many lineages that are poorly represented—or entirely absent—in existing databases. Some microbial groups known to be abundant at vents, such as Campylobacterota and certain Proteobacteria, are especially well covered, but the dataset also greatly expands the genomic coverage of underexplored archaeal branches like Thermoproteota and the tiny-celled DPANN lineages. In total, more than 29 million predicted proteins were identified, with many linked to known metabolic functions. This provides a treasure trove for understanding how vent microbes harness chemical energy, recycle elements, and interact with one another and with vent chemistry. 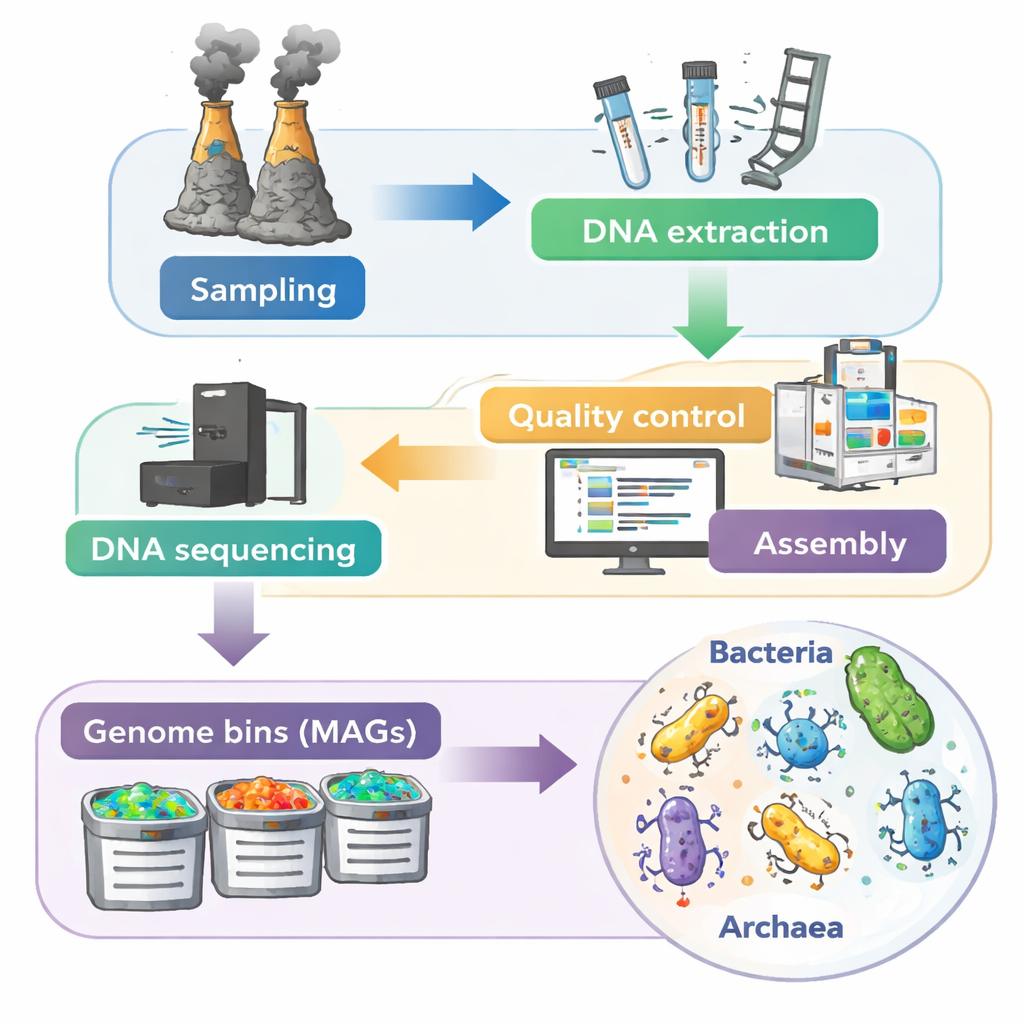
Tracking change over space, time, and temperature
Because the samples come from many locations and span more than a decade, the dataset allows scientists to ask how vent microbial communities change over time and across different geological settings. The authors combined their new sequences with earlier metagenome collections from similar habitats to create a statistically powerful foundation for comparing vents on different ridges and back-arc basins. Researchers can now explore questions such as which microbes are found everywhere versus only at certain vents, how closely related deep-sea microbes are to those in terrestrial hot springs, and how community structure might respond to events like eruptions, natural climate shifts, or human activities such as seafloor mining.
Why this genomic atlas matters
All raw sequences, assembled DNA fragments, and reconstructed genomes have been deposited in public repositories, effectively creating a global reference library for deep-sea hydrothermal microbes. For non-specialists, the key takeaway is that scientists now have a detailed genetic snapshot of life in some of Earth’s most extreme, and increasingly threatened, environments. This resource will help researchers identify new enzymes for industrial and medical applications, refine the microbial branches of the tree of life, and establish a baseline for monitoring how deep-sea ecosystems may change as the ocean warms and mining interests advance into the abyss.
Citation: St. John, E., Reysenbach, AL. Global deep-sea hydrothermal deposit metagenomes and metagenome-assembled genomes over time and space. Sci Data 13, 283 (2026). https://doi.org/10.1038/s41597-026-06612-w
Keywords: deep-sea hydrothermal vents, microbial genomes, metagenomics, extremophiles, ocean biodiversity